Abstract
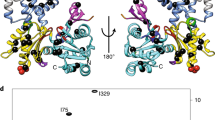
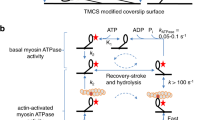
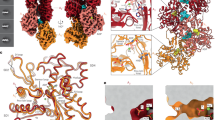
Structural insights into the interaction of smooth muscle myosin with actin have been provided by computer-based fitting of crystal structures into three-dimensional reconstructions obtained by electron cryomicroscopy, and by mapping of structural and dynamic changes in the actomyosin complex. The actomyosin structures determined in the presence and absence of MgADP differ significantly from each other, and from all crystallographic structures of unbound myosin. Coupled to a complex movement (∼34 Å) of the light chain binding domain upon MgADP release, we observed a ∼9° rotation of the myosin motor domain relative to the actin filament, and a closure of the cleft that divides the actin binding region of the myosin head. Cleft closure is achieved by a movement of the upper 50 kDa region, while parts of the lower 50 kDa region are stabilized through strong interactions with actin. This model supports a mechanism in which binding of MgATP at the active site opens the cleft and disrupts the interface, thereby releasing myosin from actin.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Geeves, M.A. The dynamics of actin and myosin association and the crossbridge model of muscle contraction. Biochem. J. 274, 1–14 (1991).
Block, S.M. Fifty ways to love your lever: myosin motors. Cell 87, 151–157 (1996).
Goldman, Y.E. Wag the tail: structural dynamics of actomyosin. Cell 93, 1–4 (1998).
Dominguez, R., Freyzon, Y., Trybus, K.M. & Cohen, C. Crystal structure of a vertebrate smooth muscle myosin motor domain and its complex with the essential light chain: visualization of the pre-power stroke state. Cell 94, 559–571 (1998).
Fisher, A.J. et al. X-ray structures of the myosin motor domain of Dictyostelium discoideum complexed with MgADP.BeFx and MgADP.AlF4 −. Biochemistry 34, 8960–8972 (1995).
Gulick, A.M., Bauer, C.B., Thoden, J.B. & Rayment, I. X-ray structures of the MgADP, MgATPγS, and MgAMPPNP complexes of the Dictyostelium discoideum myosin motor domain. Biochemistry 36, 11619–11628 (1997).
Rayment, I. et al. Three-dimensional structure of myosin subfragment-1: a molecular motor. Science 261, 50–58 (1993).
Smith, C.A. & Rayment, I. X-ray structure of the magnesium(II).ADP.vanadate complex of the Dictyostelium discoideum myosin motor domain to 1.9 Å resolution. Biochemistry 35, 5404–5417 (1996).
Lorenz, M., Popp, D. & Holmes, K.C. Refinement of the F-actin model against X-ray fiber diffraction data by the use of a directed mutation algorithm. J. Mol. Biol. 234, 826–836 (1993).
Rayment, I. et al. Structure of the actin-myosin complex and its implications for muscle contraction. Science 261, 58–65 (1993).
Schröder, R.R. et al. Three-dimensional atomic model of F-actin decorated with Dictyostelium myosin S1. Nature 364, 171–174 (1993).
Whittaker, M. et al. A 35-Å movement of smooth muscle myosin on ADP release. Nature 378, 748–751 (1995).
Hanein, D., Matsudaira, P. & DeRosier, D.J. Evidence for a conformational change in actin induced by fimbrin (N375) binding. J. Cell Biol. 139, 387–396 (1997).
Hanein, D. & DeRosier, D.J. A new algorithm to align 3-D maps of helical structures. Ultramicroscopy 76, 233–238 (1999).
Rost, L.E., Hanein, D. & DeRosier, D.J. Reconstruction of symmetry deviations: a procedure to analyze partially decorated F-actin and other incomplete structures. Ultramicroscopy 72, 187–197 (1998).
Volkmann, N. & Hanein, D. Quantitative fitting of atomic models into observed densities derived by electron microscopy. J. Struct. Biol. 125, 176–184 (1999).
Houdusse, A., Kalabokis, V.N., Himmel, D., Szent-Györgyi, A.G. & Cohen, C. Atomic structure of scallop myosin subfragment S1 complexed with MgADP: a novel conformation of the myosin head. Cell 97, 459–470 (1999).
Cremo, C.R. & Geeves, M.A. Interaction of actin and ADP with the head domain of smooth muscle myosin: implications for strain-dependent ADP release in smooth muscle. Biochemistry 37, 1969–1978 (1998).
Baker, T.S. & Johnson, J.E. Low resolution meets high: towards a resolution continuum from cells to atoms. Curr. Opin. Struct. Biol. 6, 585–594 (1996).
Corrie, J.E.T. et al. Dynamic measurement of myosin light-chain-domain tilt and twist in muscle contraction. Nature 400, 425–430 (1999).
Lauzon, A.M. et al. A 7-amino-acid loop alters the kinetics of smooth muscle myosin in the laser trap. J. Muscle Res. 19, 825–837 (1998).
Sweeney, H.L. et al. Kinetic tuning of myosin via a flexible loop adjacent to the nucleotide binding pocket. J. Biol. Chem. 273, 6262–6270 (1998).
Kurzawa-Goertz, S.E., Perreault-Micale, C.L., Trybus, K.M., Szent-Györgyi, A.G. & Geeves, M.A. Loop I can modulate ADP affinity, ATPase activity, and motility of different scallop myosins. Transient kinetic analysis of S1 isoforms. Biochemistry 37, 7517–7525 (1998).
Applegate, D. & Reisler, E. Protease-sensitive regions in myosin subfragment 1. Proc. Natl. Acad. Sci. USA 80, 7109–7112 (1983).
Mehta, A.D., Finer, J.T. & Spudich, J.A. Detection of single-molecule interactions using correlated thermal diffusion. Proc. Natl. Acad. Sci. USA 94, 7927–7931 (1997).
Molloy, J.E., Burns, J.E., Kendrick-Jones, J., Tregear, R.T. & White, D.C. Movement and force produced by a single myosin head. Nature 378, 209–212 (1995).
Tyska, M.J. et al. Two heads of myosin are better than one for generating force and motion. Proc. Natl. Acad. Sci. USA 96, 4402–4407 (1999).
Gollub, J., Cremo, C.R. & Cooke, R. ADP release produces a rotation of the neck region of smooth myosin but not skeletal myosin. Nature Struct. Biol. 3, 796–802 (1996).
Dantzig, J.A., Barsotti, R.J., Manz, S., Sweeney, H.L. & Goldman, Y.E. The ADP release step of the smooth muscle cross-bridge cycle is not directly associated with force generation. Biophys. J. 77, 386–397 (1999).
Milligan, R.A. Protein-protein interactions in the rigor actomyosin complex. Proc. Natl. Acad. Sci. USA 93, 21–26 (1996).
Volkmann, N. & Hanein, D. Actomyosin: law and order in motility. Curr. Opin. Cell Biol. 12, 26–34 (2000).
Yengo, C.M., Chrin, L., Rovner, A.S. & Berger, C.L. Intrinsic tryptophan fluorescence identifies specific conformational changes at the actomyosin interface upon actin binding and ADP-release. Biochemistry 38, 14515–14523 (1999).
Fisher, A.J. et al. Structural studies of myosin:nucleotide complexes: a revised model for the molecular basis of muscle contraction. Biophys. J. 68, 19S–26S (1995).
Holmes, K.C. Muscle proteins - their actions and interactions. Curr. Opin. Struct. Biol. 6, 781–789 (1996).
Pardee, J.D. & Spudich, J.A. Purification of muscle actin. Methods Enzymol. 85B, 164–181 (1982).
Trybus, K.M. Regulation of expressed truncated smooth muscle myosins. Role of the essential light chain and tail length. J. Biol. Chem. 269, 20819–20822 (1994).
Dubochet, J. et al. Cryo-electron microscopy of vitrified specimens. Q. Rev. Biophys. 21, 129–228 (1988).
Owen, C.H., Morgan, D.G. & DeRosier, D.J. Image analysis of helical objects: the Brandeis helical package. J. Struct. Biol. 116, 167–175 (1996).
DeRosier, D.J. & Moore, P.B. Reconstruction of three-dimensional images from electron micrographs of structures with helical symmetry. J. Mol. Biol. 52, 355–369 (1970).
Trachtenberg, S. & DeRosier, D.J. Three-dimensional structure of the frozen-hydrated flagellar filament: the left-handed filament of Salmonella typhimurium. J. Mol. Biol. 195, 581–601 (1987).
Hanein, D. et al. An atomic model of fimbrin binding to F-actin and its implications for filament crosslinking and regulation. Nature Struct. Biol. 5, 787–792 (1998).
Priestle, J.P. Stereochemical dictionaries for protein structure refinement and model building. Structure 2, 911–913 (1994).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D 53, 240–255 (1997).
von Mises, R. Mathematical theory of probability and statistics (Academic Press, New York; 1964).
Kraulis, P. MOLSCRIPT: a program to produce both detailed and schematic plots of protein structures. J. Appl. Crystallogr. 24, 946–950 (1991).
Esnouf, R. An extensively modified version of MolScript that includes greatly enhanced coloring capabilities. J. Mol. Graph. 15, 132–134 (1997).
Bacon, D. & Anderson, W. A fast algorithm for rendering space-filling molecule pictures. J. Mol. Graph. 6, 219–220 (1988).
Nicholls, A., Sharp, K.A. & Honig, B. Protein folding and association: insights from the interfacial and thermodynamic properties of hydrocarbons. Proteins 11, 281–296 (1991).
Acknowledgements
This work was supported by National Institutes of Health grants to D.J.D., S.L. and K.M.T. and a National Science Foundation grant to S.L. for the support of G.O. The authors acknowledge funds from the W.M. Keck Foundation.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Volkmann, N., Hanein, D., Ouyang, G. et al. Evidence for cleft closure in actomyosin upon ADP release. Nat Struct Mol Biol 7, 1147–1155 (2000). https://doi.org/10.1038/82008
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/82008
This article is cited by
-
The actomyosin interface contains an evolutionary conserved core and an ancillary interface involved in specificity
Nature Communications (2021)
-
Three-dimensional reconstructions of Arp2/3 complex with bound nucleation promoting factors
The EMBO Journal (2012)
-
Loop 1 dynamics in smooth muscle myosin: isoform specific differences modulate ADP release
Journal of Muscle Research and Cell Motility (2011)
-
The structure of the C-terminal actin-binding domain of talin
The EMBO Journal (2008)