Abstract
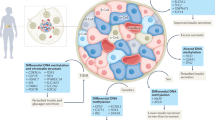
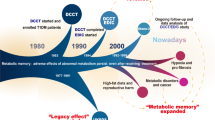
Diabetes mellitus (type 1 and type 2) and the complications associated with this condition are an urgent public health problem, as the incidence of diabetes mellitus is steadily increasing. Environmental factors, such as diet and exposure to hyperglycemia, contribute to the etiology of diabetes mellitus and its associated microvascular and macrovascular complications. These vascular complications are the main cause of the morbidity and mortality burden of diabetes mellitus. The DCCT–EDIC and UKPDS epidemiological studies correlated poor glycemic control with the development of vascular complications in patients with type 1 or type 2 diabetes mellitus. The findings of these studies suggest that early exposure to hyperglycemia predisposes individuals to the development of diabetic complications, a phenomenon referred to as metabolic memory or the legacy effect. The first experimental evidence for metabolic memory was reported >20 years ago and the underlying molecular mechanisms are currently being characterized. Interestingly, transient exposure to hyperglycemia results in long-lasting epigenetic modifications that lead to changes in chromatin structure and gene expression, which mediate these persistent metabolic characteristics.
Key Points
-
In patients with type 1 or type 2 diabetes mellitus, the clinical complications underlying morbidity and mortality have a microvascular or macrovascular nature
-
Vascular complications in patients with diabetes mellitus are primarily, although not solely, caused by hyperglycemia
-
The results of prospective, large-scale clinical trials suggest that diabetic complications attributable to poorly controlled glycemia can manifest long after initiation of an improved glycemic control regimen
-
Studies in animal models and cell cultures have demonstrated that clinical and biological responses to exposure to hyperglycemia persist even after return to normoglycemic conditions
-
In endothelial cells, hyperglycemia induces generation of reactive oxygen species and activates both the exosamine pathway and protein kinase C; these events upregulate transcription of proinflammatory mediators
-
In patients with diabetes mellitus, hyperglycemia induces long-lasting, post-translational modifications in histones that contribute (after return to normoglycemia) to the sustained expression of proinflammatory mediators involved in vascular dysfunction
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Sheetz, M. J. & King, G. L. Molecular understanding of hyperglycemia's adverse effects for diabetic complications. JAMA 288, 2579–2588 (2002).
McCarthy, M. I. & Froguel, P. Genetic approaches to the molecular understanding of type 2 diabetes. Am. J. Physiol. Endocrinol. Metab. 283, E217–E225 (2002).
Nesto, R. W. Beyond low-density lipoprotein: addressing the atherogenic lipid triad in type 2 diabetes mellitus and the metabolic syndrome. Am. J. Cardiovasc. Drugs 5, 379–387 (2005).
Mourad, J. J. & Le Jeune, S. Blood pressure control, risk factors and cardiovascular prognosis in patients with diabetes: 30 years of progress. J. Hypertens. 26 (Suppl. 3), S7–S13 (2008).
Brownlee, M. Biochemistry and molecular cell biology of diabetic complications. Nature 414, 813–820 (2001).
Forbes, J. M., Fukami, K. & Cooper, M. E. Diabetic nephropathy: where hemodynamics meets metabolism. Exp. Clin. Endocrinol. Diabetes 115, 69–84 (2007).
[No authors listed] The effect of intensive treatment of diabetes on the development and progression of long-term complications in insulin-dependent diabetes mellitus. The Diabetes Control and Complications Trial Research Group. N. Engl. J. Med. 329, 977–986 (1993).
[No authors listed] UK Prospective Diabetes Study (UKPDS). VIII. Study design, progress and performance. Diabetologia 34, 877–890 (1991).
Writing team for the Diabetes Control and Complications Trial/Epidemiology Interventions and Complications Research Group. Sustained effect of intensive treatment of type 1 diabetes mellitus on development and progression of diabetic nephropathy: the Epidemiology of Diabetes Interventions and Complications (EDIC) study. JAMA 290, 2159–2167 (2003).
Nathan, D. M. et al. Intensive diabetes treatment and cardiovascular disease in patients with type 1 diabetes. N. Engl. J. Med. 353, 2643–2653 (2005).
Nathan, D. M. et al. Modern-day clinical course of type 1 diabetes mellitus after 30 years' duration: the diabetes control and complications trial/epidemiology of diabetes interventions and complications and Pittsburgh epidemiology of diabetes complications experience (1983–2005). Arch. Intern. Med. 169, 1307–1316 (2009).
Albers, J. W. et al. Effect of prior intensive insulin treatment during the Diabetes Control and Complications Trial (DCCT) on peripheral neuropathy in type 1 diabetes during the Epidemiology of Diabetes Interventions and Complications (EDIC) Study. Diabetes Care 33, 1090–1096 (2010).
[No authors listed] Intensive blood-glucose control with sulphonylureas or insulin compared with conventional treatment and risk of complications in patients with type 2 diabetes (UKPDS 33). UK Prospective Diabetes Study (UKPDS) Group. Lancet 352, 837–853 (1998).
Holman, R. R., Paul, S. K., Bethel, M. A., Matthews, D. R. & Neil, H. A. 10-year follow-up of intensive glucose control in type 2 diabetes. N. Engl. J. Med. 359, 1577–1589 (2008).
Chalmers, J. & Cooper, M. E. UKPDS and the legacy effect. N. Engl. J. Med. 359, 1618–1620 (2008).
Marshall, S. M. & Flyvbjerg, A. Prevention and early detection of vascular complications of diabetes. BMJ 333, 475–480 (2006).
Patel, A. et al. Intensive blood glucose control and vascular outcomes in patients with type 2 diabetes. N. Engl. J. Med. 358, 2560–2572 (2008).
Gerstein, H. C. et al. Effects of intensive glucose lowering in type 2 diabetes. N. Engl. J. Med. 358, 2545–2559 (2008).
Wong, T. Y. et al. Relation between fasting glucose and retinopathy for diagnosis of diabetes: three population-based cross-sectional studies. Lancet 371, 736–743 (2008).
Writing team for the Diabetes Control and Complications Trial/Epidemiology of Diabetes Interventions and Complications Research Group. Effect of intensive therapy on the microvascular complications of type 1 diabetes mellitus. JAMA 287, 2563–2569 (2002).
Ling, C. & Groop, L. Epigenetics: a molecular link between environmental factors and type 2 diabetes. Diabetes 58, 2718–2725 (2009).
Lyssenko, V. et al. Clinical risk factors, DNA variants, and the development of type 2 diabetes. N. Engl. J. Med. 359, 2220–2232 (2008).
van Hoek, M. et al. Predicting type 2 diabetes based on polymorphisms from genome-wide association studies: a population-based study. Diabetes 57, 3122–3128 (2008).
El-Osta, A. et al. Transient high glucose causes persistent epigenetic changes and altered gene expression during subsequent normoglycemia. J. Exp. Med. 205, 2409–2417 (2008).
Engerman, R. L. & Kern, T. S. Progression of incipient diabetic retinopathy during good glycemic control. Diabetes 36, 808–812 (1987).
Kowluru, R. A. Effect of reinstitution of good glycemic control on retinal oxidative stress and nitrative stress in diabetic rats. Diabetes 52, 818–823 (2003).
Hammes, H. P. et al. Islet transplantation inhibits diabetic retinopathy in the sucrose-fed diabetic Cohen rat. Invest. Ophthalmol. Vis. Sci. 34, 2092–2096 (1993).
Brasacchio, D. et al. Hyperglycemia induces a dynamic cooperativity of histone methylase and demethylase enzymes associated with gene-activating epigenetic marks that coexist on the lysine tail. Diabetes 58, 1229–1236 (2009).
Roy, S., Sala, R., Cagliero, E. & Lorenzi, M. Overexpression of fibronectin induced by diabetes or high glucose: phenomenon with a memory. Proc. Natl Acad. Sci. USA 87, 404–408 (1990).
Ihnat, M. A. et al. Reactive oxygen species mediate a cellular 'memory' of high glucose stress signalling. Diabetologia 50, 1523–1531 (2007).
Corgnali, M., Piconi, L., Ihnat, M. & Ceriello, A. Evaluation of gliclazide ability to attenuate the hyperglycaemic 'memory' induced by high glucose in isolated human endothelial cells. Diabetes Metab. Res. Rev. 24, 301–309 (2008).
Ceriello, A., Ihnat, M. A. & Thorpe, J. E. Clinical review 2: The “metabolic memory”: is more than just tight glucose control necessary to prevent diabetic complications? J. Clin. Endocrinol. Metab. 94, 410–415 (2009).
El-Osta, A. DNMT cooperativity--the developing links between methylation, chromatin structure and cancer. Bioessays 25, 1071–1084 (2003).
Ruthenburg, A. J., Allis, C. D. & Wysocka, J. Methylation of lysine 4 on histone H3: intricacy of writing and reading a single epigenetic mark. Mol. Cell 25, 15–30 (2007).
Turner, B. M. Cellular memory and the histone code. Cell 111, 285–291 (2002).
Feng, Q. et al. Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr. Biol. 12, 1052–1058 (2002).
Wang, H. et al. Purification and functional characterization of a histone H3-lysine 4-specific methyltransferase. Mol. Cell 8, 1207–1217 (2001).
Nishioka, K. et al. Set9, a novel histone H3 methyltransferase that facilitates transcription by precluding histone tail modifications required for heterochromatin formation. Genes Dev. 16, 479–489 (2002).
Li, Y. et al. Role of the histone H3 lysine 4 methyltransferase, SET7/9, in the regulation of NF-κB-dependent inflammatory genes. Relevance to diabetes and inflammation. J. Biol. Chem. 283, 26771–26781 (2008).
Deering, T. G., Ogihara, T., Trace, A. P., Maier, B. & Mirmira, R. G. Methyltransferase Set7/9 maintains transcription and euchromatin structure at islet-enriched genes. Diabetes 58, 185–193 (2009).
Villeneuve, L. M. et al. Epigenetic histone H3 lysine 9 methylation in metabolic memory and inflammatory phenotype of vascular smooth muscle cells in diabetes. Proc. Natl Acad. Sci. USA 105, 9047–9052 (2008).
Kurash, J. K. et al. Methylation of p53 by Set7/9 mediates p53 acetylation and activity in vivo. Mol. Cell 29, 392–400 (2008).
Ea, C. K. & Baltimore, D. Regulation of NF-κB activity through lysine monomethylation of p65. Proc. Natl Acad. Sci. USA 106, 18972–18977 (2009).
Estève, P. O. et al. Regulation of DNMT1 stability through SET7-mediated lysine methylation in mammalian cells. Proc. Natl Acad. Sci. USA 106, 5076–5081 (2009).
Pradhan, S., Chin, H. G., Estève, P. O. & Jacobsen, S. E. SET7/9 mediated methylation of non-histone proteins in mammalian cells. Epigenetics 4, 383–387 (2009).
Miao, F., Li, S., Chavez, V., Lanting, L. & Natarajan, R. Coactivator-associated arginine methyltransferase-1 enhances nuclear factor-kappaB-mediated gene transcription through methylation of histone H3 at arginine 17. Mol. Endocrinol. 20, 1562–1573 (2006).
Wang, J. et al. The lysine demethylase LSD1 (KDM1) is required for maintenance of global DNA methylation. Nat. Genet. 41, 125–129 (2009).
Hotamisligil, G. S. & Spiegelman, B. M. Tumor necrosis factor alpha: a key component of the obesity-diabetes link. Diabetes 43, 1271–1278 (1994).
Nishikawa, T. et al. Normalizing mitochondrial superoxide production blocks three pathways of hyperglycaemic damage. Nature 404, 787–790 (2000).
Miao, F., Gonzalo, I. G., Lanting, L. & Natarajan, R. In vivo chromatin remodeling events leading to inflammatory gene transcription under diabetic conditions. J. Biol. Chem. 279, 18091–18097 (2004).
Xiang, K. et al. Genome-wide search for type 2 diabetes/impaired glucose homeostasis susceptibility genes in the Chinese: significant linkage to chromosome 6q21-q23 and chromosome 1q21-q24. Diabetes 53, 228–234 (2004).
van Tilburg, J. H. et al. A genome-wide scan in type 2 diabetes mellitus provides independent replication of a susceptibility locus on 18p11 and suggests the existence of novel loci on 2q12 and 19q13. J. Clin. Endocrinol. Metab. 88, 2223–2230 (2003).
Gray, S. G. & De Meyts, P. Role of histone and transcription factor acetylation in diabetes pathogenesis. Diabetes Metab. Res. Rev. 21, 416–433 (2005).
Garber, K. Purchase of Aton spotlights HDAC inhibitors. Nat. Biotechnol. 22, 364–365 (2004).
Guarente, L. Sirtuins as potential targets for metabolic syndrome. Nature 444, 868–874 (2006).
Miao, F. et al. Genome-wide analysis of histone lysine methylation variations caused by diabetic conditions in human monocytes. J. Biol. Chem. 282, 13854–13863 (2007).
Nowak, S. J. & Corces, V. G. Phosphorylation of histone H3: a balancing act between chromosome condensation and transcriptional activation. Trends Genet. 20, 214–220 (2004).
Weake, V. M. & Workman, J. L. Histone ubiquitination: triggering gene activity. Mol. Cell 29, 653–663 (2008).
Shiio, Y. & Eisenman, R. N. Histone sumoylation is associated with transcriptional repression. Proc. Natl Acad. Sci. USA 100, 13225–13230 (2003).
Ouyang, J. & Gill, G. SUMO engages multiple corepressors to regulate chromatin structure and transcription. Epigenetics 4, 440–444 (2009).
Smith, B. C. & Denu, J. M. Chemical mechanisms of histone lysine and arginine modifications. Biochim. Biophys. Acta 1789, 45–57 (2009).
An, W. Histone acetylation and methylation: combinatorial players for transcriptional regulation. Subcell. Biochem. 41, 351–369 (2007).
Imhof, A. et al. Acetylation of general transcription factors by histone acetyltransferases. Curr. Biol. 7, 689–692 (1997).
Haberland, M., Montgomery, R. L. & Olson, E. N. The many roles of histone deacetylases in development and physiology: implications for disease and therapy. Nat. Rev. Genet. 10, 32–42 (2009).
Krause, B., Sobrevia, L. & Casanello, P. Epigenetics: new concepts of old phenomena in vascular physiology. Curr. Vasc. Pharmacol. 7, 513–520 (2009).
Chen, S. et al. Transcriptional coactivator p300 regulates glucose induced gene expression in the endothelial cells. Am. J. Physiol. Endocrinol. Metab. 298, E127–E137 (2010).
Kabra, D. G., Gupta, J. & Tikoo, K. Insulin induced alteration in post-translational modifications of histone H3 under a hyperglycemic condition in L6 skeletal muscle myoblasts. Biochim. Biophys. Acta 1792, 574–583 (2009).
Mosley, A. L. & Ozcan, S. Glucose regulates insulin gene transcription by hyperacetylation of histone h4. J. Biol. Chem. 278, 19660–19666 (2003).
Mosley, A. L., Corbett, J. A. & Ozcan, S. Glucose regulation of insulin gene expression requires the recruitment of p300 by the beta-cell-specific transcription factor Pdx-1. Mol. Endocrinol. 18, 2279–2290 (2004).
Mosley, A. L. & Ozcan, S. The pancreatic duodenal homeobox-1 protein (Pdx-1) interacts with histone deacetylases Hdac-1 and Hdac-2 on low levels of glucose. J. Biol. Chem. 279, 54241–54247 (2004).
El-Assaad, W. et al. Glucolipotoxicity alters lipid partitioning and causes mitochondrial dysfunction, cholesterol, and ceramide deposition and reactive oxygen species production in INS832/13 ss-cells. Endocrinology 151, 3061–3073 (2010).
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009).
Liu, L., Li, Y. & Tollefsbol, T. O. Gene-environment interactions and epigenetic basis of human diseases. Curr. Issues Mol. Biol. 10, 25–36 (2008).
Reik, W., Dean, W. & Walter, J. Epigenetic reprogramming in mammalian development. Science 293, 1089–1093 (2001).
Maier, S. & Olek, A. Diabetes: a candidate disease for efficient DNA methylation profiling. J. Nutr. 132 (Suppl.), 2440S–2443S (2002).
Poirier, L. A. et al. Blood S-adenosylmethionine concentrations and lymphocyte methylenetetrahydrofolate reductase activity in diabetes mellitus and diabetic nephropathy. Metabolism 50, 1014–1018 (2001).
Thompson, R. F. et al. Experimental intrauterine growth restriction induces alterations in DNA methylation and gene expression in pancreatic islets of rats. J. Biol. Chem. 285, 15111–15118 (2010).
Einstein, F. et al. Cytosine methylation dysregulation in neonates following intrauterine growth restriction. PLoS ONE 5, e8887 (2010).
Orlando, V. & Paro, R. Mapping Polycomb-repressed domains in the bithorax complex using in vivo formaldehyde cross-linked chromatin. Cell 75, 1187–1198 (1993).
Sekinger, E. A., Moqtaderi, Z. & Struhl, K. Intrinsic histone-DNA interactions and low nucleosome density are important for preferential accessibility of promoter regions in yeast. Mol. Cell 18, 735–748 (2005).
Schones, D. E. & Zhao, K. Genome-wide approaches to studying chromatin modifications. Nat. Rev. Genet. 9, 179–191 (2008).
Ceriello, A., Esposito, K., Ihnat, M., Thorpe, J. & Giugliano, D. Long-term glycemic control influences the long-lasting effect of hyperglycemia on endothelial function in type 1 diabetes. J. Clin. Endocrinol. Metab. 94, 2751–2756 (2009).
Park, P. J. ChIP-seq: advantages and challenges of a maturing technology. Nat. Rev. Genet. 10, 669–680 (2009).
Lindström, J. et al. Take action to prevent diabetes--the IMAGE toolkit for the prevention of type 2 diabetes in Europe. Horm. Metab. Res. 42 (Suppl. 1), S37–S55 (2010).
Paulweber, B. et al. A European evidence-based guideline for the prevention of type 2 diabetes. Horm. Metab. Res. 42 (Suppl. 1), S3–S36 (2010).
Schwarz, P. E., Schwarz, J., Schuppenies, A., Bornstein, S. R. & Schulze, J. Development of a diabetes prevention management program for clinical practice. Public Health Rep. 122, 258–263 (2007).
Rothe, U. et al. Evaluation of a diabetes management system based on practice guidelines, integrated care, and continuous quality management in a Federal State of Germany: a population-based approach to health care research. Diabetes Care 31, 863–868 (2008).
Acknowledgements
We acknowledge grant and fellowship support from the Juvenile Research Diabetes Foundation, Diabetes Australia Research Trust, the National Health and Medical Research Council of Australia and the Franco-Australian INSERM-NHMRC cooperation program. A. Balcerczyk was supported by the PSF (Polish Science Foundation).
Author information
Authors and Affiliations
Contributions
All authors contributed equally to researching data for the article, discussions of the content, writing the article and to reviewing and editing the manuscript before submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Pirola, L., Balcerczyk, A., Okabe, J. et al. Epigenetic phenomena linked to diabetic complications. Nat Rev Endocrinol 6, 665–675 (2010). https://doi.org/10.1038/nrendo.2010.188
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrendo.2010.188
This article is cited by
-
Simultaneous attenuation of hyperglycemic memory-induced retinal, pulmonary, and glomerular dysfunctions by proinsulin C-peptide in diabetes
BMC Medicine (2023)
-
Target amplification-free detection of urinary microRNA for diabetic nephropathy diagnosis with electrocatalytic reaction
Analytical and Bioanalytical Chemistry (2022)
-
miRNA signatures in diabetic retinopathy and nephropathy: delineating underlying mechanisms
Journal of Physiology and Biochemistry (2022)
-
Polymer-Based Delivery of Peptide Drugs to Treat Diabetes: Normalizing Hyperglycemia and Preventing Diabetic Complications
BioChip Journal (2022)
-
Increased PARylation impacts the DNA methylation process in type 2 diabetes mellitus
Clinical Epigenetics (2021)