Abstract
DNA methylation occurs in CG and non-CG sequence contexts. Non-CG methylation is abundant in plants and is mediated by CHROMOMETHYLASE (CMT) and DOMAINS REARRANGED METHYLTRANSFERASE (DRM) proteins; however, its roles remain poorly understood. Here we characterize the roles of non-CG methylation in Arabidopsis thaliana. We show that a poorly characterized methyltransferase, CMT2, is a functional methyltransferase in vitro and in vivo. CMT2 preferentially binds histone H3 Lys9 (H3K9) dimethylation and methylates non-CG cytosines that are regulated by H3K9 methylation. We revealed the contributions and redundancies between each non-CG methyltransferase in DNA methylation patterning and in regulating transcription. We also demonstrate extensive dependencies of small-RNA accumulation and H3K9 methylation patterning on non-CG methylation, suggesting self-reinforcing mechanisms between these epigenetic factors. The results suggest that non-CG methylation patterns are critical in shaping the landscapes of histone modification and small noncoding RNA.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Accession codes
References
Law, J.A. & Jacobsen, S.E. Establishing, maintaining and modifying DNA methylation patterns in plants and animals. Nat. Rev. Genet. 11, 204–220 (2010).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Xie, W. et al. Base-resolution analyses of sequence and parent-of-origin dependent DNA methylation in the mouse genome. Cell 148, 816–831 (2012).
Laurent, L. et al. Dynamic changes in the human methylome during differentiation. Genome Res. 20, 320–331 (2010).
Meissner, A. et al. Reduced representation bisulfite sequencing for comparative high-resolution DNA methylation analysis. Nucleic Acids Res. 33, 5868–5877 (2005).
Ramsahoye, B.H. et al. Non-CpG methylation is prevalent in embryonic stem cells and may be mediated by DNA methyltransferase 3a. Proc. Natl. Acad. Sci. USA 97, 5237–5242 (2000).
Varley, K.E. et al. Dynamic DNA methylation across diverse human cell lines and tissues. Genome Res. 23, 555–567 (2013).
Stroud, H., Greenberg, M.V., Feng, S., Bernatavichute, Y.V. & Jacobsen, S.E. Comprehensive analysis of silencing mutants reveals complex regulation of the Arabidopsis methylome. Cell 152, 352–364 (2013).
Zemach, A. et al. The Arabidopsis nucleosome remodeler DDM1 allows DNA methyltransferases to access H1-containing heterochromatin. Cell 153, 193–205 (2013).
Du, J. et al. Dual binding of chromomethylase domains to H3K9me2-containing nucleosomes directs DNA methylation in plants. Cell 151, 167–180 (2012).
Ebbs, M.L. & Bender, J. Locus-specific control of DNA methylation by the Arabidopsis SUVH5 histone methyltransferase. Plant Cell 18, 1166–1176 (2006).
Jackson, J.P., Lindroth, A.M., Cao, X. & Jacobsen, S.E. Control of CpNpG DNA methylation by the KRYPTONITE histone H3 methyltransferase. Nature 416, 556–560 (2002).
Henikoff, S. & Comai, L. A DNA methyltransferase homolog with a chromodomain exists in multiple polymorphic forms in Arabidopsis. Genetics 149, 307–318 (1998).
Law, J.A. et al. Polymerase IV occupancy at RNA-directed DNA methylation sites requires SHH1. Nature 498, 385–389 (2013).
Zhang, H. et al. DTF1 is a core component of RNA-directed DNA methylation and may assist in the recruitment of Pol IV. Proc. Natl. Acad. Sci. USA 110, 8290–8295 (2013).
Johnson, L. et al. Mass spectrometry analysis of Arabidopsis histone H3 reveals distinct combinations of post-translational modifications. Nucleic Acids Res. 32, 6511–6518 (2004).
Roudier, F. et al. Integrative epigenomic mapping defines four main chromatin states in Arabidopsis. EMBO J. 30, 1928–1938 (2011).
Jullien, P.E., Susaki, D., Yelagandula, R., Higashiyama, T. & Berger, F. DNA methylation dynamics during sexual reproduction in Arabidopsis thaliana. Curr. Biol. 22, 1825–1830 (2012).
Henderson, I.R. & Jacobsen, S.E. Tandem repeats upstream of the Arabidopsis endogene SDC recruit non-CG DNA methylation and initiate siRNA spreading. Genes Dev. 22, 1597–1606 (2008).
Herr, A.J., Jensen, M.B., Dalmay, T. & Baulcombe, D.C. RNA polymerase IV directs silencing of endogenous DNA. Science 308, 118–120 (2005).
Pontier, D. et al. Reinforcement of silencing at transposons and highly repeated sequences requires the concerted action of two distinct RNA polymerases IV in Arabidopsis. Genes Dev. 19, 2030–2040 (2005).
Zilberman, D., Cao, X. & Jacobsen, S.E. ARGONAUTE4 control of locus-specific siRNA accumulation and DNA and histone methylation. Science 299, 716–719 (2003).
Kasschau, K.D. et al. Genome-wide profiling and analysis of Arabidopsis siRNAs. PLoS Biol. 5, e57 (2007).
Mosher, R.A., Schwach, F., Studholme, D. & Baulcombe, D.C. PolIVb influences RNA-directed DNA methylation independently of its role in siRNA biogenesis. Proc. Natl. Acad. Sci. USA 105, 3145–3150 (2008).
Zhang, X., Henderson, I.R., Lu, C., Green, P.J. & Jacobsen, S.E. Role of RNA polymerase IV in plant small RNA metabolism. Proc. Natl. Acad. Sci. USA 104, 4536–4541 (2007).
Wierzbicki, A.T. et al. Spatial and functional relationships among Pol V-associated loci, Pol IV-dependent siRNAs, and cytosine methylation in the Arabidopsis epigenome. Genes Dev. 26, 1825–1836 (2012).
McCue, A.D., Nuthikattu, S. & Slotkin, R.K. Genome-wide identification of genes regulated in trans by transposable element small interfering RNAs. RNA Biol. 10, 1379–1395 (2013).
Zhong, X. et al. DDR complex facilitates global association of RNA polymerase V to promoters and evolutionarily young transposons. Nat. Struct. Mol. Biol. 19, 870–875 (2012).
Johnson, L.M. et al. The SRA methyl-cytosine-binding domain links DNA and histone methylation. Curr. Biol. 17, 379–384 (2007).
Inagaki, S. et al. Autocatalytic differentiation of epigenetic modifications within the Arabidopsis genome. EMBO J. 29, 3496–3506 (2010).
Mathieu, O., Probst, A.V. & Paszkowski, J. Distinct regulation of histone H3 methylation at lysines 27 and 9 by CpG methylation in Arabidopsis. EMBO J. 24, 2783–2791 (2005).
Soppe, W.J. et al. DNA methylation controls histone H3 lysine 9 methylation and heterochromatin assembly in Arabidopsis. EMBO J. 21, 6549–6559 (2002).
Johnson, L., Cao, X. & Jacobsen, S. Interplay between two epigenetic marks. DNA methylation and histone H3 lysine 9 methylation. Curr. Biol. 12, 1360–1367 (2002).
Deleris, A. et al. Loss of the DNA methyltransferase MET1 induces H3K9 hypermethylation at PcG target genes and redistribution of H3K27 trimethylation to transposons in Arabidopsis thaliana. PLoS Genet. 8, e1003062 (2012).
Wierzbicki, A.T., Ream, T.S., Haag, J.R. & Pikaard, C.S. RNA polymerase V transcription guides ARGONAUTE4 to chromatin. Nat. Genet. 41, 630–634 (2009).
Chan, S.W. et al. RNAi, DRD1, and histone methylation actively target developmentally important non-CG DNA methylation in Arabidopsis. PLoS Genet. 2, e83 (2006).
Feng, S., Rubbi, L., Jacobsen, S.E. & Pellegrini, M. Determining DNA methylation profiles using sequencing. Methods Mol. Biol. 733, 223–238 (2011).
Stroud, H. et al. DNA methyltransferases are required to induce heterochromatic re-replication in Arabidopsis. PLoS Genet. 8, e1002808 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Lu, C., Meyers, B.C. & Green, P.J. Construction of small RNA cDNA libraries for deep sequencing. Methods 43, 110–117 (2007).
Moissiard, G. et al. MORC family ATPases required for heterochromatin condensation and gene silencing. Science 336, 1448–1451 (2012).
Acknowledgements
We thank M. Akhavan for Illumina sequencing. Sequencing was performed at the University of California, Los Angeles (UCLA) Broad Stem Cell Research Center BioSequencing Core Facility. We thank W. Yang and M. Pellegrini for help with the UCSC browser. We thank L. Tao and D. Chen for technical help with experiments. We thank F. Berger for helpful comments. This work was supported by the Abby Rockefeller Mauzé Trust, the Maloris and Starr foundations (D.J.P.) and US National Institutes of Health grant GM60398 and National Science Foundation grant 0701745 (S.E.J.). H.S. was supported by a UCLA Dissertation Year Fellowship. X.Z. is supported by a Ruth L. Kirschstein National Research Service Award (F32GM096483-01). S.F. is supported as a Special Fellow of the Leukemia & Lymphoma Society. S.E.J. is supported as an investigator of the Howard Hughes Medical Institute.
Author information
Authors and Affiliations
Contributions
H.S., T.D., J.D., X.Z., S.F. and L.J. performed the experiments. S.E.J. and D.J.P. oversaw the study. H.S. designed the study, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 CMT2 does not methylate CG sites in vitro, and its activity is distinct from that of CMT3.
(a) RT-PCR on CMT2 and ACTIN transcripts in wild type and cmt2-7 mutants. A no RT control for the CMT2 amplification is also shown. Cartoon of T-DNA insertion site in the CMT2 gene (At4g19020) is also shown. Thick lines, exons; thin lines, introns. (b) CMT3 in vitro methyltransferase activity assay performed in parallel to the CMT2 assay. Error bars represent SD for two technical replicates. (c) CMT2 in vitro methyltransferase activity on CG sites. Error bars represent SD for two technical replicates
Supplementary Figure 2 CMT2 preferentially binds H3K9 di- and trimethylated peptides in vitro.
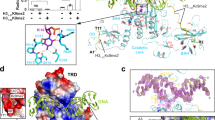
(a) Sequence comparison between CMT2, CMT3, and ZMET2 (the maize CMT3 homolog) by ClustalW (http://www.genome.jp/tools/clustalw/). BAH and CHROMO domains are shaded. (b) Single modification peptide reactivity from the peptide array generated by the manufacturer software (Active Motif).
Supplementary Figure 3 Contributions of non-CG methyltransferases in DNA methylation patterning.
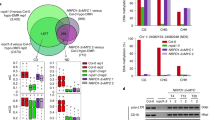
(a) CG, CHG, and CHH methylation levels across chromosomes. (b) Genome browser views of CHG and CHH methylation in chromosome 1. Blue bars, TEs; Yellow bars, genes. (c) Overlap between drm1 drm2 cmt2 and drm1 drm2 cmt2 cmt3 CHH DMRs. (d) Overlap between drm1 drm2 and cmt2 CHH DMRs.
Supplementary Figure 4 Roles of non-CG methyltransferases in gene silencing.
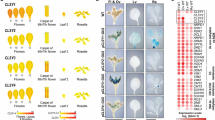
(a) Genome browser views of RNA-seq data. Blue bars, TEs; Yellow bars, genes. (b) Types of TEs upregulated in drm1 drm2 cmt2 cmt3 mutants. (c) Wild-type CHG and CHH methylation levels over genes. (d) Gene ontology analysis of genes upregulated in drm1 drm2 cmt2 cmt3 mutants using DAVID (http://david.abcc.ncifcrf.gov/). (e) Expression levels of SDC. Error bars represent SD from two biological replicates. (f) Photo of plants of indicated genotypes.
Supplementary Figure 5 24-nt siRNAs produced at CMT2 sites do not guide DNA methylation in cis.
(a) Average distribution of Pol IV over CMT2 target sites. ChIP-seq data on epitope-tagged CMT31, Pol IV2, Pol V3 relative to the respective controls, as well as Pol II relative to input genomic DNA was plotted over CMT2 target sites. (b) Heatmaps of CHH methylation levels4 within cmt2 CHH DMRs.
Supplementary Figure 6 Non-CG methylation shapes the histone-modification landscape.
(a) Genome browser views of DNA methylation, expression levels, and H3K9me2 in wild type, drm1 drm2 cmt2 cmt3, and kyp suvh5 suvh6 mutants in chromosome 1. Yellow bars, genes. (b) Average distribution of H3K9me2 over previously defined met1 CHG hypomethylation DMRs (black) and CHH hypomethylation DMRs (blue). Wild-type data is plotted in solid lines, and met1 data is plotted in faded lines. (c) Distribution of H3K9me2 over chromosomes in wild type and met1 mutants. (d) Average distribution of wild-type H3K23ac levels over genes categorized by indicated wild-type expression levels. (e) Distribution of H3K23ac relative to H3 across chromosomes. (f) Genome browser views of DNA methylation, expression levels, H3K23ac, and H3K9me2 in wild type, drm1 drm2 cmt2 cmt3, and kyp suvh5 suvh6 mutants in chromosome 1. Yellow bars, genes. (g) Chromocenter decondensation assay in indicated genotypes. >100 nuclei were assayed per genotype.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Tables 1–3 (PDF 8888 kb)
Rights and permissions
About this article
Cite this article
Stroud, H., Do, T., Du, J. et al. Non-CG methylation patterns shape the epigenetic landscape in Arabidopsis. Nat Struct Mol Biol 21, 64–72 (2014). https://doi.org/10.1038/nsmb.2735
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2735
This article is cited by
-
Contribution of RdDM to the ecotype-specific differential methylation on conserved as well as highly variable regions between Arabidopsis ecotypes
BMC Genomics (2023)
-
DDM1-mediated gene body DNA methylation is associated with inducible activation of defense-related genes in Arabidopsis
Genome Biology (2023)
-
Arabidopsis histone H3 lysine 9 methyltransferases KYP/SUVH5/6 are involved in leaf development by interacting with AS1-AS2 to repress KNAT1 and KNAT2
Communications Biology (2023)
-
Advances in DNA methylation and demethylation in medicinal plants: a review
Molecular Biology Reports (2023)
-
Epigenetic modification associated with climate regulates betulin biosynthesis in birch
Journal of Forestry Research (2023)