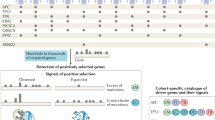
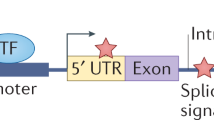
Abstract
Much of cancer genetics research has focused on the identification of the most-important somatic mutations ('major drivers') that cause tumour growth. However, many mutations found in cancer might not be major drivers or 'passenger' mutations, but instead might have relatively weak tumour-promoting effects. Our aim is to highlight the existence of these mutations (termed 'mini drivers' herein), as multiple mini-driver mutations might substitute for a major-driver change, especially in the presence of genomic instability or high mutagen exposure. The mini-driver model has clinical implications: for example, the effects of therapeutically targeting such genes may be limited. However, the main importance of the model lies in helping to provide a complete understanding of tumorigenesis, especially as we anticipate that an increasing number of mini-driver mutations will be found by cancer genome sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Garraway, L. A. & Lander, E. S. Lessons from the cancer genome. Cell 153, 17–37 (2013).
Leiserson, M. D. et al. Pan-cancer network analysis identifies combinations of rare somatic mutations across pathways and protein complexes. Nat. Genet. 47, 106–114 (2015).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Stratton, M. R., Campbell, P. J. & Futreal, P. A. The cancer genome. Nature 458, 719–724 (2009).
Marx, V. Cancer genomes: discerning drivers from passengers. Nat. Methods 11, 375–379 (2014).
Lawrence, M. S. et al. Discovery and saturation analysis of cancer genes across 21 tumour types. Nature 505, 495–501 (2014).
Greaves, M. & Maley, C. C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
Merlo, L. M., Pepper, J. W., Reid, B. J. & Maley, C. C. Cancer as an evolutionary and ecological process. Nat. Rev. Cancer 6, 924–935 (2006).
Amos-Landgraf, J. M. et al. Sex disparity in colonic adenomagenesis involves promotion by male hormones, not protection by female hormones. Proc. Natl Acad. Sci. USA 111, 16514–16519 (2014).
Figueiredo, J. C. et al. Genome-wide diet–gene interaction analyses for risk of colorectal cancer. PLoS Genet. 10, e1004228 (2014).
Beerenwinkel, N. et al. Genetic progression and the waiting time to cancer. PLoS Comput. Biol. 3, e225 (2007).
Bansal, V., Libiger, O., Torkamani, A. & Schork, N. J. Statistical analysis strategies for association studies involving rare variants. Nat. Rev. Genet. 11, 773–785 (2010).
Bodmer, W. & Bonilla, C. Common and rare variants in multifactorial susceptibility to common diseases. Nat. Genet. 40, 695–701 (2008).
Hindorff, L. A., Gillanders, E. M. & Manolio, T. A. Genetic architecture of cancer and other complex diseases: lessons learned and future directions. Carcinogenesis 32, 945–954 (2011).
Cheetham, S. W., Gruhl, F., Mattick, J. S. & Dinger, M. E. Long noncoding RNAs and the genetics of cancer. Br. J. Cancer 108, 2419–2425 (2013).
Fredriksson, N. J., Ny, L., Nilsson, J. A. & Larsson, E. Systematic analysis of noncoding somatic mutations and gene expression alterations across 14 tumor types. Nat. Genet. 46, 1258–1263 (2014).
Hayes, J., Peruzzi, P. P. & Lawler, S. MicroRNAs in cancer: biomarkers, functions and therapy. Trends Mol. Med. 20, 460–469 (2014).
Horn, S. et al. TERT promoter mutations in familial and sporadic melanoma. Science 339, 959–961 (2013).
Huang, F. W. et al. Highly recurrent TERT promoter mutations in human melanoma. Science 339, 957–959 (2013).
Weinhold, N., Jacobsen, A., Schultz, N., Sander, C. & Lee, W. Genome-wide analysis of noncoding regulatory mutations in cancer. Nat. Genet. 46, 1160–1165 (2014).
Zeron-Medina, J. et al. A polymorphic p53 response element in KIT ligand influences cancer risk and has undergone natural selection. Cell 155, 410–422 (2013).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
Imamura, Y. et al. Analyses of clinicopathological, molecular, and prognostic associations of KRAS codon 61 and codon 146 mutations in colorectal cancer: cohort study and literature review. Mol. Cancer 13, 135 (2014).
Sansom, O. Tissue-specific tumour suppression by APC. Adv. Exp. Med. Biol. 656, 107–118 (2009).
The Cancer Genome Atlas Network. Comprehensive molecular characterization of human colon and rectal cancer. Nature 487, 330–337 (2012).
Lamlum, H. et al. The type of somatic mutation at APC in familial adenomatous polyposis is determined by the site of the germline mutation: a new facet to Knudson's 'two-hit' hypothesis. Nat. Med. 5, 1071–1075 (1999).
Albuquerque, C. et al. The 'just-right' signaling model: APC somatic mutations are selected based on a specific level of activation of the β-catenin signaling cascade. Hum. Mol. Genet. 11, 1549–1560 (2002).
Schneikert, J., Grohmann, A. & Behrens, J. Truncated APC regulates the transcriptional activity of β-catenin in a cell cycle dependent manner. Hum. Mol. Genet. 16, 199–209 (2007).
Segditsas, S. et al. APC and the three-hit hypothesis. Oncogene 28, 146–155 (2009).
Tuupanen, S. et al. Allelic imbalance at rs6983267 suggests selection of the risk allele in somatic colorectal tumor evolution. Cancer Res. 68, 14–17 (2008).
Dewhurst, S. M. et al. Tolerance of whole-genome doubling propagates chromosomal instability and accelerates cancer genome evolution. Cancer Discov. 4, 175–185 (2014).
Pugh, C. W. & Ratcliffe, P. J. The von Hippel–Lindau tumor suppressor, hypoxia-inducible factor-1 (HIF-1) degradation, and cancer pathogenesis. Semin. Cancer Biol. 13, 83–89 (2003).
Shen, C. et al. Genetic and functional studies implicate HIF1α as a 14q kidney cancer suppressor gene. Cancer Discov. 1, 222–235 (2011).
McFarland, C. D., Korolev, K. S., Kryukov, G. V., Sunyaev, S. R. & Mirny, L. A. Impact of deleterious passenger mutations on cancer progression. Proc. Natl Acad. Sci. USA 110, 2910–2915 (2013).
Favero, F. et al. Glioblastoma adaptation traced through decline of an IDH1 clonal driver and macroevolution of a double minute chromosome. Ann. Oncol. 26, 880–887 (2015).
Korolev, K. S., Xavier, J. B. & Gore, J. Turning ecology and evolution against cancer. Nat. Rev. Cancer 14, 371–380 (2014).
Marusyk, A. et al. Non-cell-autonomous driving of tumour growth supports sub-clonal heterogeneity. Nature 514, 54–58 (2014).
Reilly, K. M. & Van Dyke, T. It takes a (dysfunctional) village to raise a tumor. Cell 135, 408–410 (2008).
Tabassum, D. P. & Polyak, K. Tumorigenesis: it takes a village. Nat. Rev. Cancer 15, 473–483 (2015).
Tomlinson, I. P. Game-theory models of interactions between tumour cells. Eur. J. Cancer 33, 1495–1500 (1997).
Tomlinson, I. P. & Bodmer, W. F. Modelling the consequences of interactions between tumour cells. Br. J. Cancer 75, 157–160 (1997).
Miller, B. E., Miller, F. R., Wilburn, D. & Heppner, G. H. Dominance of a tumor subpopulation line in mixed heterogeneous mouse mammary tumors. Cancer Res. 48, 5747–5753 (1988).
Axelrod, R., Axelrod, D. E. & Pienta, K. J. Evolution of cooperation among tumor cells. Proc. Natl Acad. Sci. USA 103, 13474–13479 (2006).
Cleary, A. S., Leonard, T. L., Gestl, S. A. & Gunther, E. J. Tumour cell heterogeneity maintained by cooperating subclones in Wnt-driven mammary cancers. Nature 508, 113–117 (2014).
Inda, M. M. et al. Tumor heterogeneity is an active process maintained by a mutant EGFR-induced cytokine circuit in glioblastoma. Genes Dev. 24, 1731–1745 (2010).
Turke, A. B. et al. Preexistence and clonal selection of MET amplification in EGFR mutant NSCLC. Cancer Cell 17, 77–88 (2010).
Aceto, N. et al. Circulating tumor cell clusters are oligoclonal precursors of breast cancer metastasis. Cell 158, 1110–1122 (2014).
Calbo, J. et al. A functional role for tumor cell heterogeneity in a mouse model of small cell lung cancer. Cancer Cell 19, 244–256 (2011).
Marusyk, A. & Polyak, K. Tumor heterogeneity: causes and consequences. Biochim. Biophys. Acta 1805, 105–117 (2010).
Parsons, D. W. et al. An integrated genomic analysis of human glioblastoma multiforme. Science 321, 1807–1812 (2008).
Mardis, E. R. et al. Recurring mutations found by sequencing an acute myeloid leukemia genome. N. Engl. J. Med. 361, 1058–1066 (2009).
Davoli, T. et al. Cumulative haploinsufficiency and triplosensitivity drive aneuploidy patterns and shape the cancer genome. Cell 155, 948–962 (2013).
Creixell, P. et al. Pathway and network analysis of cancer genomes. Nat. Methods 12, 615–621 (2015).
Krogan, N. J., Lippman, S., Agard, D. A., Ashworth, A. & Ideker, T. The cancer cell map initiative: defining the hallmark networks of cancer. Mol. Cell 58, 690–698 (2015).
Pe'er, D. & Hacohen, N. Principles and strategies for developing network models in cancer. Cell 144, 864–873 (2011).
Gonzalez-Perez, A. & Lopez-Bigas, N. Functional impact bias reveals cancer drivers. Nucleic Acids Res. 40, e169 (2012).
Lawrence, M. S. et al. Mutational heterogeneity in cancer and the search for new cancer-associated genes. Nature 499, 214–218 (2013).
Tamborero, D., Lopez-Bigas, N. & Gonzalez-Perez, A. Oncodrive-CIS: a method to reveal likely driver genes based on the impact of their copy number changes on expression. PLoS ONE 8, e55489 (2013).
Gundem, G. et al. IntOGen: integration and data mining of multidimensional oncogenomic data. Nat. Methods 7, 92–93 (2010).
Salk, J. J., Fox, E. J. & Loeb, L. A. Mutational heterogeneity in human cancers: origin and consequences. Annu. Rev. Pathol. 5, 51–75 (2010).
Balmain, A., Gray, J. & Ponder, B. The genetics and genomics of cancer. Nat. Genet. 33, 238–244 (2003).
Heitzer, E. & Tomlinson, I. Replicative DNA polymerase mutations in cancer. Curr. Opin. Genet. Dev. 24, 107–113 (2014).
Anderson, A. R., Weaver, A. M., Cummings, P. T. & Quaranta, V. Tumor morphology and phenotypic evolution driven by selective pressure from the microenvironment. Cell 127, 905–915 (2006).
Sidow, A. & Spies, N. Concepts in solid tumor evolution. Trends Genet. 31, 208–214 (2015).
Reiter, J. G., Bozic, I., Allen, B., Chatterjee, K. & Nowak, M. A. The effect of one additional driver mutation on tumor progression. Evol. Appl. 6, 34–45 (2013).
Bozic, I. et al. Accumulation of driver and passenger mutations during tumor progression. Proc. Natl Acad. Sci. USA 107, 18545–18550 (2010).
Crespi, B. The evolutionary biology of child health. Proc. Biol. Sci. 278, 1441–1449 (2011).
Peer, P. G., van Dijck, J. A., Hendriks, J. H., Holland, R. & Verbeek, A. L. Age-dependent growth rate of primary breast cancer. Cancer 71, 3547–3551 (1993).
Egeblad, M., Nakasone, E. S. & Werb, Z. Tumors as organs: complex tissues that interface with the entire organism. Dev. Cell 18, 884–901 (2010).
Kandoth, C. et al. Integrated genomic characterization of endometrial carcinoma. Nature 497, 67–73 (2013).
Abbott, K. L. et al. The Candidate Cancer Gene Database: a database of cancer driver genes from forward genetic screens in mice. Nucleic Acids Res. 43, D844–D848 (2015).
Shain, A. & Pollack, J. The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS ONE 8, e55119 (2013).
Church D. N. et al. Prognostic significance of POLE proofreading mutations in endometrial cancer. J. Natl Cancer Inst. 107, dju402 (2015).
Acknowledgements
The authors are very grateful to T. Graham for comments and criticism. F.C.-G. and I.T. were supported by the Oxford National Institute for Health Research Comprehensive Biomedical Research Centre. Core funding to the Wellcome Trust Centre for Human Genetics was provided by the Wellcome Trust (090532/Z/09/Z).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Rights and permissions
About this article
Cite this article
Castro-Giner, F., Ratcliffe, P. & Tomlinson, I. The mini-driver model of polygenic cancer evolution. Nat Rev Cancer 15, 680–685 (2015). https://doi.org/10.1038/nrc3999
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc3999
This article is cited by
-
APOBEC mediated mutagenesis drives genomic heterogeneity in endometriosis
Journal of Human Genetics (2022)
-
Biased expression of mutant alleles in cancer-related genes in esophageal squamous cell carcinoma
Esophagus (2022)
-
Phosphoproteomic analysis of neoadjuvant breast cancer suggests that increased sensitivity to paclitaxel is driven by CDK4 and filamin A
Nature Communications (2022)
-
Network propagation-based prioritization of long tail genes in 17 cancer types
Genome Biology (2021)
-
Clonal evolution driven by superdriver mutations
BMC Evolutionary Biology (2020)