Key Points
-
Ribosomal translocation and back-translocation refer to the forward and backward movements of the tRNA2–mRNA complex on the ribosome during elongation. Elongation factor G (EF-G) catalyses translocation and EF4 catalyses the reverse reaction.
-
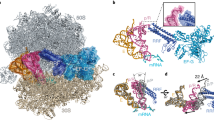
EF-G and EF4 have very similar structures and differ by each having only one factor-specific domain; domain IV of EF-G and the carboxy-terminal domain of EF4. The domains that are common to both factors are responsible for binding to the ribosome and GTPase activity, the specific domains are responsible for mediating translocation and back-translocation.
-
Evidence so far suggests that EF4 mobilizes stalled ribosomes either before or after translocation, thus accelerating protein synthesis.
-
Translocation and back-translocation are controlled by a constriction of the the neck of the small subunit to approximately 14 Å, which is termed the A790 gate. This gap is too narrow for movement of the anticodon stem of the tRNA as it has a diameter of 20 Å.
-
The A790 gate is closed in all functional states except during an intermediate translocation state, in which the gate is open to a width of more than 23 Å. EF-G, and probably also EF4, cause a rotation of the 30S head relative to the 30S body (termed swivelling), which opens the A790 gate and enables the subsequent translocation and back-translocation of the tRNA2–mRNA complex.
-
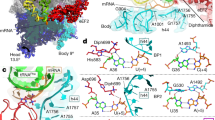
Domain IV of EF-G flips into the A-site as soon this site becomes free during translocation. This 'doorstop' function of domain IV prevents tRNA back-translocation, as long as the A790 gate is open. On the mRNA side, this function is supported by the 16S rRNA bases C1397 and A1505 that intercalate with bases of the mRNA, thereby blocking its movement.
-
The available structures suggest that the orientation of His92 of EF-G (Escherichia coli nomenclature) is a diagnostic feature of the activity state of the GTPase centre. An active state is observed exclusively in the translocation intermediate state, in which His92 is located 3 Å away from the bound GTP and points towards the γ-phosphate of GTP; whereas in the inactive state (after translocation), His92 has moved further away (at a distance of 9 Å) and turns away from the γ-phosphate of GTP.
Abstract
Ribosomes translate the codon sequence of an mRNA into the amino acid sequence of the corresponding protein. One of the most crucial events is the translocation reaction, which involves movement of both the mRNA and the attached tRNAs by one codon length and is catalysed by the GTPase elongation factor G (EF-G). Interestingly, recent studies have identified a structurally related GTPase, EF4, that catalyses movement of the tRNA2–mRNA complex in the opposite direction when the ribosome stalls, which is known as back-translocation. In this Review, we describe recent insights into the mechanistic basis of both translocation and back-translocation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Kaltschmidt, E. & Wittmann, H. G. Ribosomal proteins XII. Number of proteins in small and large subunits of Escherichia coli as determined by two-dimensional gel electrophoresis. Proc. Natl Acad. Sci. USA 67, 1276–1282 (1970).
Ilag, L. L. et al. Heptameric (L12)6/L10 rather than canonical pentameric complexes are found by tandem MS of intact ribosomes from thermophilic bacteria. Proc. Natl Acad. Sci. USA 102, 8192–8197 (2005).
Diaconu, M. et al. Structural basis for the function of the ribosomal L7/12 stalk in factor binding and GTPase activation. Cell 121, 991–1004 (2005).
Maki, Y. et al. Three binding sites for stalk protein dimers are generally present in ribosomes from archaeal organism. J. Biol. Chem. 282, 32827–32833 (2007).
Naganuma, T. et al. Structural basis for translation factor recruitment to the eukaryotic/archaeal ribosomes. J. Biol. Chem. 285, 4747–4756 (2010).
Pettersson, I., Hardy, S. J. S. & Liljas, A. The ribosomal protein L8 is a complex of L7/L12 and L10. FEBS Lett. 6, 135–138 (1976).
Ogle, J. M. et al. Recognition of cognate transfer RNA by the 30S ribosomal subunit. Science 292, 897–902 (2001).
Demeshkina, N., Jenner, L., Westhof, E., Yusupov, M. & Yusupova, G. A new understanding of the decoding principle on the ribosome. Nature 484, 256–259 (2012).
Nissen, P., Hansen, J., Ban, N., Moore, P. B. & Steitz, T. A. The structural basis of ribosome activity in peptide bond synthesis. Science 289, 920–930 (2000).
Voorhees, R. M., Weixlbaumer, A., Loakes, D., Kelley, A. C. & Ramakrishnan, V. Insights into substrate stabilization from snapshots of the peptidyl transferase center of the intact 70S ribosome. Nature Struct. Mol. Biol. 16, 528–533 (2009).
Hiller, D. A., Singh, V., Zhong, M. & Strobel, S. A. A two-step chemical mechanism for ribosome-catalysed peptide bond formation. Nature 476, 236–239 (2011).
Pech, M. & Nierhaus, K. H. The thorny way to the mechanism of ribosomal peptide-bond formation. Chembiochem 13, 189–192 (2012).
Nomura, M. Assembly of bacterial ribosomes. Science 179, 864–873 (1973).
Nierhaus, K. H. The assembly of prokaryotic ribosomes. Biochimie 73, 739–755 (1991).
Takyar, S., Hickerson, R. P. & Noller, H. F. mRNA helicase activity of the ribosome. Cell 120, 49–58 (2005).
Wower, I., Wower, J. & Zimmermann, R. Ribosomal protein L27 participates in both 50S subunit assembly and the peptidyl transferase reaction. J. Biol. Chem. 273, 19847–19852 (1998).
Diedrich, G. et al. Ribosomal protein L2 is involved in the association of the ribosomal subunits, tRNA binding to A and P sites and peptidyl transfer. EMBO J. 19, 5241–5250 (2000).
Schmeing, T. & Ramakrishnan, V. What recent ribosome structures have revealed about the mechanism of translation. Nature 461, 1234–1242 (2009).
Robertson, J. M. & Wintermeyer, W. Mechanism of ribosomal translocation. tRNA binds transiently to an exit site before leaving the ribosome during translocation. J. Mol. Biol. 196, 525–540 (1987).
Uemura, S. et al. Real-time tRNA transit on single translating ribosomes at codon resolution. Nature 464, 1012–1017 (2010).
Chen, C. et al. Allosteric versus spontaneous exit-site (E-site) tRNA dissociation early in protein synthesis. Proc. Natl Acad. Sci. USA 108, 16980–16985 (2011).
Triana-Alonso, F. J., Chakraburtty, K. & Nierhaus, K. H. The elongation factor 3 unique in higher fungi and essential for protein biosynthesis is an E site factor. J. Biol. Chem. 270, 20473–20478 (1995).
Dinos, G., Kalpaxis, D. L., Wilson, D. N. & Nierhaus, K. H. Deacylated tRNA is released from the E site upon A site occupation but before GTP is hydrolyzed by EF-Tu. Nucleic Acids Res. 33, 5291–5296 (2005).
Qin, Y. et al. The highly conserved LepA is a ribosomal elongation factor that back-translocates the ribosome. Cell 127, 721–733 (2006). This paper reports the discovery that EF4 is a ribosomal back-translocase.
Liu, H., Pan, D., Pech, M. & Cooperman, B. S. Interrupted catalysis: the EF4 (LepA) effect on back-translocation. J. Mol. Biol. 396, 1043–1052 (2010).
Pech, M. et al. Elongation factor 4 (EF4/LepA) accelerates protein synthesis at increased Mg2+ concentrations. Proc. Natl Acad. Sci. USA 108, 3199–3203 (2011). An in vivo and in vitro study suggesting that EF4 rescues stalled ribosomes.
Liu, H. et al. The conserved protein EF4 (LepA) modulates the elongation cycle of protein synthesis. Proc. Natl Acad. Sci. USA 108, 16223–16228 (2011). This study suggests that the PRE-state is the preferential target for EF4.
Connell, S. R. et al. A new tRNA intermediate revealed on the ribosome during EF4-mediated back-translocation. Nature Struct. Mol. Biol. 15, 910–915 (2008). This study shows the cryo-EM structure of EF4 in action.
Datta, P. P., Sharma, M. R., Qi, L., Frank, J. & Agrawal, R. K. Interaction of the G′ domain of elongation factor G and the C-terminal domain of ribosomal protein L7/L12 during translocation as revealed by cryo-EM. Mol. Cell 20, 723–731 (2005).
Yokoyama, T. et al. Structural insights into initial and intermediate steps of the ribosome-recycling process. EMBO J. 31, 1836–1846 (2012).
Gao, Y. G. et al. The structure of the ribosome with elongation factor G trapped in the posttranslocational state. Science 326, 694–699 (2009). This paper shows the crystal structure of an EF-G–70S complex in the POST-state.
Feng, S., Chen, Y. & Gao, Y. G. Crystal structure of 70S ribosome with both cognate tRNAs in the E and P sites representing an authentic elongation complex. PLoS ONE 8, e58829 (2013).
Wang, L. et al. A conserved proline switch on the ribosome facilitates the recruitment and binding of trGTPases. Nature Struct. Mol. Biol. 19, 403–410 (2012).
Zhang, D. et al. Common chaperone activity in the G-domain of trGTPase protects L11-L12 interaction on the ribosome. Nucleic Acids Res. 40, 10851–10865 (2012).
Ratje, A. H. et al. Head swivel on the ribosome facilitates translocation by means of intra-subunit tRNA hybrid sites. Nature 468, 713–716 (2010). This paper reports the first visualization of distinct structural intermediates during translocation.
Connell, S. R. et al. Structural basis for interaction of the ribosome with the switch regions of GTP-bound elongation factors. Mol. Cell 25, 751–764 (2007).
Khade, P. K., Shi, X. & Joseph, S. Steric complementarity in the decoding center Is Important for tRNA selection by the ribosome. J. Mol. Biol. (2013).
Chen, Y., Feng, S., Kumar, V. & Gao, Y.-G. Structure of EF-G–ribosome complex in a pre-translocation state. Nature Struct. Mol. Biol. 20, 1077–1084 (2013). This paper (along with references 39, 68 and 88) is one of four simultaneously published crystal structures of translocation intermediates.
Tourigny, D. S., Fernandez, I. S., Kelley, A. C. & Ramakrishnan, V. Elongation factor G bound to the ribosome in an intermediate state of translocation. Science 340, 1235490 (2013).
Martemyanov, K. A. & Gudkov, A. T. Domain IV of elongation factor G from Thermus thermophilus is strictly required for translocation. FEBS Lett. 452, 155–159 (1999).
Evans, R. N., Blaha, G., Bailey, S. & Steitz, T. A. The structure of LepA, the ribosomal back translocase. Proc. Natl Acad. Sci. USA 105, 4673–4678 (2008). This paper shows the crystal structure of EF4.
Schäfer, M. A. et al. Codon–anticodon interaction at the P site is a prerequisite for tRNA interaction with the small ribosomal subunit. J. Biol. Chem. 277, 19095–19105 (2002).
Frank, J. & Agrawal, R. K. A ratchet-like inter-subunit reorganization of the ribosome during translocation. Nature 406, 318–322 (2000).
Munro, J. B., Altman, R. B., O'Connor, N. & Blanchard, S. C. Identification of two distinct hybrid state intermediates on the ribosome. Mol. Cell 25, 505–517 (2007). This study is the first demonstration of the two ribosomal hybrid states, H1 and H2.
Moazed, D. & Noller, H. F. Intermediate states in the movement of transfer RNA in the ribosome. Nature 342, 142–148 (1989).
Agirrezabala, X. et al. Structural characterization of mRNA–tRNA translocation intermediates. Proc. Natl Acad. Sci. USA 109, 6094–6099 (2012).
Chen, C. et al. Single-molecule fluorescence measurements of ribosomal translocation dynamics. Mol. Cell 42, 367–377 (2011).
Cornish, P. V., Ermolenko, D. N., Noller, H. F. & Ha, T. Spontaneous intersubunit rotation in single ribosomes. Mol. Cell 30, 578–588 (2008).
Munro, J. B., Altman, R. B., Tung, C. S., Sanbonmatsu, K. Y. & Blanchard, S. C. A fast dynamic mode of the EF-G-bound ribosome. EMBO J. 29, 770–781 (2010).
Horan, L. H. & Noller, H. F. Intersubunit movement is required for ribosomal translocation. Proc. Natl Acad. Sci. USA 104, 4881–4885 (2007).
Warner, J. R. & Rich, A. The number of soluble RNA molecules on reticulocyte polyribosomes. Proc. Natl Acad. Sci. USA 51, 1134–1141 (1964).
Remme, J., Margus, T., Villems, R. & Nierhaus, K. H. The third ribosomal tRNA-binding site, the E site, is occupied in native polysomes. Eur. J. Biochem. 183, 281–284 (1989).
Rheinberger, H.-J. & Nierhaus, K. H. Testing an alternative model for the ribosomal peptide elongation cycle. Proc. Natl Acad. Sci. USA 80, 4213–4217 (1983).
Spahn, C. M. et al. Localization of the ribosomal protection protein Tet(O) on the ribosome and the mechanism of tetracycline resistance. Mol. Cell. 7, 1037–1045 (2001).
Spahn, C. M. et al. Domain movements of elongation factor eEF2 and the eukaryotic 80S ribosome facilitate tRNA translocation. EMBO J. 23, 1008–1019 (2004).
Zhang, W., Dunkle, J. A. & Cate, J. H. Structures of the ribosome in intermediate states of ratcheting. Science 325, 1014–1017 (2009).
Guo, Z. & Noller, H. F. Rotation of the head of the 30S ribosomal subunit during mRNA translocation. Proc. Natl Acad. Sci. USA 109, 20391–22034 (2012).
Ramrath, D. J. et al. The complex of tmRNA-SmpB and EF-G on translocating ribosomes. Nature 485, 526–529 (2012).
Alexeeva, E. V., Shpanchenko, O. V., Dontsova, O. A., Bogdanov, A. A. & Nierhaus, K. H. Interaction of mRNA with the Escherichia coli ribosome: accessibility of phosphorothioate-containing mRNA bound to ribosomes for iodine cleavage. Nucleic Acids Res. 24, 2228–2235 (1996).
Wurmbach, P. & Nierhaus, K. H. Codon–anticodon interaction at the ribosomal P (peptidyl-tRNA) site. Proc. Natl Acad. Sci. USA 76, 2143–2147 (1979).
Lührmann, R., Eckhardt, H. & Stöffler, G. Codon–anticodon interaction at the ribosomal peptidyl-site. Nature 280, 423–425 (1979).
Jenner, L., Rees, B., Yusupov, M. & Yusupova, G. Messenger RNA conformations in the ribosomal E site revealed by X-ray crystallography. EMBO Rep. 8, 846–850 (2007).
Schilling-Bartetzko, S., Bartetzko, A. & Nierhaus, K. H. Kinetic and thermodynamic parameters for transfer RNA binding to the ribosome and for the translocation reaction. J. Biol. Chem. 267, 4703–4712 (1992).
Shoji, S., Walker, S. E. & Fredrick, K. Ribosomal translocation: one step closer to the molecular mechanism. ACS Chem. Biol. 20, 93–107 (2009).
Cukras, A. R., Southworth, D. R., Brunelle, J. L., Culver, G. M. & Green, R. Ribosomal proteins S12 and S13 function as control elements for translocation of the mRNA:tRNA complex. Mol. Cell 12, 321–328 (2003).
Liu, Q. & Fredrick, K. Contribution of intersubunit bridges to the energy barrier of ribosomal translocation. Nucleic Acids Res. 41, 565–574 (2013).
Schuwirth, B. S. et al. Structures of the bacterial ribosome at 3.5 Å resolution. Science 310, 827–834 (2005). This paper reports the first crystal structure of an E. coli ribosome and also demonstrates the closure of the A790 gate.
Zhou, J., Lancaster, L., Donohue, J. P. & Noller, H. F. Crystal structures of EF-G-ribosome complexes trapped in intermediate states of translocation. Science 340, 1236086 (2013). This paper reports the crystal structure of a translocation intermediate showing 16S rRNA bases C1397 and A1503 intercalating with the mRNA, thereby preventing back-translocation.
Valle, M. et al. Locking and unlocking of ribosomal motions. Cell 114, 123–134 (2003).
Schmeing, T. M. et al. The crystal structure of the ribosome bound to EF-Tu and aminoacyl-tRNA. Science 326, 688–694 (2009).
Trabuco, L. G. et al. The role of L1 stalk-tRNA interaction in the ribosome elongation cycle. J. Mol. Biol. 402, 741–760 (2010).
Nierhaus, K. H. The allosteric three-site model for the ribosomal elongation cycle: features and future. Biochemistry 29, 4997–5008 (1990).
Rheinberger, H.-J. & Nierhaus, K. H. Allosteric interactions between the ribosomal transfer RNA-binding sites A and E. J. Biol. Chem. 261, 9133–9139 (1986).
Cornish, P. V. et al. Following movement of the L1 stalk between three functional states in single ribosomes. Proc. Natl Acad. Sci. USA 106, 2571–2576 (2009).
Munro, J. B. et al. Spontaneous formation of the unlocked state of the ribosome is a multistep process. Proc. Natl Acad. Sci. USA 107, 709–714 (2010).
Subramanian, A. R. & Dabbs, E. R. Functional studies on ribosomes lacking protein L1 from mutant Escherichia coli. Eur. J. Biochem. 112, 425–430 (1980).
Sander, G. Ribosomal protein L1 from Escherichia coli. Its role in the binding of tRNA to the ribosome and in elongation factor G-dependent GTP hydrolysis. J. Biol. Chem. 258, 10098–10103 (1983).
Rodnina, M. V., Savelsbergh, A., Katunin, V. I. & Wintermeyer, W. Hydrolysis of GTP by elongation factor G drives tRNA movement on the ribosome. Nature 385, 37–41 (1997).
Zavialov, A. V. & Ehrenberg, M. Peptidyl-tRNA regulates the GTPase activity of translational factors. Cell 114, 113–122 (2003).
Pan, D., Kirillov, S. V. & Cooperman, B. S. Kinetically competent intermediates in the translocation step of protein synthesis. Mol. Cell 25, 519–529 (2007).
Peske, F., Savelsbergh, A., Katunin, V. I., Rodnina, M. V. & Wintermeyer, W. Conformational changes of the small ribosomal subunit during elongation factor G-dependent tRNA–mRNA translocation. J. Mol. Biol. 343, 1183–1194 (2004).
Hausner, T. P., Atmadja, J. & Nierhaus, K. H. Evidence that the G2661 region of 23S rRNA is located at the ribosomal binding sites of both elongation factors. Biochimie 69, 911–923 (1987).
Clementi, N. & Polacek, N. Ribosome-associated GTPases: the role of RNA for GTPase activation. RNA Biol. 7, 521–527 (2010).
Endo, Y. & Wool, I. G. The site of action of α-sarcin on eukaryotic ribosomes. The sequence at the α-sarcin cleavage site in 28S rRNA. J. Biol. Chem. 257, 9054–9060 (1982).
Garcia-Ortega, L., Alvarez-Garcia, E., Gavilanes, J. G., Martinez-del-Pozo, A. & Joseph, S. Cleavage of the sarcin-ricin loop of 23S rRNA differentially affects EF-G and EF-Tu binding. Nucleic Acids Res. 38, 4108–4119 (2010).
Shi, X., Khade, P. K., Sanbonmatsu, K. Y. & Joseph, S. Functional role of the sarcin–ricin loop of the 23S rRNA in the elongation cycle of protein synthesis. J. Mol. Biol. 419, 125–138 (2012).
Clementi, N., Chirkova, A., Puffer, B., Micura, R. & Polacek, N. Atomic mutagenesis reveals A2660 of 23S ribosomal RNA as key to EF-G GTPase activation. Nature Chem. Biol. 6, 344–351 (2010).
Pulk, A. & Cate, J. H. Control of ribosomal subunit rotation by elongation factor G. Science 340, 1235970 (2013).
Berchtold, H. et al. Crystal structure of active elongation factor Tu reveals major domain rearrangements. Nature 365, 126–132 (1993).
Dibb, N. J. & Wolfe, P. B. lep operon proximal gene is not required for growth or secretion by Escherichia coli. J. Bacteriol. 166, 83–87 (1986).
Bijlsma, J. J., Lie, A. L. M., Nootenboom, I. C., Vandenbroucke-Grauls, C. M. & Kusters, J. G. Identification of loci essential for the growth of Helicobacter pylori under acidic conditions. J. Infect. Dis. 182, 1566–1569 (2000).
Froschauer, E. M., Kolisek, M., Dieterich, F., Schweigel, M. & Schweyen, R. J. Fluorescence measurements of free [Mg2+] by use of mag-fura 2 in Salmonella enterica. FEMS Microbiol. Lett. 237, 49–55 (2004).
Richey, B. et al. Variability of intracellular ionic environment of Escherichia coli. J. Biol. Chem. 262, 7157–7164 (1987).
Bartetzko, A. & Nierhaus, K. H. A simple Mg2+/NH4+/polyamine system for poly(U) dependent poly(Phe) synthesis with near in vivo characteristics. Methods Enzymol. 164, 650–658 (1988).
Szaflarski, W. et al. New features of the ribosome and ribosomal inhibitors: non-enzymatic recycling, misreading and back-translocation. J. Mol. Biol. 380, 193–205 (2008).
Shoji, S., Janssen, B. D., Hayes, C. S. & Fredrick, K. Translation factor LepA contributes to tellurite resistance in Escherichia coli but plays no apparent role in the fidelity of protein synthesis. Biochimie 92, 157–163 (2010).
Ogle, J. M. & Ramakrishnan, V. Structural insights into translational fidelity. Annu. Rev. Biochem. 74, 129–177 (2005).
Zhang, G., Hubalewska, M. & Ignatova, Z. Transient ribosomal attenuation coordinates protein synthesis and co-translational folding. Nature Struct. Mol. Biol. 16, 274–280 (2009).
Kunte, H. Osmoregulation in bacteria: compatible solute accumulation and osmosensing. Environ. Chem. 3, 94–99 (2006).
Bauerschmitt, H., Funes, S. & Herrmann, J. M. The membrane-bound GTPase Guf1 promotes mitochondrial protein synthesis under suboptimal conditions. J. Biol. Chem. 283, 17139–17146 (2008).
Gavrilova, L. P. & Spirin, A. S. “Nonenzymatic” translation. Methods Enzymol. 30, 452–462 (1974).
Bergemann, K. & Nierhaus, K. H. Spontaneous, elongation factor G independent translocation of Escherichia coli ribosomes. J. Biol. Chem. 258, 15105–15113 (1983).
Gavrilova, L. P. & Spirin, A. S. Stimulation of 'non-enzymic' translocation in ribosome by p-chloromercuribenzoate. FEBS Lett. 17, 324–326 (1971).
Shoji, S., Walker, S. E. & Fredrick, K. Reverse translocation of tRNA in the ribosome. Mol. Cell 24, 931–942 (2006).
Konevega, A. L. et al. Spontaneous reverse movement of mRNA-bound tRNA through the ribosome. Nature Struct. Mol. Biol. 14, 318–324 (2007).
Stapulionis, R. & Deutscher, M. P. A channeled tRNA cycle during mammalian protein synthesis. Proc. Natl Acad. Sci. USA 92, 7158–7161 (1995).
Yegian, C. D., Stent, G. S. & Martin, E. M. Intracellular condition of Escherichia coli transfer RNA. Proc. Natl Acad. Sci. USA 55, 839–846 (1966).
Dittmar, K. A., Sorensen, M. A., Elf, J., Ehrenberg, M. & Pan, T. Selective charging of tRNA isoacceptors induced by amino-acid starvation. EMBO Rep. 6, 151–157 (2005).
Fredrick, K. & Noller, H. F. Catalysis of ribosomal translocation by sparsomycin. Science 300, 1159–1162 (2003).
Fischer, N., Konevega, A. L., Wintermeyer, W., Rodnina, M. V. & Stark, H. Ribosome dynamics and tRNA movement by time-resolved electron cryomicroscopy. Nature 466, 329–333 (2010).
Munro, J. B., Sanbonmatsu, K. Y., Spahn, C. M. & Blanchard, S. C. Navigating the ribosome's metastable energy landscape. Trends Biochem. Sci. 34, 390–400 (2009).
Kavran, J. M. & Steitz, T. A. Structure of the base of the L7/L12 stalk of the Haloarcula marismortui large ribosomal subunit: analysis of L11 movements. J. Mol. Biol. 371, 1047–1059 (2007).
Bocharov, E. V. et al. From structure and dynamics of protein L7/L12 to molecular switching in ribosome. J. Biol. Chem. 279, 17697–17706 (2004).
Budkevich, T. et al. Structure and dynamics of the mammalian ribosomal pretranslocation complex. Mol. Cell 44, 214–224 (2011). This paper reports cryo-EM analyses of various functional states of rabbit liver ribosomes.
Brilot, A.F., Korostelev, A.A., Ermolenko, D.N. & Grigorieff, N. Structure of the ribosome with elongation factor G trapped in the pretranslocation state. Proc. Natl Acad. Sci USA http://dx.doi.org/10.1073/pnas.1311423110 (2013).
Ramrath, D.J. et al. Visualization of two transfer RNAs trapped in transit during elongation factor G-mediated translocation. Proc. Natl Acad. Sci. USA http://dx.doi.org/10.1073/pnas.1320387110 (2013).
Acknowledgements
The authors thank J. Harms (Hamburg) N. Polacek (University Bern, Switzerland) and T. Sprink (Charité, Berlin) for help and discussions. H.Y. and C.M.T.S acknowledge the support of the Deutsche Forschergruppe (DFG), Forschergruppe 1805, and Y.Q. is grateful for research grants from the Major State Basic Research of China 973 project (grant 2012CB911000) and the National Natural Science Foundation of China (grants 31270847 and 31322015).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
DATABASES
Supplementary information
Supplementary information S1 (figure)
First contacts of elongation factor G (EF-G) with the ribosome. (PDF 304 kb)
Supplementary information S2 (figure)
EF-G and EF4 on the ribosome. (PDF 273 kb)
Supplementary information S3 (figure)
tRNAs adopt classical and hybrid binding states on the ribosome. (PDF 307 kb)
Supplementary information S4 (figure)
16rRNA interactions with mRNA in the TIpost transition station. (PDF 301 kb)
Glossary
- Decoding
-
Selection of the cognate ternary complex of aminoacyl- tRNA–EF-Tu–GTP on the basis of correct codon-anticodon interactions between the mRNA and tRNA, respectively.
- Single-turnover experiments
-
Experiments in which the conditions are set such that the catalyst (for example, the ribosome) only undergoes a single round of catalysis.
- Single-molecule FRET
-
(Single-molecule Förster resonance energy transfer). A phenomenon in which energy induced by light excitation is transferred from one fluorophore to another in a distance-dependent manner, observed on a single complex or molecule.
- Peptidyl-prolyl cis–trans isomerase
-
An enzyme that belongs to the peptidyl-prolyl isomerase (PPIase) family that catalyses the transition of a proline residue between cis and trans conformations by reducing the activation-energy barrier that separates these two conformations.
- Sarcin–ricin loop
-
(SRL). The loop of helix H95 (G2654–A2665; E. coli nomenclature), which contains the longest universally conserved ribosomal RNA (rRNA) sequence. Its name derives from the observations that removing base A2660 by the N-glycosidase ricin or cleaving the 23S rRNA after G2661 by the RNase α-sarcin impairs the binding and GTPase activity of both elongation factor Tu (EF-Tu) and EF-G, thereby blocking translation.
- Activation-energy barrier
-
The energy barrier that separates reactants and products in a chemical reaction.
- Polysomes
-
mRNAs to which more than one ribosome is bound.
- Exocyclic group
-
A chemical group attached to a cyclic structure. For example, adenine contains an exocyclic amino group at position 6, and guanine contains a hydroxyl group at the same position.
Rights and permissions
About this article
Cite this article
Yamamoto, H., Qin, Y., Achenbach, J. et al. EF-G and EF4: translocation and back-translocation on the bacterial ribosome. Nat Rev Microbiol 12, 89–100 (2014). https://doi.org/10.1038/nrmicro3176
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3176
This article is cited by
-
Structural insights of the elongation factor EF-Tu complexes in protein translation of Mycobacterium tuberculosis
Communications Biology (2022)
-
A genetic system for targeted mutations to disrupt and restore genes in the obligate bacterium, Ehrlichia chaffeensis
Scientific Reports (2017)
-
EF4 disengages the peptidyl-tRNA CCA end and facilitates back-translocation on the 70S ribosome
Nature Structural & Molecular Biology (2016)
-
Steric interactions lead to collective tilting motion in the ribosome during mRNA–tRNA translocation
Nature Communications (2016)
-
Mammalian elongation factor 4 regulates mitochondrial translation essential for spermatogenesis
Nature Structural & Molecular Biology (2016)