Key Points
-
Initiator proteins direct the assembly of DNA synthesis enzymes at chromosomal sites in a highly regulated manner. In bacteria, the initiator DnaA cooperatively oligomerizes at the site of replication fork formation to direct melting of DNA duplex strands and loading of the replicative helicase DnaB. DnaA belongs to the AAA+ superfamily of ATPases, and the binding and hydrolysis of nucleotides have a crucial role in controlling DnaA activity.
-
The involvement of DnaA at multiple stages of replication initiation is facilitated by its highly modular domain architecture. Interactions with DnaB are mediated by the extreme N-terminal region, whereas a helix–turn–helix motif located in the C-terminal domain confers the sequence-specific DNA-binding activity needed to recognize the origin of replication. The central region of the protein comprises the highly conserved AAA+ fold and primary oligomerization site.
-
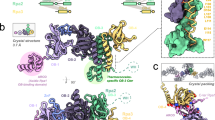
Recent structural studies have helped to reveal how ATP facilitates DnaA auto-assembly. Oligomerized DnaA forms a right-handed helical filament that is stabilized through the formation of a bipartite nucleotide-binding pocket between two successive AAA+ domains. The structural features that allow for DnaA filament formation seem to be conserved among cellular initiators from archaea and eukaryotes.
-
Bacteria use many regulatory strategies that are dedicated to controlling the initiation of DNA replication. Among these, origin sequestration, DnaA titration, dnaA autoregulation and DnaA inactivation closely monitor events that occur at the bacterial replication origin to optimize the intracellular levels and activity of DnaA.
-
The involvement of DNA architectural factors at oriC highlight the cellular context in which initiation takes place. DNA-bending proteins such as Fis (factor for inversion stimulation) and IHF (integration host factor) appear to regulate and fine tune the assembly of DnaA at the replication origin during the cell cycle.
Abstract
In all organisms, multi-subunit replicases are responsible for the accurate duplication of genetic material during cellular division. Initiator proteins control the onset of DNA replication and direct the assembly of replisomal components through a series of precisely timed protein–DNA and protein–protein interactions. Recent structural studies of the bacterial protein DnaA have helped to clarify the molecular mechanisms underlying initiator function, and suggest that key structural features of cellular initiators are universally conserved. Moreover, it appears that bacteria use a diverse range of regulatory strategies dedicated to tightly controlling replication initiation; in many cases, these mechanisms are intricately connected to the activities of DnaA at the origin of replication. This Review presents an overview of both the mechanism and regulation of bacterial DNA replication initiation, with emphasis on the features that are similar in eukaryotic and archaeal systems.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jacob, F., Brenner, S. & Couzin, F. On the regulation of DNA replication in bacteria. Cold Spring Harb. Symp. Quant. Biol. 28, 329–348 (1963).
Fuller, R. S., Funnell, B. E. & Kornberg, A. The dnaA protein complex with the E. coli chromosomal replication origin (oriC) and other DNA sites. Cell 38, 889–900 (1984).
Matsui, M., Oka, A., Takanami, M., Yasuda, S. & Hirota, Y. Sites of dnaA protein-binding in the replication origin of the Escherichia coli K-12 chromosome. J. Mol. Biol. 184, 529–533 (1985).
Schaper, S. & Messer, W. Interaction of the initiator protein DnaA of Escherichia coli with its DNA target. J. Biol. Chem. 270, 17622–17626 (1995).
Grimwade, J. E., Ryan, V. T. & Leonard, A. C. IHF redistributes bound initiator protein, DnaA, on supercoiled oriC of Escherichia coli. Mol. Microbiol. 35, 835–844 (2000).
Ryan, V. T., Grimwade, J. E., Nievera, C. J. & Leonard, A. C. IHF and HU stimulate assembly of pre-replication complexes at Escherichia coli oriC by two different mechanisms. Mol. Microbiol. 46, 113–124 (2002).
McGarry, K. C., Ryan, V. T., Grimwade, J. E. & Leonard, A. C. Two discriminatory binding sites in the Escherichia coli replication origin are required for DNA strand opening by initiator ATP–DnaA. Proc. Natl Acad. Sci. USA 101, 2811–2816 (2004).
Speck, C. & Messer, W. Mechanism of origin unwinding: sequential binding of DnaA to double- and single-stranded DNA. EMBO J. 20, 1469–1476 (2001).
Speck, C., Weigel, C. & Messer, W. ATP- and ADP–dnaA protein, a molecular switch in gene regulation. EMBO J. 18, 6169–6176 (1999).
Cassler, M. R., Grimwade, J. E. & Leonard, A. C. Cell cycle-specific changes in nucleoprotein complexes at a chromosomal replication origin. EMBO J. 14, 5833–5841 (1995).
Samitt, C. E., Hansen, F. G., Miller, J. F. & Schaechter, M. In vivo studies of DnaA binding to the origin of replication of Escherichia coli. EMBO J. 8, 989–993 (1989).
Leonard, A. C. & Grimwade, J. E. Building a bacterial orisome: emergence of new regulatory features for replication origin unwinding. Mol. Microbiol. 55, 978–985 (2005).
Funnell, B. E, Baker, T. A. & Kornberg, A. In vitro assembly of a prepriming complex at the origin of the Escherichia coli chromosome. J. Biol. Chem. 262, 10327–10334 (1987).
Bramhill, D. & Kornberg, A. Duplex opening by dnaA protein at novel sequences in initiation of replication at the origin of the E. coli chromosome. Cell 52, 743–755 (1988). A classic paper that defines many of the biochemical steps in DnaA function at oriC.
Katayama, T., Kubota, T., Kurokawa, K., Crooke, E. & Sekimizu K. The initiator function of DnaA protein is negatively regulated by the sliding clamp of the E. coli chromosomal replicase. Cell 94, 61–71 (1998).
Crooke, E., Castuma, C. E. & Kornberg, A. The chromosome origin of Escherichia coli stabilizes DnaA protein during rejuvenation by phospholipids. J. Biol. Chem. 267, 16779–16782 (1992).
Sekimizu, K. & Kornberg, A. Cardiolipin activation of dnaA protein, the initiation protein of replication in Escherichia coli. J. Biol. Chem. 263, 7131–7135 (1988).
Xia, W. & Dowhan, W. In vivo evidence for the involvement of anionic phospholipids in initiation of DNA replication in Escherichia coli. Proc. Natl Acad. Sci. USA 92, 783–787 (1995).
Messer, W. The bacterial replication initiator DnaA. DnaA and oriC, the bacterial mode to initiate DNA replication. FEMS Microbiol. Rev. 26, 355–374 (2002).
Zawilak-Pawlik, A. et al. Architecture of bacterial replication initiation complexes: orisomes from four unrelated bacteria. Biochem J. 389, 471–481 (2005). Compares the architecture of DnaA oriC nucleoprotein complexes among four evolutionarily distantly related bacterial species that possess diverse origin organization.
Marczynski, G. T. & Shapiro, L. Control of chromosome replication in Caulobacter crescentus. Annu. Rev. Microbiol. 56, 625–656 (2002).
Quon, K. C., Marczynski, G. T. & Shapiro, L. Cell cycle control by an essential bacterial two-component signal transduction protein. Cell 84, 83–93 (1996).
Quon, K. C., Yang, B, Domian, I. J., Shapiro, L & Marczynski, G. T. Negative control of bacterial DNA replication by a cell cycle regulatory protein that binds at the chromosome origin. Proc. Natl Acad. Sci. USA 95, 120–125 (1998).
Siam, R. & Marczynski, G. T. Cell cycle regulator phosphorylation stimulates two distinct modes of binding at a chromosome replication origin. EMBO J. 19, 1138–1147 (2000).
Simmons, L. A., Felczak, M. & Kaguni, J. M. DnaA protein of Escherichia coli: oligomerization at the E. coli chromosomal origin is required for initiation and involves specific N-terminal amino acids. Mol. Microbiol. 49, 849–858 (2003).
Seitz, H., Weigel, C. & Messer, W. The interaction domains of the DnaA and DnaB replication proteins of Escherichia coli. Mol. Microbiol. 37, 1270–1279 (2000).
Sutton, M. D., Carr, K. M., Vicente, M. & Kaguni, J. M. Escherichia coli DnaA protein. The N-terminal domain and loading of DnaB helicase at the E. coli chromosomal origin. J. Biol. Chem. 273, 34255–34262 (1998).
Weigel, C., Schmidt, A., Seitz, H., Tungler, D., Welzeck, M. & Messer, W. The N-terminus promotes oligomerization of the Escherichia coli initiator protein DnaA. Mol. Microbiol. 34, 53–66 (1999).
Messer, W. et al. Functional domains of DnaA proteins. Biochimie 81, 819–825 (1999).
Roth, A. & Messer, W. The DNA binding domain of the initiator protein DnaA. EMBO J. 14, 2106–2111 (1995).
Sutton, M. D. & Kaguni, J. M. Threonine 435 of Escherichia coli DnaA protein confers sequence-specific DNA binding activity. J. Biol. Chem. 272, 23017–23024 (1997).
Blaesing, F., Weigel, C., Welzeck, M. & Messer, W. Analysis of the DNA-binding domain of Escherichia coli DnaA protein. Mol. Microbiol. 36, 36557–36569 (2000).
Fujikawa, N. et al. Structural basis of replication origin recognition by the DnaA protein. Nucleic Acids Res. 31, 2077–2086 (2003).
Erzberger, J. P., Pirruccello, M. M. & Berger, J. M. The structure of bacterial DnaA: implications for general mechanisms underlying DNA replication initiation. EMBO J. 21, 4763–4773 (2002). The high-resolution crystal structure of A. aeolicus ADP–DnaA showing the spatial arrangement of the AAA+ and DNA-binding domains. Revealed a remarkable degree of structural homology with the archaeal initiator Cdc6/Orc1 from reference 45.
Erzberger, J. P., Mott, M. L. & Berger, J. M. Structural basis for ATP-dependent DnaA assembly and replication-origin remodeling. Nature Struct. Mol. Biol. 8, 676–683 (2006). The high-resolution crystal structure revealed the molecular basis for ATP-dependent oligomerization by DnaA. The right-handed helical filament of A. aeolicus DnaA bound to AMP-PCP represents a novel AAA+ assembly mode mediated by a helical insert specific to the initiator AAA+ clade.
Garner, J., Durrer, P., Kitchen, J., Brunner, J. & Crooke, E. Membrane-mediated release of nucleotide from an initiator of chromosomal replication, Escherichia coli DnaA, occurs with insertion of a distinct region of the protein into the lipid bilayer. J. Biol. Chem. 273, 5167–5173 (1998).
Hase, M. et al. Site-directed mutational analysis for the membrane binding of DnaA protein. Identification of amino acids involved in the functional interaction between DnaA protein and acidic phospholipids. J. Biol. Chem. 273, 28651–28656 (1998).
Makise, M., Mima, S., Tsuchiya, T. & Mizushima, T. Molecular mechanism for functional interaction between DnaA protein and acidic phospholipids: identification of important amino acids. J. Biol. Chem. 276, 7450–7456 (2001).
Makise, M., Mima, S., Tsuchiya, T. & Mizushima, T. Identification of amino acids involved in the functional interaction between DnaA protein and acidic phospholipids. J. Biol. Chem. 275, 4513–4518 (2000).
Zheng, W., Li, Z., Skarstad, K. & Crooke, E. Mutations in DnaA protein suppress the growth arrest of acidic phospholipid-deficient Escherichia coli cells. EMBO J. 20, 1164–1172 (2001).
Koonin, E. V. DnaC protein contains a modified ATP-binding motif and belongs to a novel family of ATPases including also DnaA. Nucleic Acids Res. 20, 1997 (1992).
Iyer, L. M., Leipe, D. D., Koonin, E. V. & Aravind, L. Evolutionary history and higher order classification of AAA+ ATPases. J. Struct. Biol. 146, 11–31 (2004). Provides a phylogenetic comparison of AAA+ proteins and distinguishes subclasses on the basis of distinct structural elements.
Neuwald, A. F., Aravind, L., Spouge, J. L. & Koonin, E. V. AAA+: A class of chaperone-like ATPases associated with the assembly, operation, and disassembly of protein complexes. Genome Res. 9, 27–43 (1999).
Felczak, M. M. & Kaguni, J. M. The box VII motif of Escherichia coli DnaA protein is required for DnaA oligomerization at the E. coli replication origin. J. Biol. Chem. 279, 51156–51162 (2004).
Liu J. et al. Structure and function of Cdc6/Cdc18: implications for origin recognition and checkpoint control. Mol. Cell 6, 637–648 (2000).
Kong, X. P., Onrust, R., O'Donnell, M. & Kuriyan, J. Three-dimensional structure of the β subunit of E. coli DNA polymerase III holoenzyme: a sliding DNA clamp. Cell 69, 425–437 (1992).
Krishna, T. S., Kong, X. P., Gary, S., Burgers, P. M. & Kuriyan J. Crystal structure of the eukaryotic DNA polymerase processivity factor PCNA. Cell 79, 1233–1243 (1994).
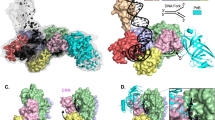
Clarey, M. G. et al. Nucleotide-dependent conformational changes in the DnaA-like core of the origin recognition complex. Nature Struct. Mol. Biol. 8, 684–690 (2006). Electron microscopy reconstructions depicting nucleotide-dependent conformational changes of the D. melanogaster ORC. Docking studies indicate that ORC can accommodate a DnaA helical pentamer, suggesting a similar arrangement of the ORC AAA+ domains.
Speck, C., Chen, Z., Li, H. & Stillman B. ATPase-dependent cooperative binding of ORC and Cdc6 to origin DNA. Nature Struct. Mol. Biol. 12, 965–971 (2005).
Donachie, W. D. Relationship between cell size and time of initiation of DNA replication. Nature 219, 1077–1079 (1968).
Skarstad, K., Boye, E. & Steen, H. B. Timing of initiation of chromosome replication in individual Escherichia coli cells. EMBO J. 5, 1711–1717 (1986).
Zyskind, J. W. & Smith, D. W. The bacterial origin of replication, oriC. Cell 46, 489–490 (1986).
Brendler, T., Abeles, A. & Austin, S. A protein that binds to the P1 origin core and the oriC 13mer region in a methylation-specific fashion is the product of the host seqA gene. EMBO J. 14, 4083–4089 (1995).
Slater, S., Wold, S., Lu, M., Boye, E., Skarstad, K. & Kleckner, N. E. coli SeqA protein binds oriC in two different methyl-modulated reactions appropriate to its roles in DNA replication initiation and origin sequestration. Cell 82, 927–936 (1995).
Nievera, C., Torgue, J. J., Grimwade, J. E. & Leonard, A C. SeqA blocking of DnaA-oriC interactions ensures staged assembly of the E. coli pre-RC. Mol. Cell 24, 581–592 (2006). Presents evidence that SeqA-mediated origin sequestration blocks the binding of DnaA to several low-affinity sites following the completion of initiation, preventing further rounds of initiation and resetting DnaA occupancy at oriC to a pre-initiation state.
Brendler, T., Sawitzke, J., Sergueev, K. & Austin, S. A case for sliding SeqA tracts at anchored replication forks during Escherichia coli chromosome replication and segregation. EMBO J. 19, 6249–6258 (2000).
Lu, M., Campbell, J. L., Boye, E. & Kleckner, N. SeqA: a negative modulator of replication initiation in E. coli. Cell 77, 413–426 (1994).
von Freiesleben, U., Rasmussen, K. V. & Schaechter, M. SeqA limits DnaA activity in replication from oriC in Escherichia coli. Mol. Microbiol. 14, 763–772 (1994).
Boye, E., Stokke, T., Kleckner, N. & Skarstad, K. Coordinating DNA replication initiation with cell growth: differential roles for DnaA and SeqA proteins. Proc. Natl Acad. Sci. USA 93, 12206–12211 (1996).
Brendler, T. & Austin, S. Binding of SeqA protein to DNA requires interaction between two or more complexes bound to separate hemimethylated GATC sequences. EMBO J. 18, 2304–2310 (1999).
Guarne, A., Zhao, Q., Ghirlando, R. & Yang, W. Insights into negative modulation of E. coli replication initiation from the structure of SeqA-hemimethylated DNA complex. Nature Struct. Biol. 9, 839–843 (2002).
Skarstad, K., Lueder, G., Lurz, R., Speck, C. & Messer, W. The Escherichia coli SeqA protein binds specifically and co-operatively to two sites in hemimethylated and fully methylated oriC. Mol. Microbiol. 36, 1319–1326 (2000).
Guarne, A. et al. Crystal structure of a SeqA-N filament: implications for DNA replication and chromosome organization. EMBO J. 24, 1502–1511 (2005). Identifies the regions of the SeqA N-terminal domain that are important for oligomerization and, together with the structural data on the C-terminal domain (in reference 61), presents an intriguing, left-handed filament model for the full-length SeqA protein.
Odsbu, I., Klungsoyr, H. K., Fossum, S. & Skarstad, K. Specific N-terminal interactions of the Escherichia coli SeqA protein are required to form multimers that restrain negative supercoils and form foci. Genes Cells. 10, 1039–1049 (2005).
Roth, A. & Messer, W. High-affinity binding sites for the initiator protein DnaA on the chromosome of Escherichia coli. Mol. Microbiol. 28, 395–401 (1998).
Kitagawa, R., Mitsuki, H., Okazaki, T. & Ogawa, T. A novel DnaA protein-binding site at 94.7 min on the Escherichia coli chromosome. Mol. Microbiol. 19, 1137–1147 (1996).
Kitagawa, R., Ozaki, T., Moriya, S. & Ogawa, T. Negative control of replication initiation by a novel chromosomal locus exhibiting exceptional affinity for Escherichia coli DnaA protein. Genes Dev. 12, 3032–3043 (1998).
Messer, W. & Weigel, C. DnaA initiator — also a transcription factor. Mol. Microbiol. 24, 1–6 (1997).
Atlung, T., Clausen, E. S. & Hansen, F. G. Autoregulation of the dnaA gene of Escherichia coli K12. Mol. Gen. Genet. 200, 442–450 (1985).
Braun, R. E., O'Day, K. & Wright, A. Autoregulation of the DNA replication gene dnaA in E. coli K-12. Cell 40, 159–169 (1985).
Nishida S. et al. A nucleotide switch in the Escherichia coli DnaA protein initiates chromosomal replication: evidence from a mutant DnaA protein defective in regulatory ATP hydrolysis in vitro and in vivo. J. Biol. Chem. 277, 14986–14995 (2002).
Kato, J. & Katayama, T. Hda, a novel DnaA-related protein, regulates the replication cycle in Escherichia coli. EMBO J. 20, 4253–4262 (2001).
Camara, J. E., Skarstad, K. & Crooke, E. Controlled initiation of chromosomal replication in Escherichia coli requires functional Hda protein. J. Bacteriol. 185, 3244–3248 (2003).
Su'etsugu, M., Shimuta, T. R., Ishida, T., Kawakami, H. & Katayama, T. Protein associations in ATP–DnaA hydrolysis mediated by the Hda-replicase clamp complex. J. Biol. Chem. 280, 6528–6536 (2005). Shows that the conserved arginine finger of the Hda box VII motif is essential for stimulating DnaA ATP hydrolysis.
Hayashi, M., Ogura, Y., Harry, E. J., Ogasawara, N. & Moriya, S. Bacillus subtilis YabA is involved in determining the timing and synchrony of replication initiation. FEMS Microbiol. Lett. 247, 73–79 (2005).
Noirot-Gros, M. F. et al. An expanded view of bacterial DNA replication. Proc. Natl Acad. Sci. USA 99, 8342–8347 (2002).
Noirot-Gros, M. F. et al. Functional dissection of YabA, a negative regulator of DNA replication initiation in Bacillus subtilis. Proc. Natl Acad. Sci. USA 103, 2368–2373 (2006).
Schwob, E. Flexibility and governance in eukaryotic DNA replication. Curr. Opin. Microbiol. 7, 680–690 (2004).
Weinreich, M., Palacios DeBeer, M. A. & Fox, C. A. The activities of eukaryotic replication origins in chromatin. Biochim. Biophys. Acta 1677, 142–157 (2004).
Remus, D., Beall, E. L. & Botchan, M. R. DNA topology, not DNA sequence, is a critical determinant for Drosophila ORC-DNA binding. EMBO J. 23, 897–907 (2004).
Finkel, S. E. & Johnson, R. C. The Fis protein: it's not just for DNA inversion anymore. Mol. Microbiol. 6, 3257–3265 (1992).
Friedman, D. I. Integration host factor: a protein for all reasons. Cell 55, 545–554 (1988).
Schmid, M. B. More than just histone-like proteins. Cell. 63, 451–453 (1990).
Hwang, D. S. & Kornberg, A. Opening of the replication origin of Escherichia coli by DnaA protein with protein HU or IHF. J. Biol. Chem. 267, 23083–23086 (1992).
Zhang, W. et al. The Bacillus subtilis DnaD and DnaB proteins exhibit different DNA remodelling activities. J. Mol. Biol. 351, 66–75 (2005).
Goosen, N. & van de Putte, P. The regulation of transcription initiation by integration host factor. Mol. Microbiol. 16, 1–7 (1995).
Johnson, R. C. & Simon, M. I. Hin-mediated site-specific recombination requires two 26 bp recombination sites and a 60 bp recombinational enhancer. Cell 41, 781–791 (1985).
Koch, C. & Kahmann, R. Purification and properties of the Escherichia coli host factor required for inversion of the G segment in bacteriophage Mu. J. Biol. Chem. 261, 15673–15678 (1986).
Lorenz, M., Hillisch, A., Goodman, S. D. & Diekmann, S. Global structure similarities of intact and nicked DNA complexed with IHF measured in solution by fluorescence resonance energy transfer. Nucleic Acids Res. 27, 4619–4625 (1999).
Pan, C. Q. et al. Structure of the Escherichia coli Fis–DNA complex probed by protein conjugated with 1,10-phenanthroline copper(I) complex. Proc. Natl Acad. Sci. USA 91, 1721–1725 (1994).
Rice, P. A., Yang, S., Mizuuchi, K. & Nash, H. A. Crystal structure of an IHF–DNA complex: a protein-induced DNA U-turn. Cell 87, 1295–1306 (1996).
Filutowicz, M. & Roll, J. The requirement of IHF protein for extrachromosomal replication of the Escherichia coli oriC in a mutant deficient in DNA polymerase I activity. New Biol. 2, 818–827 (1990).
Gille, H., Egan, J. B., Roth, A. & Messer, W. The FIS protein binds and bends the origin of chromosomal DNA replication, oriC, of Escherichia coli. Nucleic Acids Res. 19, 4167–4172 (1991).
Polaczek, P. Bending of the origin of replication of E. coli by binding of IHF at a specific site. New Biol. 2, 265–271 (1990).
Boye, E., Lyngstadaas, A., Lobner-Olesen, A., Skarstad, K. & Wold, S. in DNA Replication and the Cell Cycle (eds Fanning, E., Knippers, R & Winnacker, E. L.) 15–26 (Springer, Berlin, 1993).
Hiasa, H. & Marians, K. J. Fis cannot support oriC DNA replication in vitro. J. Biol. Chem. 69, 24999–25003 (1994).
Wold, S., Crooke, E. & Skarstad, K. The Escherichia coli Fis protein prevents initiation of DNA replication from oriC in vitro. Nucleic Acids Res. 24, 3527–3532 (1996).
Ryan, V. T., Grimwade, J. E., Camara, J. E., Crooke, E., Leonard, A. C. Escherichia coli prereplication complex assembly is regulated by dynamic interplay among Fis, IHF and DnaA. Mol. Microbiol. 51, 1347–1359 (2004). Examines how Fis and IHF binding at oriC influences the progression of DnaA nucleoprotein formation during initiation.
Margulies, C. & Kaguni, J. M. The FIS protein fails to block the binding of DnaA protein to oriC, the Escherichia coli chromosomal origin. Nucleic Acids Res. 26, 5170–5175 (1998).
Schneider, R., Travers, A. & Muskhelishvili G. FIS modulates growth phase-dependent topological transitions of DNA in Escherichia coli. Mol. Microbiol. 26, 519–530 (1997).
Fekete, R. A., Venkova-Canova, T., Park, K. & Chattoraj, D. K. IHF-dependent activation of P1 plasmid origin by DnaA Mol. Microbiol. 62, 1739–1751 (2006).
Siam, R., Brassinga, A. K. C & Marczynski, G. T. A dual binding site for integration host factor and the response regulator CtrA inside the Caulobacter crescentus replication origin. J. Bacteriol. 185, 5563–5572 (2003).
Mackiewicz, P., Zakrzewska-Czerwinska, J., Zawilak, A., Dudek, M. R. & Cebrat, S. Where does bacterial replication start? Rules for predicting the oriC region. Nucleic Acids Res. 32, 3781–3791 (2004).
Erzberger, J. P. & Berger, J. M. Evolutionary relationships and structural mechanisms of AAA+ proteins. Annu. Rev. Biophys. Biomol. Struct. 35, 93–114 (2006).
Ogura, T., Whitehaeart, S. W. & Wilkinson, A. J. Conserved arginine residues implicated in ATP hydrolysis, nucleotide-sensing, and inter-subunit interactions in AAA and AAA+ ATPases. J. Struct. Biol. 146, 106–112 (2004).
Hanson, P. I. & Whiteheart, S. W. AAA+ proteins: have engine, will work. Nature Rev. Mol. Cell Biol. 6, 519–529 (2005).
Acknowledgements
M.L.M. acknowledges support from the National Science Foundation predoctoral fellows programme. Support for this work was provided to J.M.B. by the National Institutes of Health (GM071747).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
RCSB Protein Data Bank
FURTHER INFORMATION
Glossary
- Negatively supercoiled DNA
-
Supercoiled DNA with a negative average linking difference; DNA that is underwound with respect to a relaxed duplex.
- Replisome
-
A multi-protein complex assembled at oriC that contains all the enzymes required for DNA replication.
- Dimethylsulphate (DMS) footprinting
-
An assay that identifies a region of DNA that is involved in DNA–protein interactions. DMS methylates guanine nucleotides at the N7 position in the major groove. Protein binding protects this site from methylation, creating a characteristic pattern or footprint.
- AAA+ ATPases
-
A superfamily of proteins with one or two nucleotide-binding domains (AAA modules), which often form ring-like oligomers and function as molecular remodelling factors for diverse cellular processes.
- Positive DNA supercoils
-
Supercoiled DNA with a positive average linking difference; DNA that is overwound with respect to a relaxed duplex.
- Negative writhe
-
Writhe is the geometric parameter used to describe the axial paths of a DNA superhelix (two DNA duplexes) as they wrap around each other in space. To maintain a constant helical pitch (or twist), negatively supercoiled DNA often adopts a negative writhe.
- Positive torodial wrap
-
Some proteins can spatially bend or wrap DNA around their surface. A toroidal wrap refers to the bending of a DNA duplex into a near-circle, much like the spiral coiling of a telephone cord. A positive wrap indicates that the coils have a right-handed pitch.
- DNA sliding clamp
-
A doughnut-shaped protein complex that threads the DNA through its hole while tethering the polymerase to DNA.
- Cracked-ring particles
-
The vast majority of AAA+ proteins form ring-shaped hexamers. A few (such as the processivity clamp loaders) do not seal completely, leaving a notch or crack in the ring.
- Dam methyltransferase
-
An N6-adenine methyltransferase found in bacteria.
Rights and permissions
About this article
Cite this article
Mott, M., Berger, J. DNA replication initiation: mechanisms and regulation in bacteria. Nat Rev Microbiol 5, 343–354 (2007). https://doi.org/10.1038/nrmicro1640
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1640
This article is cited by
-
Escherichia coli cell factories with altered chromosomal replication scenarios exhibit accelerated growth and rapid biomass production
Microbial Cell Factories (2022)
-
Controllable DNA hybridization by host–guest complexation-mediated ligand invasion
Nature Communications (2022)
-
NapM enhances the survival of Mycobacterium tuberculosis under stress and in macrophages
Communications Biology (2019)
-
Elucidating functions of DP1 and DP2 subunits from the Thermococcus kodakarensis family D DNA polymerase
Extremophiles (2019)