Abstract
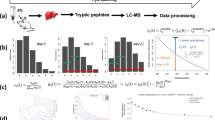
The protein ubiquitin is an important post-translational modifier that regulates a wide variety of biological processes. In cells, ubiquitin is apportioned among distinct pools, which include a variety of free and conjugated species. Although maintenance of a dynamic and complex equilibrium among ubiquitin pools is crucial for cell survival, the tools necessary to quantify each cellular ubiquitin pool have been limited. We have developed a quantitative mass spectrometry approach to measure cellular concentrations of ubiquitin species using isotope-labeled protein standards and applied it to characterize ubiquitin pools in cells and tissues. Our method is convenient, adaptable and should be a valuable tool to facilitate our understanding of this important signaling molecule.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Ravid, T. & Hochstrasser, M. Diversity of degradation signals in the ubiquitin-proteasome system. Nat. Rev. Mol. Cell Biol. 9, 679–690 (2008).
Mukhopadhyay, D. & Riezman, H. Proteasome-independent functions of ubiquitin in endocytosis and signaling. Science 315, 201–205 (2007).
Zeng, W. et al. Reconstitution of the RIG-I pathway reveals a signaling role of unanchored polyubiquitin chains in innate immunity. Cell 141, 315–330 (2010).
Hoeller, D. & Dikic, I. Targeting the ubiquitin system in cancer therapy. Nature 458, 438–444 (2009).
Lee, B.H. et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 467, 179–184 (2010).
Takada, K., Hibi, N., Tsukada, Y., Shibasaki, T. & Ohkawa, K. Ability of ubiquitin radioimmunoassay to discriminate between monoubiquitin and multi-ubiquitin chains. Biochim. Biophys. Acta 1290, 282–288 (1996).
Haas, A.L. & Bright, P.M. The immunochemical detection and quantitation of intracellular ubiquitin-protein conjugates. J. Biol. Chem. 260, 12464–12473 (1985).
Bennett, E.J. et al. Global changes to the ubiquitin system in Huntington's disease. Nature 448, 704–708 (2007).
Gerber, S.A., Rush, J., Stemman, O., Kirschner, M.W. & Gygi, S.P. Absolute quantification of proteins and phosphoproteins from cell lysates by tandem MS. Proc. Natl. Acad. Sci. USA 100, 6940–6945 (2003).
Kirkpatrick, D.S. et al. Quantitative analysis of in vitro ubiquitinated cyclin B1 reveals complex chain topology. Nat. Cell Biol. 8, 700–710 (2006).
Brun, V. et al. Isotope-labeled protein standards: toward absolute quantitative proteomics. Mol. Cell. Proteomics 6, 2139–2149 (2007).
Catanzariti, A.M., Soboleva, T.A., Jans, D.A., Board, P.G. & Baker, R.T. An efficient system for high-level expression and easy purification of authentic recombinant proteins. Protein Sci. 13, 1331–1339 (2004).
Reyes-Turcu, F.E. et al. The ubiquitin binding domain ZnF UBP recognizes the C-terminal diglycine motif of unanchored ubiquitin. Cell 124, 1197–1208 (2006).
Ryu, K.Y., Baker, R.T. & Kopito, R.R. Ubiquitin-specific protease 2 as a tool for quantification of total ubiquitin levels in biological specimens. Anal. Biochem. 353, 153–155 (2006).
Ryu, K.Y. et al. The mouse polyubiquitin gene UbC is essential for fetal liver development, cell-cycle progression and stress tolerance. EMBO J. 26, 2693–2706 (2007).
Dantuma, N.P., Groothuis, T.A., Salomons, F.A. & Neefjes, J. A dynamic ubiquitin equilibrium couples proteasomal activity to chromatin remodeling. J. Cell Biol. 173, 19–26 (2006).
Mimnaugh, E.G., Chen, H.Y., Davie, J.R., Celis, J.E. & Neckers, L. Rapid deubiquitination of nucleosomal histones in human tumor cells caused by proteasome inhibitors and stress response inducers: effects on replication, transcription, translation, and the cellular stress response. Biochemistry 36, 14418–14429 (1997).
Xu, P. et al. Quantitative proteomics reveals the function of unconventional ubiquitin chains in proteasomal degradation. Cell 137, 133–145 (2009).
Meierhofer, D., Wang, X., Huang, L. & Kaiser, P. Quantitative analysis of global ubiquitination in HeLa cells by mass spectrometry. J. Proteome Res. 7, 4566–4576 (2008).
Phu, L. et al. Improved quantitative mass spectrometry methods for characterizing complex ubiquitin signals. Mol. Cell. Proteomics 10, M110.003756 (2011).
Kim, H.T. et al. Certain pairs of ubiquitin-conjugating enzymes (E2s) and ubiquitin-protein ligases (E3s) synthesize nondegradable forked ubiquitin chains containing all possible isopeptide linkages. J. Biol. Chem. 282, 17375–17386 (2007).
Raasi, S., Varadan, R., Fushman, D. & Pickart, C.M. Diverse polyubiquitin interaction properties of ubiquitin-associated domains. Nat. Struct. Mol. Biol. 12, 708–714 (2005).
Zhang, D., Raasi, S. & Fushman, D. Affinity makes the difference: nonselective interaction of the UBA domain of Ubiquilin-1 with monomeric ubiquitin and polyubiquitin chains. J. Mol. Biol. 377, 162–180 (2008).
Raasi, S., Orlov, I., Fleming, K.G. & Pickart, C.M. Binding of polyubiquitin chains to ubiquitin-associated (UBA) domains of HHR23A. J. Mol. Biol. 341, 1367–1379 (2004).
Song, J. et al. Stability of thioester intermediates in ubiquitin-like modifications. Protein Sci. 18, 2492–2499 (2009).
Newton, K. et al. Ubiquitin chain editing revealed by polyubiquitin linkage-specific antibodies. Cell 134, 668–678 (2008).
Riley, B.E. et al. Ubiquitin accumulation in autophagy-deficient mice is dependent on the Nrf2-mediated stress response pathway: a potential role for protein aggregation in autophagic substrate selection. J. Cell Biol. 191, 537–552 (2010).
Hosokawa, N., Hara, Y. & Mizushima, N. Generation of cell lines with tetracycline-regulated autophagy and a role for autophagy in controlling cell size. FEBS Lett. 580, 2623–2629 (2006).
Eftekharzadeh, B., Maghsoudi, N. & Khodagholi, F. Stabilization of transcription factor Nrf2 by tBHQ prevents oxidative stress-induced amyloid beta formation in NT2N neurons. Biochimie 92, 245–253 (2010).
Shechter, D., Dormann, H.L., Allis, C.D. & Hake, S.B. Extraction, purification and analysis of histones. Nat. Protoc. 2, 1445–1457 (2007).
Acknowledgements
We thank E. Bennett, M. Reese and M. Bowen for discussions and critical reading of the manuscript. This work was funded in part by a grant (NS 04842) from the US National Institute of Neurological Disorders and Stroke.
Author information
Authors and Affiliations
Contributions
S.E.K. and R.R.K. devised the Ub-PSAQ strategy with contributions from B.E.R. and T.A.S.; S.E.K. prepared all protein affinity reagents and standards, performed the experiments and analyzed the data with input from B.E.R. and T.A.S.; B.E.R. prepared all cellular samples; T.A.S. performed all mass spectrometry analyses; and R.S.T. contributed to mass spectrometry data analysis. S.E.K. and R.R.K. wrote the manuscript, and B.E.R. contributed to figure preparation. B.E.R., T.A.S. and R.S.T. contributed to editing. C.H.B. and H.S. contributed to conceptual and experimental design. All authors discussed the results and manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 2003 kb)
Supplementary Data 1
Overview of Ub-PSAQ analysis using the samples described in Figure 3b as an example. (XLS 225 kb)
Rights and permissions
About this article
Cite this article
Kaiser, S., Riley, B., Shaler, T. et al. Protein standard absolute quantification (PSAQ) method for the measurement of cellular ubiquitin pools. Nat Methods 8, 691–696 (2011). https://doi.org/10.1038/nmeth.1649
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1649
This article is cited by
-
Current methodologies in protein ubiquitination characterization: from ubiquitinated protein to ubiquitin chain architecture
Cell & Bioscience (2022)
-
Cryo-EM structures of Gid12-bound GID E3 reveal steric blockade as a mechanism inhibiting substrate ubiquitylation
Nature Communications (2022)
-
Deubiquitinating enzymes and the proteasome regulate preferential sets of ubiquitin substrates
Nature Communications (2022)
-
A Dual Functional-Group Derivatization Liquid Chromatography–Tandem Mass Spectrometry Method: Application for Quantification of Human Insulin
Chromatographia (2022)
-
The murine ortholog of Kaufman oculocerebrofacial syndrome protein Ube3b regulates synapse number by ubiquitinating Ppp3cc
Molecular Psychiatry (2021)