Abstract
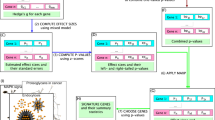
This protocol details the use of the mode-of-action by network identification (MNI) algorithm to identify the gene targets of a drug treatment based on gene-expression data. Investigators might also use the MNI algorithm to identify the gene mediators of a disease or the physiological state of cells and tissues. The MNI algorithm uses a training data set of hundreds of expression profiles to construct a statistical model of gene-regulatory networks in a cell or tissue. The model describes combinatorial influences of genes on one another. The algorithm then uses the model to filter the expression profile of a particular experimental treatment and thereby distinguish the molecular targets or mediators of the treatment response from hundreds of additional genes that also exhibit expression changes. It takes ∼1 h per run, although run time is significantly affected by the size of the genome and data set.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Similar content being viewed by others
References
Ambesi-Impiombato, A. & di Bernado, D. Computational biology and drug discovery: from single-target to network-target. Curr. Bioinform. 1, 3–13 (2006).
di Bernardo, D. et al. Chemogenomic profiling on a genome-wide scale using reverse-engineered gene networks. Nat. Biotech. 23, 377–383 (2005).
Bolstad, B.M., Irizarry, R.A., Astrand, M. & Speed, T.P. A comparison of normalization methods for high density oligonucleotide array data based on bias and variance. Bioinformatics 19, 185–193 (2003).
Yang, Y.H. et al. Normalization for cDNA microarray data: a robust composite method addressing single and multiple slide systematic variation. Nucleic Acids Res. 30, e15 (2002).
Ashburner, M. et al. Gene Ontology: tool for the unification of biology. Nat. Genet. 25, 25–29 (2000).
Curtis, R.K., Oresic, M. & Vidal-Puig, A. Pathways to the analysis of microarray data. Trends Biotech. 23, 429–435 (2005).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
H.X. is an employee of Cellicon Biotechnologies, Inc. T.S.G. is a founder of Cellicon Biotechnologies, Inc.
Rights and permissions
About this article
Cite this article
Xing, H., Gardner, T. The mode-of-action by network identification (MNI) algorithm: a network biology approach for molecular target identification. Nat Protoc 1, 2551–2554 (2006). https://doi.org/10.1038/nprot.2006.300
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2006.300
This article is cited by
-
Learning perturbation-inducible cell states from observability analysis of transcriptome dynamics
Nature Communications (2023)
-
Olfactory receptor 10J5 responding to α-cedrene regulates hepatic steatosis via the cAMP–PKA pathway
Scientific Reports (2017)
-
Network based elucidation of drug response: from modulators to targets
BMC Systems Biology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.