Key Points
-
Divergent eukaryotic pathogens use common host-targeting signals in both animal and plant hosts. The first identification of a eukaryotic pathogen host-targeting signal was made in the human malaria parasite Plasmodium falciparum. This led to the first definition of the host-targeted 'secretome' of a eukaryotic pathogen. A functionally equivalent signal is conserved in the Irish potato famine pathogen Phytophthora infestans, showing that the host-targeting signal is shared in an apicomplexan and an oomycete.
-
The size and complexity of the predicted host-targeted secretome is significantly greater than that predicted for signal-mediated effectors in prokaryotic pathogens. The predicted involvement of hundreds of effectors indicates diverse mechanisms are involved in host remodelling by animal and plant pathogens. Analyses of common and contrasting domains in the effector secretomes reveal ABC transporters, post-translational or protein folding events in host remodelling, combined with expansions in specific gene families linked to virulence and evolution.
-
Analyses of the invasion mechanisms and secreted proteases of P. infestans and P. falciparum indicate a broader similarity in a wide range of pathogenicity mechanisms between animal and plant parasites than is currently appreciated.
-
Phylogenetic trees highlighting the major groups of animal and plant pathogens show that apicomplexans and oomycetes are divergent but more related to each other than to other pathogen groups, such as fungi, suggesting the need for a re-evaluation of their similarities, based on phylogeny and cellular similarities.
-
With the emerging genomics resources for eukaryotic parasites, it is important to appreciate that, like prokaryotes, similarities exist across divergent eukaryotic microorganisms and these could be exploited to elucidate new concepts in eukaryotic pathogenesis and potential new treatments for difficult infections. We are on the cusp of unique opportunities for the comparative genomics of eukaryotic microorganisms and these will be augmented by an understanding of common pathogenicity mechanisms in divergent species and an appreciation how such conserved mechanisms manifest themselves.
Abstract
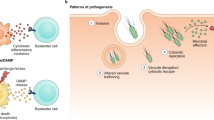
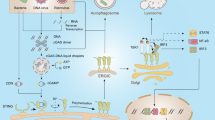
Pathogenic eukaryotes belong to several distinct phylogenetic lineages and have evolved the ability to colonize a range of hosts, including animals and plants. Pathogenic lifestyles have evolved repeatedly in eukaryotes, indicating that unique molecular processes are involved in host infection. However, evidence is now emerging that divergent eukaryotic pathogens might share common mechanisms of pathogenicity. The results from recent studies demonstrate that Plasmodium falciparum and Phytophthora infestans use equivalent host-targeting signals to deliver virulence adhesins and avirulence gene products into human and plant cells, respectively. Remodelling of host cells by different eukaryotic pathogens might therefore share some common features.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Preston, G. M., Studholme, D. J. & Caldelari, I. Profiling the secretomes of plant pathogenic Proteobacteria. FEMS Microbiol. Rev. 29, 331–360 (2005).
Christie, P. J. et al. Biogenesis, architecture, and function of bacterial type IV secretion systems. Annu. Rev. Microbiol. 59, 451–485 (2005).
Lammertyn, E. & Anne, J. Protein secretion in Legionella pneumophila and its relation to virulence. FEMS Microbiol. Lett. 238, 273–279 (2004).
Journet, L., Hughes, K. T. & Cornelis, G. R. Type III secretion: a secretory pathway serving both motility and virulence. Mol. Membr. Biol. 22, 41–50 (2005).
Miller, L. H. et al. The pathogenic basis of malaria. Nature 415, 673–679 (2002).
Haldar, K. et al. Protein and lipid trafficking induced in erythrocytes infected by malaria parasites. Cell. Microbiol. 4, 383–395 (2002).
Su, X.-Z. et al. The large diverse gene family var encodes proteins involved in cytoadherence and antigenic variation of Plasmodium falciparum-infected erythrocytes. Cell 82, 89–100 (1995).
Fernandez, V. et al. Small, clonally variant antigens expressed on the surface of the Plasmodium falciparum-infected erythrocyte are encoded by the rif gene family and are the target of human immune responses. J. Exp. Med. 190, 1393–1404 (1999).
Cooke, B. M. et al. Protein trafficking in Plasmodium falciparum-infected red blood cells. Trends Parasitol. 20, 581–589 (2004).
Baldauf, S. L. et al. A kingdom-level phylogeny of eukaryotes based on combined protein data. Science 290, 972–977 (2000).
Kamoun, S. Molecular genetics of pathogenic oomycetes. Euk. Cell 2, 191–199 (2003).
Huitema, E. et al. Linking sequence to phenotype in Phytophthora-plant interactions. Trends Microbiol. 12, 193–200 (2004).
Ellis, J., Catanzariti, A. M. & Dodds, P. The problem of how fungal and oomycete avirulence proteins enter plant cells. Trends Plant Sci. 11, 61–63 (2006).
Kamoun, S. A catalogue of the effector secretome of plant pathogenic oomycetes. Annu. Rev. Phytopathol. 44, 41–60 (2006). This review highlights recent findings on the structure of oomycete effectors, such as the RxLR effectors.
Birch, P. R. et al. Trafficking arms: oomycete effectors enter host plant cells. Trends Microbiol. 14, 8–11 (2006).
Allen, R. L. et al. Host-parasite coevolutionary conflict between Arabidopsis and downy mildew. Science 306, 1957–1960 (2004). The cloning of Hyaloperonospora parasitica ATR1 along with other oomycete avirulence genes resulted in the discovery of the conserved RxLR motif.
Rehmany, A. P. et al. Differential recognition of highly divergent downy mildew avirulence gene alleles by RPP1 resistance genes from two Arabidopsis lines. Plant Cell 17, 1839–1850 (2005).
Armstrong, M. R. et al. An ancestral oomycete locus contains late blight avirulence gene Avr3a, encoding a protein that is recognized in the host cytoplasm. Proc. Natl Acad. Sci. USA 102, 7766–7771 (2005).
Lopez-Estrano, C. et al. Cooperative domains define a unique host cell-targeting signal in Plasmodium falciparum-infected erythrocytes. Proc. Natl Acad. Sci. USA 100, 12402–12407 (2003). This study first defined minimal vacuolar translocation sequences (VTS) of about 35–40 amino acids necessary and sufficient for the export of P. falciparum proteins to the host erythrocyte.
Hiller, N. L. et al. A host-targeting signal in virulence proteins reveals a secretome in malarial infection. Science 306, 1934–1937 (2004). This study along with Ref. 21 simultaneously described a host-cell-targeting signal present in hundreds of parasite secretory proteins and thereby predicted a 'secretome' for malarial infection.
Marti, M. et al. Targeting malaria virulence and remodeling proteins to the host erythrocyte. Science 306, 1930–1933 (2004).
Schwarze, S. R. et al. In vivo protein transduction: delivery of a biologically active protein into the mouse. Science 285, 1569–1572 (1999).
Muller, M. & Klosgen, R. B. The Tat pathway in bacteria and chloroplasts. Mol. Membr. Biol. 22, 113–121 (2005).
Sargeant, T. J. et al. Lineage-specific expansion of proteins exported to erythrocytes in malaria parasites. Genome Biol. 7, R12 (2006).
Bos, J. I. et al. The C-terminal half of Phytophthora infestans RXLR effector AVR3a is sufficient to trigger R3a-mediated hypersensitivity and suppress INF1-induced cell death in Nicotiana benthamiana. Plant J. 48, 165–176 (2006).
Bhattacharjee, S. et al. The malarial host-targeting signal is conserved in the Irish potato famine pathogen. PLoS Pathog. 2, e50 (2006). This study established the molecular equivalence between host-targeting signals of Plasmodium and Phytopthora species.
Neduva, V. & Russell, R. B. Linear motifs: evolutionary interaction switches. FEBS Lett. 579, 3342–3345 (2005).
Ward, P. et al. Protein kinases of the human malaria parasite Plasmodium falciparum: the kinome of a divergent eukaryote. BMC Genomics 5, 79 (2004).
Sam-Yellowe, T. Y. et al. A Plasmodium gene family encoding Maurer's cleft membrane proteins: structural properties and expression profiling. Genome Res. 14, 1052–1059 (2004).
Schneider, A. G. & Mercereau-Puijalon, O. A new Apicomplexa-specific protein kinase family: multiple members in Plasmodium falciparum, all with an export signature. BMC Genomics 6, 30 (2005).
Janssen, C. S. et al. Plasmodium interspersed repeats: the major multigene superfamily of malaria parasites. Nucleic Acids Res. 32, 5712–5720 (2004).
Kooij, T. W. et al. A Plasmodium whole-genome synteny map: indels and synteny breakpoints as foci for species-specific genes. PLoS Pathog. 1, e44 (2005).
Brunner, F. et al. Pep-13, a plant defense-inducing pathogen-associated pattern from Phytophthora transglutaminases. EMBO J. 21, 6681–6688 (2002).
Sibley, L. D. Intracellular parasite invasion strategies. Science 304, 248–253 (2004). This is an excellent review for apicomplexan invasion strategies.
Judelson, H. S. & Blanco, F. A. The spores of Phytophthora: weapons of the plant destroyer. Nature Rev. Microbiol. 3, 47–58 (2005).
Robold, A. V. & Hardham, A. R. During attachment Phytophthora spores secrete proteins containing thrombospondin type 1 repeats. Curr. Genet. 47, 307–315 (2005). Shows that PcVsv1, a 260-kDa protein secreted by Phytophthora spores is similar to apicomplexan adhesins.
Deng, M. et al. Cryptosporidium parvum genes containing thrombospondin type 1 domains. Infect. Immun. 70, 6987–6995 (2002).
Kappe, S. H., Buscaglia, C. A. & Nussenzweig, V. Plasmodium sporozoite molecular cell biology. Annu. Rev. Cell. Dev. Biol. 20, 29–59 (2004).
Kappe, S. et al. Conservation of a gliding motility and cell invasion machinery in Apicomplexan parasites. J. Cell Biol. 147, 937–944 (1999).
Sultan, A. A. et al. TRAP is necessary for gliding motility and infectivity of Plasmodium sporozoites. Cell 90, 511–522 (1997).
Menard, R. Gliding motility and cell invasion by Apicomplexa: insights from the Plasmodium sporozoite. Cell. Microbiol. 3, 63–73 (2001).
Baum, J. et al. A conserved molecular motor drives cell invasion and gliding motility across malaria life cycle stages and other apicomplexan parasites. J. Biol. Chem. 281, 5197–5208 (2006).
Brecht, S. et al. The Toxoplasma micronemal protein MIC4 is an adhesin composed of six conserved apple domains. J. Biol. Chem. 276, 4119–4127 (2001).
Brown, P. J. et al. A microneme protein from Eimeria tenella with homology to the Apple domains of coagulation factor XI and plasma pre-kallikrein. Mol. Biochem. Parasitol. 107, 91–102 (2000).
Brown, P.J. et al. Domains of invasion organelle proteins from apicomplexan parasites are homologous with the Apple domains of blood coagulation factor XI and plasma pre-kallikrein and are members of the PAN module superfamily. FEBS Lett. 497, 31–38 (2001).
Pizarro, J.C. et al. Crystal structure of the malaria vaccine candidate apical membrane antigen 1. Science 308, 408–411 (2005). This is an important paper for the development of a malaria vaccine.
Mital, J. et al. Conditional expression of Toxoplasma gondii apical membrane antigen-1 (TgAMA1) demonstrates that TgAMA1 plays a critical role in host cell invasion. Mol. Biol. Cell 16, 4341–4349 (2005).
Mullen, G. E. et al. Enhancement of functional antibody responses to AMA1-C1/Alhydrogel, a Plasmodium falciparum malaria vaccine, with CpG oligodeoxynucleotide. Vaccine 24, 2497–2505 (2006).
Gaulin, E. et al. The CBEL glycoprotein of Phytophthora parasitica var. nicotianae is involved in cell wall deposition and adhesion to cellulosic substrates. J. Cell Sci. 115, 4565–4575 (2002).
Torto, T. A. et al. EST mining and functional expression assays identify extracellular effector proteins from the plant pathogen Phytophthora. Genome Res. 13, 1675–1685 (2003).
Pszenny, V. et al. Molecular cloning, sequencing and expression of a serine proteinase inhibitor gene from Toxoplasma gondii. Mol. Biochem. Parasitol. 107, 241–249 (2000).
Lindh, J. G. et al. A protease inhibitor associated with the surface of Toxoplasma gondii. Mol. Biochem. Parasitol. 116, 137–145 (2001).
Morris, M. T. et al. Neospora caninum expresses an unusual single-domain Kazal protease inhibitor that is discharged into the parasitophorous vacuole. Int. J. Parasitol. 34, 693–701 (2004).
Bruno, S. et al. Identification and characterization of serine proteinase inhibitors from Neospora caninum. Mol. Biochem. Parasitol. 136, 101–107 (2004).
Abrahamsen, M. S. et al. Complete genome sequence of the apicomplexan, Cryptosporidium parvum. Science 304, 441–445 (2004).
Tian, M. et al. A Kazal-like extracellular serine protease inhibitor from Phytophthora infestans targets the tomato pathogenesis-related protease P69B. J. Biol. Chem. 279, 26370–26377 (2004). In this first report of a protease inhibitor from a plant pathogenic microorganism, the similarity with animal parasites is highlighted.
Santos, C. C. et al. Chagasin, the endogenous cysteine-protease inhibitor of Trypanosoma cruzi, modulates parasite differentiation and invasion of mammalian cells. J. Cell Sci. 118, 901–915 (2005).
Besteiro, S., Coombs, G. H. & Mottram, J. C. A potential role for ICP, a leishmanial inhibitor of cysteine peptidases, in the interaction between host and parasite. Mol. Microbiol. 54, 1224–1236 (2004).
Rigden, D.J., Mosolov, V. V. & Galperin, M. Y. Sequence conservation in the chagasin family suggests a common trend in cysteine proteinase binding by unrelated protein inhibitors. Protein Sci. 11, 1971–1977 (2002).
Torto-Alalibo, T. et al. Expressed sequence tags from the oomycete fish pathogen Saprolegnia parasitica reveal putative virulence factors. BMC Microbiol. 5, 46 (2005).
Win, J. et al. Computational and comparative analyses of 150 full-length cDNA sequences from the oomycete plant pathogen Phytophthora infestans. Fungal Genet. Biol. 43, 20–33 (2006).
Orbach, M. J. et al. A telomeric avirulence gene determines efficacy for the rice blast resistance gene Pi-ta. Plant Cell 12, 2019–2032 (2000).
Dodds, P. N. et al. The Melampsora lini AvrL567 avirulence genes are expressed in haustoria and their products are recognized inside plant cells. Plant Cell 16, 755–768 (2004).
Kemen, E. et al. Identification of a protein from rust fungi transferred from haustoria into infected plant cells. Mol. Plant-Microbe Interact. 18, 1130–1139 (2005).
Embley, T. M. & Martin, W. Eukaryotic evolution, changes and challenges. Nature 440, 623–630 (2006).
Acknowledgements
This work was supported by grants from the National Institutes of Health (K.H.), American Heart Foundation (NLH) and the National Science Foundation (S.K.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Related links
DATABASES
Entrez Genome Project
FURTHER INFORMATION
Glossary
- Avirulence protein
-
An effector protein of plant pathogens that triggers disease resistance.
- Parasitophorous vacuole
-
A vacuole within the host cell in which the parasite resides.
- Maurer's clefts
-
Single-membrane-limited structures in the erythrocyte cytoplasm.
- Biotrophic
-
A biotrophic pathogen invades host tissue without killing host cells and feeds on living cells.
- Sporangium
-
A sac-like structure that is capable of converting its cytoplasm into multiple spores.
- Zoospore
-
A flagellated, wall-less spore, specialized for dispersal.
- Isoflavones
-
Specialized polyphenolic metabolites produced by plants of the legume family.
- Appressoria
-
Specialized infection structures of oomycete and fungal plant pathogens that enable host penetration.
- Hypersensitive response
-
A programmed cell-death response of plants that is associated with resistance.
- General secretory pathway
-
Pathway for protein secretion through the endoplasmic reticulum and Golgi to the plasma membrane.
- Twin-arginine transport
-
(TAT). A protein transport system that transports proteins across membranes in a fully folded state.
- Syntenous orthologue
-
An orthologue in which the gene locus is conserved among species.
- Micronemes
-
Micronemes are located at the apical end of all three invasive stages of apicomplexan parasites and have an important role in gliding motility, host-cell adhesion and invasion.
- Rhoptries
-
Rhoptries are located at the apical end of certain invasive stages of apicomplexan parasites, and have an important role in host-cell adhesion and invasion, and in the establishment of the parasitophorous vacuole.
- Dense granules
-
Secretory granules, the contents of which are released from the parasites after discharge from the rhoptries.
- Gliding motility
-
A form of substrate-dependent locomotion in which the parasite maintains a fixed shape.
- Haustoria
-
Specialized infection structures of biotrophic oomycete and fungal plant pathogens that invaginate into plant cells but remain enveloped by a modified host-cell membrane.
Rights and permissions
About this article
Cite this article
Haldar, K., Kamoun, S., Hiller, N. et al. Common infection strategies of pathogenic eukaryotes. Nat Rev Microbiol 4, 922–931 (2006). https://doi.org/10.1038/nrmicro1549
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro1549
This article is cited by
-
Breeding of Genetically Engineered Indonesian Potato, Bio Granola, Resistant to Late Blight Pathogen Phytophthora infestans
Potato Research (2024)
-
Durable Late Blight Resistance in Potato Through Dynamic Varieties Obtained by Cisgenesis: Scientific and Societal Advances in the DuRPh Project
Potato Research (2016)
-
Oomycete pathogens encode RNA silencing suppressors
Nature Genetics (2013)
-
Genome evolution in filamentous plant pathogens: why bigger can be better
Nature Reviews Microbiology (2012)
-
Computational models in plant-pathogen interactions: the case of Phytophthora infestans
Theoretical Biology and Medical Modelling (2009)