Abstract
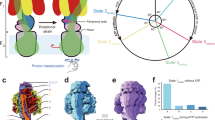
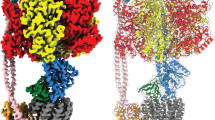
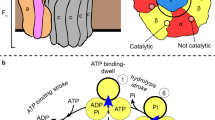
ATP synthase is a membrane-bound rotary motor enzyme that is critical for cellular energy metabolism in all kingdoms of life. Despite conservation of its basic structure and function, autoinhibition by one of its rotary stalk subunits occurs in bacteria and chloroplasts but not in mitochondria. The crystal structure of the ATP synthase catalytic complex (F1) from Escherichia coli described here reveals the structural basis for this inhibition. The C-terminal domain of subunit ɛ adopts a heretofore unknown, highly extended conformation that inserts deeply into the central cavity of the enzyme and engages both rotor and stator subunits in extensive contacts that are incompatible with functional rotation. As a result, the three catalytic subunits are stabilized in a set of conformations and rotational positions distinct from previous F1 structures.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Boyer, P.D. The ATP synthase–-a splendid molecular machine. Annu. Rev. Biochem. 66, 717–749 (1997).
Duncan, T.M. The ATP synthase: parts and properties of a rotary motor. in The Enzymes, vol. XXIII: Energy Coupling and Molecular Motors Vol. 23 (eds. Hackney, D.D. & Tamanoi, F.) 203–275 (Elsevie, New York, 2004).
Junge, W., Sielaff, H. & Engelbrecht, S. Torque generation and elastic power transmission in the rotary FoF1-ATPase. Nature 459, 364–370 (2009).
Abrahams, J.P., Leslie, A.G., Lutter, R. & Walker, J.E. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994).
Bowler, M.W., Montgomery, M.G., Leslie, A.G. & Walker, J.E. Ground state structure of F1-ATPase from bovine heart mitochondria at 1.9 Å resolution. J. Biol. Chem. 282, 14238–14242 (2007).
Gledhill, J.R., Montgomery, M.G., Leslie, A.G. & Walker, J.E. How the regulatory protein, IF1, inhibits F1-ATPase from bovine mitochondria. Proc. Natl. Acad. Sci. USA 104, 15671–15676 (2007).
Menz, R.I., Walker, J.E. & Leslie, A.G. Structure of bovine mitochondrial F1-ATPase with nucleotide bound to all three catalytic sites: implications for the mechanism of rotary catalysis. Cell 106, 331–341 (2001).
Hausrath, A.C., Gruber, G., Matthews, B.W. & Capaldi, R.A. Structural features of the γ subunit of the Escherichia coli F1 ATPase revealed by a 4.4-Å resolution map obtained by x-ray crystallography. Proc. Natl. Acad. Sci. USA 96, 13697–13702 (1999).
Stocker, A., Keis, S., Vonck, J., Cook, G.M. & Dimroth, P. The structural basis for unidirectional rotation of thermoalkaliphilic F1-ATPase. Structure 15, 904–914 (2007).
Shirakihara, Y. et al. The crystal structure of the nucleotide-free α3β3 subcomplex of F1-ATPase from the thermophilic Bacillus PS3 is a symmetric trimer. Structure 5, 825–836 (1997).
Feniouk, B.A., Suzuki, T. & Yoshida, M. The role of subunit ɛ in the catalysis and regulation of FoF1-ATP synthase. Biochim. Biophys. Acta 1757, 326–338 (2006).
Richter, M.L. Gamma-epsilon interactions regulate the chloroplast ATP synthase. Photosynth. Res. 79, 319–329 (2004).
Gibbons, C., Montgomery, M.G., Leslie, A.G. & Walker, J.E. The structure of the central stalk in bovine F1-ATPase at 2.4 Å resolution. Nat. Struct. Biol. 7, 1055–1061 (2000).
Campanella, M., Parker, N., Tan, C.H., Hall, A.M. & Duchen, M.R. IF1: setting the pace of the F1Fo-ATP synthase. Trends Biochem. Sci. 34, 343–350 (2009).
Andries, K. et al. A diarylquinoline drug active on the ATP synthase of Mycobacterium tuberculosis. Science 307, 223–227 (2005).
Uhlin, U., Cox, G.B. & Guss, J.M. Crystal structure of the ɛ subunit of the proton-translocating ATP synthase from Escherichia coli. Structure 5, 1219–1230 (1997).
Wilkens, S. & Capaldi, R.A. Solution structure of the ɛ subunit of the F1-ATPase from Escherichia coli and interactions of this subunit with β subunits in the complex. J. Biol. Chem. 273, 26645–26651 (1998).
Yagi, H. et al. Structures of the thermophilic F1-ATPase ɛ subunit suggesting ATP-regulated arm motion of its C-terminal domain in F1 . Proc. Natl. Acad. Sci. USA 104, 11233–11238 (2007).
Schulenberg, B. & Capaldi, R.A. The ɛ subunit of the F1Fo complex of Escherichia coli. Cross-linking studies show the same structure in situ as when isolated. J. Biol. Chem. 274, 28351–28355 (1999).
Dallmann, H.G., Flynn, T.G. & Dunn, S.D. Determination of the 1-ethyl-3-[(3-dimethylamino)propyl]-carbodiimide-induced cross-link between the β and ɛ subunits of Escherichia coli F1-ATPase. J. Biol. Chem. 267, 18953–18960 (1992).
Rodgers, A.J. & Wilce, M.C. Structure of the γ-ɛ complex of ATP synthase. Nat. Struct. Biol. 7, 1051–1054 (2000).
Saita, E.I. et al. Activation and stiffness of the inhibited states of F1-ATPase probed by single-molecule manipulation. J. Biol. Chem. 285, 11441–11447 (2010).
Iino, R., Hasegawa, R., Tabata, K.V. & Noji, H. Mechanism of inhibition by C-terminal α-helices of the ɛ subunit of Escherichia coli FoF1-ATP synthase. J. Biol. Chem. 284, 17457–17464 (2009).
Kabaleeswaran, V. et al. Asymmetric structure of the yeast F1 ATPase in the absence of bound nucleotides. J. Biol. Chem. 284, 10546–10551 (2009).
Yasuda, R., Noji, H., Yoshida, M., Kinosita, K. Jr. & Itoh, H. Resolution of distinct rotational substeps by submillisecond kinetic analysis of F1-ATPase. Nature 410, 898–904 (2001).
Nishizaka, T. et al. Chemomechanical coupling in F1-ATPase revealed by simultaneous observation of nucleotide kinetics and rotation. Nat. Struct. Mol. Biol. 11, 142–148 (2004).
Kabaleeswaran, V., Puri, N., Walker, J.E., Leslie, A.G. & Mueller, D.M. Novel features of the rotary catalytic mechanism revealed in the structure of yeast F1 ATPase. EMBO J. 25, 5433–5442 (2006).
Pu, J. & Karplus, M. How subunit coupling produces the gamma-subunit rotary motion in F1-ATPase. Proc. Natl. Acad. Sci. USA 105, 1192–1197 (2008).
Sielaff, H. et al. Domain compliance and elastic power transmission in rotary FoF1-ATPase. Proc. Natl. Acad. Sci. USA 105, 17760–17765 (2008).
Wilkens, S. & Capaldi, R.A. Asymmetry and structural changes in ECF1 examined by cryoelectronmicroscopy. Biol. Chem. Hoppe Seyler 375, 43–51 (1994).
Zimmermann, B., Diez, M., Zarrabi, N., Graber, P. & Börsch, M. Movements of the ɛ-subunit during catalysis and activation in single membrane-bound H+-ATP synthase. EMBO J. 24, 2053–2063 (2005).
Cui, Q., Li, G., Ma, J. & Karplus, M. A normal mode analysis of structural plasticity in the biomolecular motor F1-ATPase. J. Mol. Biol. 340, 345–372 (2004).
Müller, M. et al. Rotary F1-ATPase. Is the C-terminus of subunit γ fixed or mobile? Eur. J. Biochem. 271, 3914–3922 (2004).
Keis, S., Stocker, A., Dimroth, P. & Cook, G.M. Inhibition of ATP hydrolysis by thermoalkaliphilic F1Fo-ATP synthase is controlled by the C terminus of the ɛ subunit. J. Bacteriol. 188, 3796–3804 (2006).
Slonczewski, J.L., Fujisawa, M., Dopson, M. & Krulwich, T.A. Cytoplasmic pH measurement and homeostasis in bacteria and archaea. Adv. Microb. Physiol. 55, 1–79 (2009).
Feniouk, B.A. & Junge, W. Regulation of the FoF1-ATP synthase: the conformation of subunit ɛ might be determined by directionality of subunit γ rotation. FEBS Lett. 579, 5114–5118 (2005).
Foster, J.W. Escherichia coli acid resistance: tales of an amateur acidophile. Nat. Rev. Microbiol. 2, 898–907 (2004).
Mendel-Hartvig, J. & Capaldi, R.A. Nucleotide-dependent and dicyclohexylcarbodiimide-sensitive conformational changes in the ɛ subunit of Escherichia coli ATP synthase. Biochemistry 30, 10987–10991 (1991).
Lippe, G., Sorgato, M.C. & Harris, D.A. The binding and release of the inhibitor protein are governed independently by ATP and membrane potential in ox-heart submitochondrial vesicles. Biochim. Biophys. Acta 933, 12–21 (1988).
Fischer, S., Graber, P. & Turina, P. The activity of the ATP synthase from Escherichia coli is regulated by the transmembrane proton motive force. J. Biol. Chem. 275, 30157–30162 (2000).
Feniouk, B.A., Suzuki, T. & Yoshida, M. Regulatory interplay between proton motive force, ADP, phosphate, and subunit ɛ in bacterial ATP synthase. J. Biol. Chem. 282, 764–772 (2007).
Xiong, H., Zhang, D. & Vik, S.B. Subunit ɛ of the Escherichia coli ATP synthase: novel insights into structure and function by analysis of thirteen mutant forms. Biochemistry 37, 16423–16429 (1998).
Aggeler, R. & Capaldi, R.A. Nucleotide-dependent movement of the ɛ subunit between α and β subunits in the Escherichia coli F1Fo-type ATPase. J. Biol. Chem. 271, 13888–13891 (1996).
Ferrándiz, M.J. & de la Campa, A.G. The membrane-associated FoF1 ATPase is essential for the viability of Streptococcus pneumoniae. FEMS Microbiol. Lett. 212, 133–138 (2002).
Rao, S.P.S., Alonso, S., Rand, L., Dick, T. & Pethe, K. The protonmotive force is required for maintaining ATP homeostasis and viability of hypoxic, nonreplicating Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 105, 11945–11950 (2008).
Jones, S.A. et al. Respiration of Escherichia coli in the mouse intestine. Infect. Immun. 75, 4891–4899 (2007).
Dautant, A., Velours, J. & Giraud, M.-F. Crystal structure of the Mg•ADP-inhibited state of the yeast F1c10-ATP synthase. J. Biol. Chem. 285, 29502–29510 (2010).
Wise, J.G. Site-directed mutagenesis of the conserved β subunit tyrosine 331 of Escherichia coli ATP synthase yields catalytically active enzymes. J. Biol. Chem. 265, 10403–10409 (1990).
Lee, R.S., Pagan, J., Wilke Mounts, S. & Senior, A.E. Characterization of Escherichia coli ATP synthase β-subunit mutations using a chromosomal deletion strain. Biochemistry 30, 6842–6847 (1991).
Schaefer, E.M., Hartz, D., Gold, L. & Simoni, R.D. Ribosome-binding sites and RNA-processing sites in the transcript of the Escherichia coli unc operon. J. Bacteriol. 171, 3901–3908 (1989).
Hendrickson, W.A., Horton, J.R. & LeMaster, D.M. Selenomethionyl proteins produced for analysis by multiwavelength anomalous diffraction (MAD): a vehicle for direct determination of three-dimensional structure. EMBO J. 9, 1665–1672 (1990).
Duncan, T.M., Bulygin, V.V., Zhou, Y., Hutcheon, M.L. & Cross, R.L. Rotation of subunits during catalysis by Escherichia coli F1-ATPase. Proc. Natl. Acad. Sci. USA 92, 10964–10968 (1995).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. Solving structures of protein complexes by molecular replacement with Phaser. Acta Crystallogr. D Biol. Crystallogr. 63, 32–41 (2007).
Bowler, M.W., Montgomery, M.G., Leslie, A.G. & Walker, J.E. How azide inhibits ATP hydrolysis by the F-ATPases. Proc. Natl. Acad. Sci. USA 103, 8646–8649 (2006).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Duncan, T.M., Zhou, Y., Bulygin, V.V., Hutcheon, M.L. & Cross, R.L. Probing interactions of the Escherichia coli FoF1 ATP synthase β and γ subunits with disulphide cross-links. Biochem. Soc. Trans. 23, 736–741 (1995).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Acknowledgements
We dedicate this work to the memory of Vladimir Bulygin, Ph.D. (1964–2009), who was instrumental in early stages of the project. We thank M. Hutcheon for skillful protein purification. Financial support was provided by the US National Institutes of Health (R01GM083088). We thank the staff at NSLS beamlines X6A, X25 (Brookhaven National Laboratory, Upton, New York, USA) and at macCHESS (Cornell University, Ithaca, New York, USA) for beam time and assistance in data collection.
Author information
Authors and Affiliations
Contributions
T.M.D. developed a purification protocol to obtain homogeneous EF1-δ. T.M.D. and G.C. crystallized EF1-δ. G.C. collected X-ray data and determined the crystal structure. T.M.D. wrote the manuscript with the help of G.C.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Supplementary Methods (PDF 2356 kb)
Rights and permissions
About this article
Cite this article
Cingolani, G., Duncan, T. Structure of the ATP synthase catalytic complex (F1) from Escherichia coli in an autoinhibited conformation. Nat Struct Mol Biol 18, 701–707 (2011). https://doi.org/10.1038/nsmb.2058
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2058
This article is cited by
-
Molecular mechanism on forcible ejection of ATPase inhibitory factor 1 from mitochondrial ATP synthase
Nature Communications (2023)
-
Structural basis of redox modulation on chloroplast ATP synthase
Communications Biology (2020)
-
Cryo-EM structures provide insight into how E. coli F1Fo ATP synthase accommodates symmetry mismatch
Nature Communications (2020)
-
Distance measurements in the F0F1-ATP synthase from E. coli using smFRET and PELDOR spectroscopy
European Biophysics Journal (2020)
-
ATP synthesis at physiological nucleotide concentrations
Scientific Reports (2019)