Key Points
-
Many cancer genes remain functionally uncharacterized. Experimental methods to characterize their functions are inefficient, time consuming and expensive.
-
The increasing availability of diverse molecular profiles and functional-interaction data make the prediction of cancer-gene functions possible.
-
New computational prediction methods now enable the automated assessment of cancer-gene function.
-
The main difficulties are how to simultaneously integrate different high-throughput data sources and dependably assign multiple functions to a cancer gene.
-
Trustworthy gene annotations are crucial to achieving the best possible functional predictions for newly discovered or uncharacterized cancer genes.
-
Rigorous evaluation of the accuracy of functional predictions generated by computational methods is vital for formulating biologically relevant hypotheses to direct further rounds of experimentation.
Abstract
Most cancer genes remain functionally uncharacterized in the physiological context of disease development. High-throughput molecular profiling and interaction studies are increasingly being used to identify clusters of functionally linked gene products related to neoplastic cell processes. However, in vivo determination of cancer-gene function is laborious and inefficient, so accurately predicting cancer-gene function is a significant challenge for oncologists and computational biologists alike. How can modern computational and statistical methods be used to reliably deduce the function(s) of poorly characterized cancer genes from the newly available genomic and proteomic datasets? We explore plausible solutions to this important challenge.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Hanash, S. Integrated global profiling of cancer. Nature Rev. Cancer 4, 638–644 (2004).
Rhodes, D. R. & Chinnaiyan, A. M. Integrative analysis of the cancer transcriptome. Nature Genet. 37 (Suppl.), S31–S37 (2005).
Segal, E., Friedman, N., Kaminski, N., Regev, A. & Koller, D. From signatures to models: understanding cancer using microarrays. Nature Genet. 37, S38–S45 (2005).
Vogelstein, B. & Kinzler, K. W. Cancer genes and the pathways they control. Nature Med. 10, 789–799 (2004).
van't Veer, L. J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Kastan, M. B. & Bartek, J. Cell-cycle checkpoints and cancer. Nature 432, 316–323 (2004).
Roberts, R. J. Identifying protein function — a call for community action. PLoS Biology 2, E42 (2004).
Alm, E. & Arkin, A. P. Biological networks. Curr. Opin. Struct. Biol. 13, 193–202 (2003).
Barabasi, A. & Oltvai, Z. N. Network biology: understanding the cell's functional organization. Nature Rev. Genet. 5, 101–113 (2004). The authors review current network tools that can be used to understand the cell's functional organization and evolution.
Mateos, A. et al. Systematic learning of gene functional classes from DNA array expression data by using multilayer perceptrons. Genome Res. 12, 1703–1715 (2002).
Pavlidis, P., Weston, J., Cai, J. & Noble, W. S. Learning gene functional classifications from multiple data types. J. Comp. Biol. 9, 401–411 (2002).
Troyanskaya, O. G., Dolinski, K., Owen, A. B., Altman, R. B. & Botstein, D. A Bayesian framework for combining heterogeneous data source for gene function prediction (in Saccharomyces cerevisiae). Proc. Natl Acad. Sci USA 100, 8348–8353 (2003). The authors present an effective computational method to integrate different functional-association data sets for gene-function prediction.
Jansen, R., Greenbaum, D. & Gerstein, M. Relating whole-genome expression data with protein–protein interactions. Genome Res. 12, 37–46 (2002).
Lee, L., Date, S. V., Adai, A. T. & Marcotte, E. M. A probabilistic functional network of yeast genes. Science 306, 1555–1558 (2004).
Zhang, W. et al. The functional landscape of mouse gene expression. J. Biol. 3, 21 (2004).
Lanckriet, G. R. G., Deng, M., Gristianini, N., Jordan, M. I. & Noble, W. S. Kernel-based data fusion and its application to protein function prediction in yeast. Proceedings of the Pacific Symposium on Biocomputing (PSB), 300–311 (2004).
Nabieva, E., Jim, K., Agarwal, A., Chazelle, B. & Singh, M. Whole-proteome prediction of protein function via graph-theoretic analysis of interaction maps. Bioinformatics 21 (suppl. 1), i302–i310 (2005). The authors present one of the most efficient network-based label-propagation methods to make gene-function predictions using functional-association data.
Barutcuoglu, Z., Schapire, R. E. & Troyanskaya, O. G. Hierarchical multi-label prediction of gene function. Bioinformatics 22, 830–836 (2006).
Vidal, M. Interactome modeling. FEBS Lett. 579, 1834–1838 (2005).
Futreal, P. A. et al. A census of human cancer genes. Nature Rev. Cancer 4, 177–183 (2004).
Strausberg, R. L., Simpson, A. J. & Wooster, R. Sequence-based cancer genomics: progress, lessons and opportunities. Nature Rev. Genet. 4, 409–418 (2003).
Koenig, M. et al. Complete cloning of the Duchenne muscular dystrophy (DMD) cDNA and preliminary genomic organization of the DMD gene in normal and affected individuals. Cell 50, 509–517 (1987).
Tannock, I. F., Hill, R. P., Bristow, R. G. & Harrington, L. The basic science of oncology 4th ed. (McGraw Hill Companies Inc., New York, 2005).
Clark, J. et al. Genome-wide screening for complete genetic loss in prostate cancer by comparative hybridization onto cDNA microarrays. Oncogene 22, 1247–1252 (2003).
American Cancer Society. Cancer Facts and Figures 2006. American Cancer Society [online], http://www.cancer.org/downloads/STT/CAFF2006PWSecured.pdf
Balmain, A., Gray, J. & Ponder, B. The genetics and genomics of cancer. Nature Genet. 33 (Suppl.), 238–244 (2003).
Demant, P. Cancer susceptibility in the mouse: genetics, biology and implications for human cancer. Nature Rev. Genet. 4, 721–734 (2003).
Segal, E., Friedman, N., Koller, D. & Regev, A. A module map showing conditional activity of expression modules in cancer. Nature Genet. 36, 1090–1098 (2004). The authors develop a strategy to identify functional modules that are common among, or unique to, different types of tumours. The set of genes in each module can also be treated as a gold standard for cancer-gene-function prediction.
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acid Res. 25, 3389–3402 (1997).
Wiseman, B. S. & Werb, Z. Stromal effects on mammary gland development and breast cancer. Science 296, 1046–1049 (2002).
Sawyers, C. L. Chronic myeloid leukemia. N. Engl. J. Med. 340, 1330–1340 (1999).
Harris, M. A. et al. The Gene Ontology (GO) database and informatics resource. Nucleic Acids Res. 32 (Database issue), D258–D261 (2004).
Chen, Y. & Xu, D. Global protein function annotation through mining genome-scale data in yeast Saccharomyces cerevisiae. Nucleic Acids Res. 32, 6414–6424 (2004).
Wu, H., Su, Z., Mao, F., Olman, V. & Xu, Y. Prediction of functional modules based on comparative genome analysis and gene ontology application. Nucleic Acids Res. 33, 2822–2837 (2005).
Ronald, L. et al. Human homolog of patched, a candidate gene for the basal cell nevus syndrome. Science 272, 1668–1671 (1996).
Zhang, B. & Horvath, S. A general framework for weighted gene co-expression network analysis. Stat. Appl. Genet. Mol. Biol. 4, Article 17 (2005).
Pawson, T. & Nash, P. Assembly of cell regulatory systems through protein interaction domains. Science 300, 445–452 (2003).
Barrios-Rodiles, M. et al. High-throughput mapping of a dynamic signaling network in mammalian cells. Science 307, 1621–1625 (2005).
Bouwmeester, T. et al. A physical and functional map of the human TNF-α/NF-κB signal transduction pathway. Nature Cell Biol. 6, 97–105 (2004).
Stelzl, U. et al. A human protein–protein interaction network: a resource for annotating the proteome. Cell 122, 957–968 (2005).
Rual, J. F. et al. Towards a proteome-scale map of the human protein–protein interaction network. Nature 437, 1173–1178 (2005).
Boyer, L. A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Wu, L. F. et al. Large-scale prediction of Saccharomyces cerevisiae gene function using overlapping transcriptional clusters. Nature Genet. 31, 255–265 (2002).
Kislinger, T. et al. Global survey of organ and organelle selective protein expression in mouse: integrated proteomic, genomic and bioinformatic analysis. Cell 125, 173–186 (2006).
Bandyopadhyay, S., Sharan, R. & Ideker, T. Systematic identification of functional orthologs based on protein network comparison. Genome Res. 16, 428–435 (2006).
Stuart, J. M., Segal, E., Koller, D. & Kim, S. K. A gene-coexpression network for global discovery of conserved genetic modules. Science 302, 249–255 (2003).
Jonsson, P. F. & Bates, P. A. Global topological features of cancer proteins in the human interactome. Bioinformatics 22, 2291–2297 (2006). The authors show that human proteins translated from known cancer genes have a protein–protein interaction network topology that is different from that of proteins not documented as being mutated in cancer.
Bader, G. D., Cary, M. P. & Sander, C. Pathguide: a pathway resource list. Nucleic Acids Res. 34 (Database issue), D504–D506 (2006).
Chua, H. N., Sung, W. & Wong, L. Exploiting indirect neighbours and topological weight to predict protein function from protein–protein interactions. Bioinformatics 22, 1623–1630 (2006).
Brun, C., Herrmann, C. & Guenoche, A. Clustering proteins from interaction networks for the prediction of cellular functions. BMC Bioinformatics 5, 95 (2004).
Pereira-Leal, J. B., Enright, A. J. & Quzounis, C. A. Detection of functional modules from protein interaction networks. Proteins 54, 49–57 (2004).
Farutin, V. et al. Edge-count probabilities for the identification of local protein communities and their organization. Proteins 62, 800–818 (2006).
Adamcsek, B. et al. CFinder: locating cliques and overlapping modules in biological networks. Bioinformatics 22, 1021–1023 (2006).
Aittokallio, T. & Schwikowski, B. Graph-based methods for analyzing networks in cell biology. Brief. Bioinformatics 7, 243–255 (2006).
Schwikowski, B., Uetz, P. & Fields, S. A network of protein–protein interactions in yeast. Nature Biotechnol. 18, 1257–1261 (2000).
Tsuda, K. & Noble, W. S. Learning kernels from biological networks by maximizing entropy. Bioinformatics 20 (Suppl.1), I326–I333 (2004).
Goldstein, D. R., Ghosh, D. & Conlon, E. M. Statistical issues in the clustering of gene expression data. Statistica Sinica 12, 219–240 (2002).
Jansen, R. & Gerstein, M. Analyzing protein function on a genomic scale: the importance of gold-standard positives and negatives for network prediction. Curr. Opin. Microbiol. 7, 535–545 (2004). The authors discuss how to define protein functions and select gold standards for protein-function prediction using functional-association data.
Myers, C. L., Barrett, D. R., Hibbs, M. A., Huttenhower, C. & Troyanskaya, O. G. Finding function: evaluation methods for functional genomic data. BMC Genomics 7, 187 (2006). The authors discuss the deficiencies of current computational methods to infer functions from functional-association data, and outline new approaches to deal with these problems.
Devos, D. & Valencia, A. Intrinsic errors in genome annotation. Trends Genet. 17, 429–431 (2001).
Letovsky, S. & Kasif, S. Predicting protein function from protein/protein interaction data: a probabilistic approach. Bioinformatics 19 (Suppl.1), i197–i204 (2003).
Tsuda, K., Uda, S., Kin, T. & Asai, K. Minimizing the cross validation error to mix kernel matrices of heterogeneous biological data. Neural Process. Lett. 19, 63–72 (2004).
Boocock, G. R. et al. Mutations in SBDS are associated with Shwachman–Diamond syndrome. Nature Genet. 33, 97–101 (2003).
Woloszynek, J. R. et al. Mutations of the SBDS gene are present in most patients with Shwachman–Diamond syndrome. Blood 104, 3588–3590 (2004).
Austin, K. M., Leary, R. J. & Shimamura, A. The Shwachman–Diamond SBDS protein localizes to the nucleolus. Blood 106, 1253–1258 (2005).
von Mering, C. et al. STRING: a database of predicted functional associations between proteins. Nucleic Acids Res. 31, 258–261 (2003).
Boeckmann, B. et al. The SWISS-PROT protein knowledgebase and its supplement TrEMBL in 2003. Nucleic Acids Res. 31, 365–370 (2003).
Savchenko, A. et al. The Shwachman–Bodian–Diamond syndrome protein family is involved in RNA metabolism. J. Biol. Chem. 280, 19213–19220 (2005).
Martinez, N. et al. The molecular signature of mantle cell lymphoma reveals multiple signals favoring cell survival. Cancer Res. 63, 8226–8232 (2003).
Yamamoto, S. et al. High frequency of fusion transcripts of exon 11 and exon 4/5 in AF-4 gene is observed in cord blood, as well as leukemic cells from infant leukemia patients with t(4;11)(q21;q23). Leukemia 12, 1398–1403 (1998).
Zhu, X., Ghahramani, Z. & Lafferty, J. Semi-supervised learning using Gaussian fields and harmonic functions. Proc. Twentieth Int. Conf. Machine Learning 20, 912–919 (2003).
Hanahan, D. & Weinberg, R. A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Karaoz, U. et al. Whole-genome annotation by using evidence integration in functional – linkage networks. Proc. Natl Acad. Sci. USA 101, 2883–2893 (2004).
Khalil, I. G. & Hill, C. Systems biology for cancer. Curr. Opin. Oncol. 17, 44–48 (2005).
Deng, M. & Chen, T. S. & Sun,F. An integrated probabilistic model for functional prediction of proteins. Proc. Seventh Ann. Int. Conf. Res. Comp. Mol. Biol. (RECOMB), Berlin, Germany, 95–103 (2003).
Vazquez, A., Flammini, A., Maritan, A. & Vespignani, A. Global protein function prediction from protein-protein interaction networks. Nature Biotechnol. 21, 697–700 (2003).
Mewes, H. W. MIPS: a database for genomes and protein sequences. Nucleic Acids Res. 30, 31–34 (2002).
Dahlquist, K. D., Salomonis, N., Vranizan, K., Lawlor, S. C. & Conklin, B. R. GenMAPP, a new tool for viewing and analyzing microarray data on biological pathways. Nature Genet. 31, 19–20 (2002).
Kanehisa, M., Goto, S., Kawashima, S., Okuno, Y. & Hattori, M. The KEGG resource for deciphering the genome. Nucleic Acids Res. 32 (Database issue), D277−D280 (2004).
Bader, G. D., Betel, D. & Hogue, C. W. BIND: the biomolecular interaction network database. Nucleic Acids Res. 31, 248–250 (2003).
Hermjakob, H. et al. IntAct: an open source molecular interaction database. Nucleic Acids Res. 32 (Database issue), D452–D455 (2004).
Peri, S. et al. Development of human protein reference database as an initial platform for approaching systems biology in humans. Genome Res. 13, 2363–2371 (2003).
Xenarios, I. et al. DIP, the Database of Interacting Proteins: a research tool for studying cellular networks of protein interactions. Nucleic Acids Res. 30, 303–305 (2002).
Zanzoni, A. et al. MINT: a Molecular INTeraction database. FEBS Lett. 513, 135–140 (2002).
Dennis, G. Jr et al. DAVID: database for annotation, visualization, and Integrated discovery. Genome Biol. 4, R60 (2003).
Jiang, T. & Keating, A. E. AVID: an integrative framework for discovering functional relationships among proteins. BMC Bioinformatics 6, 136 (2005).
Date, S. V. & Marcotte, E. M. Protein function prediction using the protein link explorer (PLEX). Bioinformatics 21, 2558–2559 (2005).
Brown, K.R. & Jurisica, I. Online predicted human interaction database. Bioinformatics 21, 2076–2082 (2005).
Maere, S., Heymans, K. & Kuiper, M. BINGO: a cytoscape plugin to assess overrepresentation of gene ontology categories in biological networks. Bioinformatics 21, 3448–3449 (2005).
AI-Sharour, F., Minguez, P., Vaquerizas, J.M., Conde, L. & Dopazo, J. Babelomics: a suite of web/tools for functional annotation and analysis of groups of genes in high-thoughout experiments, Nucleic Acids Res. 33, W460–W464 (2005).
Acknowledgements
We thank H. Jiang, Q. Morris and B. Noble for their critical feedback and thoughtful suggestions, R. Isserlin for skillful preparation of the GO-tree analysis and M. Maris for expert computational support. This work was supported in part by funds from Genome Canada and the Ontario Genomics Institute to A.E.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Related links
Glossary
- Global
-
A large-scale or genome-wide biological perspective, often with reference to high-throughput experimental datasets.
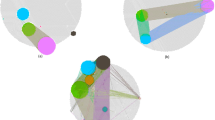
- Interaction network
-
A graphical description of a large ensemble of molecular associations, the nodes of which correspond to gene products, and the edges of which reflect direct links or connections between the gene products.
- Hierarchical clustering
-
A statistical method for finding relatively homogeneous clusters of gene products based on some measure of similarity.
- Functional module
-
A set of gene products that together function in a single process.
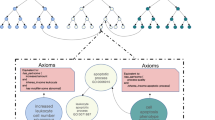
- Directed acyclic graph
-
A network data structure used to represent a gene-function classification system in the Gene Ontology database, having ordered relationships between nodes (for example, parent and child terms, wherein the graph direction indicates which term is subsumed by the other), and no cycles (no path returns to the same node twice). Nested terms can have several parents.
- Supervised learning
-
A computational procedure to identify sets of gene products that are similar to a reference set of manually-defined examples using a principled-prediction rule or criteria. Any genes of unknown function that are grouped with the set of pre-defined genes are deemed similar in function.
- Unsupervised learning
-
A computational procedure to identify subsets of gene products that are more similar to each other than to others. The function of unknown genes can then be predicted based on the functions of other known genes within a given cluster.
- Functional label
-
The function terms, such as Gene Ontology terms, that are assigned to cancer genes.
- Functional-association network
-
An interaction network in which gene products are linked if they have experimentally measured or predicted functional associations.
- Gold standard
-
A reference gene set used for labelling learning data, both for building prediction models and for creating test data to evaluate classifier performance.
- Cross-validation
-
A statistical method for evaluating a classifier model. The input-association data is randomly partitioned into at least two or more subsets such that the analysis is initially performed on a single subset (learning set), whereas the other subset(s) (test set) is retained for subsequent use in testing and validating the initial analysis. This splitting can be done many times independently to better assess the accuracy of the classifier.
- Over-fitting
-
The phenomenon in which a model has too many free parameters relative to the amount of data, which results in the learning of not only the true functional associations, but also noise and other spurious correlations. A model which has been over-fitted will not make good predictions on fresh (previously unseen) data — that is, the classifier will not generalize well.
- Receiver operating characteristic
-
ROC curves are usually drawn by plotting sensitivity versus specificity or positive predictive value versus recall to evaluate the performance of computational methods in the cross-validation procedure.
- Sensitivity
-
Also called recall. A measure of the ability of a classifier to assign all appropriate genes present in the test dataset the correct relevant functional label. Sensitivity is the proportion of all known members of a functional category for which there is a positive assignment, as determined by the number of true positives divided by the sum of true positives and false negatives. (Contrast with specificity.)
- Specificity
-
An operating characteristic of a functional-prediction procedure that measures the ability of a classifier to exclude the presence of a label when it is truly not warranted. Specificity is defined as the number of true negatives divided by the sum of true negatives and false positives. (Contrast with sensitivity and recall.)
- Precision
-
Also called 'positive predictive value'. The proportion of gene products with a predicted function that truly have the assigned biological attributes, as determined by the number of true positives divided by the sum of true positives and false positives.
- Discriminant value
-
A relative measure of confidence that the cancer gene is in the functional category in question.
- Genomic context
-
Similarity among the evolutionary attributes of gene products, such as the propensity of functionally linked gene products to co-occur across the genomes of several species, to be involved in gene-fusion events, or to be conserved in close chromosomal proximity.
- Multi-function prediction
-
A computational procedure wherein a cancer gene product is assigned to at least two or more functional classes.
- Correlation structure
-
A statistical measure of the relationships observed between all pair-wise functional classes examined.
- Support vector machine
-
A popular learning algorithm that performs binary or multi-class supervised classification tasks.
Rights and permissions
About this article
Cite this article
Hu, P., Bader, G., Wigle, D. et al. Computational prediction of cancer-gene function. Nat Rev Cancer 7, 23–34 (2007). https://doi.org/10.1038/nrc2036
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrc2036
This article is cited by
-
Impact of the Continuous Evolution of Gene Ontology on the Performance of Similarity Measures for Scoring Confidence of Protein Interactions
SN Computer Science (2020)
-
A model to predict the function of hypothetical proteins through a nine-point classification scoring schema
BMC Bioinformatics (2019)
-
A machine-learned computational functional genomics-based approach to drug classification
European Journal of Clinical Pharmacology (2016)
-
Computational functional genomics based analysis of pain-relevant micro-RNAs
Human Genetics (2015)
-
What do all the (human) micro-RNAs do?
BMC Genomics (2014)