Abstract
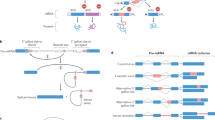
The Down syndrome cell adhesion molecule (Dscam) gene has essential roles in neural wiring and pathogen recognition in Drosophila melanogaster. Dscam encodes 38,016 distinct isoforms via extensive alternative splicing. The 95 alternative exons in Dscam are organized into clusters that are spliced in a mutually exclusive manner. The exon 6 cluster contains 48 variable exons and uses a complex system of competing RNA structures to ensure that only one variable exon is included. Here we show that the heterogeneous nuclear ribonucleoprotein hrp36 acts specifically within, and throughout, the exon 6 cluster to prevent the inclusion of multiple exons. Moreover, hrp36 prevents serine/arginine-rich proteins from promoting the ectopic inclusion of multiple exon 6 variants. Thus, the fidelity of mutually exclusive splicing in the exon 6 cluster is governed by an intricate combination of alternative RNA structures and a globally acting splicing repressor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Blencowe, B.J. Alternative splicing: new insights from global analyses. Cell 126, 37–47 (2006).
Xing, Y. & Lee, C. Alternative splicing and RNA selection pressure–evolutionary consequences for eukaryotic genomes. Nat. Rev. Genet. 7, 499–509 (2006).
Black, D.L. Protein diversity from alternative splicing: a challenge for bioinformatics and post-genome biology. Cell 103, 367–370 (2000).
Graveley, B.R. Alternative splicing: increasing diversity in the proteomic world. Trends Genet. 17, 100–107 (2001).
Schmucker, D. et al. Drosophila Dscam is an axon guidance receptor exhibiting extraordinary molecular diversity. Cell 101, 671–684 (2000).
Zipursky, S.L., Wojtowicz, W.M. & Hattori, D. Got diversity? Wiring the fly brain with Dscam. Trends Biochem. Sci. 31, 581–588 (2006).
Hummel, T. et al. Axonal targeting of olfactory receptor neurons in Drosophila is controlled by Dscam. Neuron 37, 221–231 (2003).
Zhu, H. et al. Dendritic patterning by Dscam and synaptic partner matching in the Drosophila antennal lobe. Nat. Neurosci. 9, 349–355 (2006).
Wang, J. et al. Transmembrane/juxtamembrane domain-dependent Dscam distribution and function during mushroom body neuronal morphogenesis. Neuron 43, 663–672 (2004).
Wang, J., Zugates, C.T., Liang, I.H., Lee, C.H. & Lee, T. Drosophila Dscam is required for divergent segregation of sister branches and suppresses ectopic bifurcation of axons. Neuron 33, 559–571 (2002).
Zhan, X.L. et al. Analysis of Dscam diversity in regulating axon guidance in Drosophila mushroom bodies. Neuron 43, 673–686 (2004).
Chen, B.E. et al. The molecular diversity of Dscam is functionally required for neuronal wiring specificity in Drosophila. Cell 125, 607–620 (2006).
Watson, F.L. et al. Extensive diversity of Ig-superfamily proteins in the immune system of insects. Science 309, 1874–1878 (2005).
Dong, Y., Taylor, H.E. & Dimopoulos, G. AgDscam, a hypervariable immunoglobulin domain-containing receptor of the Anopheles gambiae innate immune system. PLoS Biol. 4, e229 (2006).
Smith, C.W. Alternative splicing–when two's a crowd. Cell 123, 1–3 (2005).
Anastassiou, D., Liu, H. & Varadan, V. Variable window binding for mutually exclusive alternative splicing. Genome Biol. 7, R2 (2006).
Graveley, B.R. Mutually exclusive splicing of the insect Dscam pre-mRNA directed by competing intronic RNA secondary structures. Cell 123, 65–73 (2005).
Fan, J.B. et al. A versatile assay for high-throughput gene expression profiling on universal array matrices. Genome Res. 14, 878–885 (2004).
Yeakley, J.M. et al. Profiling alternative splicing on fiber-optic arrays. Nat. Biotechnol. 20, 353–358 (2002).
Park, J.W., Parisky, K., Celotto, A.M., Reenan, R.A. & Graveley, B.R. Identification of alternative splicing regulators by RNA interference in Drosophila. Proc. Natl. Acad. Sci. USA 101, 15974–15979 (2004).
Zhu, J., Mayeda, A. & Krainer, A.R. Exon identity established through differential antagonism between exonic splicing silencer-bound hnRNP A1 and enhancer-bound SR proteins. Mol. Cell 8, 1351–1361 (2001).
Matlin, A.J., Clark, F. & Smith, C.W. Understanding alternative splicing: towards a cellular code. Nat. Rev. Mol. Cell Biol. 6, 386–398 (2005).
Graveley, B.R. Sorting out the complexity of SR protein functions. RNA 6, 1197–1211 (2000).
Shi, H., Hoffman, B.E. & Lis, J.T. A specific RNA hairpin loop structure binds the RNA recognition motifs of the Drosophila SR protein B52. Mol. Cell. Biol. 17, 2649–2657 (1997).
Haynes, S.R., Johnson, D., Raychaudhuri, G. & Beyer, A.L. The Drosophila Hrb87F gene encodes a new member of the A and B hnRNP protein group. Nucleic Acids Res. 19, 25–31 (1991).
Zu, K., Sikes, M.L., Haynes, S.R. & Beyer, A.L. Altered levels of the Drosophila HRB87F/hrp36 hnRNP protein have limited effects on alternative splicing in vivo. Mol. Biol. Cell 7, 1059–1073 (1996).
Neves, G., Zucker, J., Daly, M. & Chess, A. Stochastic yet biased expression of multiple Dscam splice variants by individual cells. Nat. Genet. 36, 240–246 (2004).
Hattori, D. et al. Dscam diversity is essential for neuronal wiring and self-recognition. Nature 449, 223–227 (2007).
Hughes, M.E. et al. Homophilic Dscam interactions control complex dendrite morphogenesis. Neuron 54, 417–427 (2007).
Matthews, B.J. et al. Dendrite self-avoidance is controlled by Dscam. Cell 129, 593–604 (2007).
Soba, P. et al. Drosophila sensory neurons require Dscam for dendritic self-avoidance and proper dendritic field organization. Neuron 54, 403–416 (2007).
Piñol-Roma, S., Swanson, M.S., Matunis, M.J. & Dreyfuss, G. Purification and characterization of proteins of heterogeneous nuclear ribonucleoprotein complexes by affinity chromatography. Methods Enzymol. 181, 326–331 (1990).
Dignam, J.D., Lebovitz, R.M. & Roeder, R.G. Accurate transcription initiation by RNA polymerase II in a soluble extract from isolated mammalian nuclei. Nucleic Acids Res. 11, 1475–1489 (1983).
Acknowledgements
We thank members of the Graveley lab for discussions and comments on the manuscript. This work was funded by the Crick-Jacobs Center for Theoretical and Computational Biology (G.W.Y.), a long-term fellowship from the Human Frontier Science Program to M.B., and grants from the US National Institutes of Health to D.C.R. (GM61987) and B.R.G. (GM62561 and GM67842).
Author information
Authors and Affiliations
Contributions
S.O., M.B., J.P., Y.S. and J.M.Y. performed the experiments; S.O., M.B., J.P., Y.S., J.M.Y., D.C.R. and B.R.G. designed the experiments; S.O., M.B., J.P., Y.S., J.M.Y., G.W.Y., D.C.R. and B.R.G. analyzed the data; B.R.G. wrote the paper with input from all authors.
Corresponding author
Ethics declarations
Competing interests
J.M.Y. is employed by Illumina, Inc. She also performed assays and provided reagents for the DASL assays.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 and Supplementary Discussion (PDF 1194 kb)
Rights and permissions
About this article
Cite this article
Olson, S., Blanchette, M., Park, J. et al. A regulator of Dscam mutually exclusive splicing fidelity. Nat Struct Mol Biol 14, 1134–1140 (2007). https://doi.org/10.1038/nsmb1339
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1339
This article is cited by
-
Acute thiamethoxam toxicity in honeybees is not enhanced by common fungicide and herbicide and lacks stress-induced changes in mRNA splicing
Scientific Reports (2019)
-
Revisiting Dscam diversity: lessons from clustered protocadherins
Cellular and Molecular Life Sciences (2019)
-
Predicted networks of protein-protein interactions in Stegodyphus mimosarum by cross-species comparisons
BMC Genomics (2017)
-
The determinants of alternative RNA splicing in human cells
Molecular Genetics and Genomics (2017)
-
Diverse regulation of 3′ splice site usage
Cellular and Molecular Life Sciences (2015)