Abstract
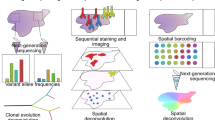
Intratumor heterogeneity (ITH) drives neoplastic progression and therapeutic resistance. We used the bioinformatics tools 'expanding ploidy and allele frequency on nested subpopulations' (EXPANDS) and PyClone to detect clones that are present at a ≥10% frequency in 1,165 exome sequences from tumors in The Cancer Genome Atlas. 86% of tumors across 12 cancer types had at least two clones. ITH in the morphology of nuclei was associated with genetic ITH (Spearman's correlation coefficient, ρ = 0.24–0.41; P < 0.001). Mutation of a driver gene that typically appears in smaller clones was a survival risk factor (hazard ratio (HR) = 2.15, 95% confidence interval (CI): 1.71–2.69). The risk of mortality also increased when >2 clones coexisted in the same tumor sample (HR = 1.49, 95% CI: 1.20–1.87). In two independent data sets, copy-number alterations affecting either <25% or >75% of a tumor's genome predicted reduced risk (HR = 0.15, 95% CI: 0.08–0.29). Mortality risk also declined when >4 clones coexisted in the sample, suggesting a trade-off between the costs and benefits of genomic instability. ITH and genomic instability thus have the potential to be useful measures that can universally be applied to all cancers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Marusyk, A. & Polyak, K. Tumor heterogeneity: causes and consequences. Biochim. Biophys. Acta 1805, 105–117 (2010).
Bonavia, R., Inda, M.-M., Cavenee, W.K. & Furnari, F.B. Heterogeneity maintenance in glioblastoma: a social network. Cancer Res. 71, 4055–4060 (2011).
Wang, Y. et al. Clonal evolution in breast cancer revealed by single-nucleus genome sequencing. Nature 512, 155–160 (2014).
Nowell, P.C. The clonal evolution of tumor cell populations. Science 194, 23–28 (1976).
Greaves, M. & Maley, C.C. Clonal evolution in cancer. Nature 481, 306–313 (2012).
McGranahan, N. et al. Clonal status of actionable driver events and the timing of mutational processes in cancer evolution. Sci. Transl. Med. 7, 283ra54 (2015).
Nik-Zainal, S. et al. The life history of 21 breast cancers. Cell 149, 994–1007 (2012).
Landau, D.A. et al. Evolution and impact of subclonal mutations in chronic lymphocytic leukemia. Cell 152, 714–726 (2013).
Mroz, E.A., Tward, A.D., Hammon, R.J., Ren, Y. & Rocco, J.W. Intratumor genetic heterogeneity and mortality in head and neck cancer: analysis of data from the Cancer Genome Atlas. PLoS Med. 12, e1001786 (2015).
Zack, T.I. et al. Pan-cancer patterns of somatic copy number alteration. Nat. Genet. 45, 1134–1140 (2013).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Ciriello, G. et al. Emerging landscape of oncogenic signatures across human cancers. Nat. Genet. 45, 1127–1133 (2013).
Almendro, V. et al. Inference of tumor evolution during chemotherapy by computational modeling and in situ analysis of genetic and phenotypic cellular diversity. Cell Rep. 6, 514–527 (2014).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Oesper, L., Satas, G. & Raphael, B.J. Quantifying tumor heterogeneity in whole-genome and whole-exome sequencing data. Bioinformatics 30, 3532–3540 (2014).
Li, B. & Li, J.Z. A general framework for analyzing tumor subclonality using SNP array and DNA sequencing data. Genome Biol. 15, 473 (2014).
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nat. Methods 11, 396–398 (2014).
Andor, N., Harness, J.V., Müller, S., Mewes, H.W. & Petritsch, C. EXPANDS: expanding ploidy and allele frequency on nested subpopulations. Bioinformatics 30, 50–60 (2014).
Ha, G. et al. TITAN: inference of copy number architectures in clonal cell populations from tumor whole-genome sequence data. Genome Res. 24, 1881–1893 (2014).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Sathirapongsasuti, J.F. et al. Exome sequencing–based copy-number variation and loss of heterozygosity detection: ExomeCNV. Bioinformatics 27, 2648–2654 (2011).
Alexandrov, L.B. et al. Signatures of mutational processes in human cancer. Nature 500, 415–421 (2013).
Vogelstein, B. et al. Cancer genome landscapes. Science 339, 1546–1558 (2013).
Barber, L.J., Davies, M.N. & Gerlinger, M. Dissecting cancer evolution at the macro-heterogeneity and micro-heterogeneity scale. Curr. Opin. Genet. Dev. 30, 1–6 (2015).
Yadav, V.K. & De, S. An assessment of computational methods for estimating purity and clonality using genomic data derived from heterogeneous tumor tissue samples. Brief. Bioinform. 16, 232–241 (2015).
Yoshihara, K. et al. Inferring tumor purity, and stromal and immune cell admixture from expression data. Nat. Commun. 4, 2612 (2013).
Tajiri, R. et al. Intratumoral heterogeneous amplification of ERBB2 and subclonal genetic diversity in gastric cancers revealed by multiple ligation-dependent probe amplification and fluorescence in situ hybridization. Hum. Pathol. 45, 725–734.
Sakurada, A., Lara-Guerra, H., Liu, N., Shepherd, F.A. & Tsao, M.-S. Tissue heterogeneity of EGFR mutation in lung adenocarcinoma. J. Thorac. Oncol. 3, 527–529 (2008).
Imielinski, M. et al. Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150, 1107–1120 (2012).
Vitale, M. Intratumor BRAFV600E heterogeneity and kinase inhibitors in the treatment of thyroid cancer: a call for participation. Thyroid 23, 517–519 (2013).
Carpenter, A.E. et al. CellProfiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biol. 7, R100 (2006).
Wang, W., Ozolek, J.A. & Rohde, G.K. Detection and classification of thyroid follicular lesions based on nuclear structure from histopathology images. Cytometry A 77, 485–494 (2010).
Hartwell, K.A. et al. Niche-based screening identifies small-molecule inhibitors of leukemia stem cells. Nat. Chem. Biol. 9, 840–848 (2013).
Yamamoto, S. et al. Clinical relevance of Ki67 gene expression analysis using formalin-fixed paraffin-embedded breast cancer specimens. Breast Cancer 20, 262–270 (2013).
Cazier, J.-B. et al. Whole-genome sequencing of bladder cancers reveals somatic CDKN1A mutations and clinicopathological associations with mutation burden. Nat. Commun. 5, 3756 (2014).
Swanton, C. Cancer evolution constrained by mutation order. N. Engl. J. Med. 372, 661–663 (2015).
Cancer Genome Atlas Research Network. et al. Comprehensive, integrative genomic analysis of diffuse lower-grade gliomas. N. Engl. J. Med. 372, 2481–2498 (2015).
Roylance, R. et al. Relationship of extreme chromosomal instability with long-term survival in a retrospective analysis of primary breast cancer. Cancer Epidemiol. Biomarkers Prev. 20, 2183–2194 (2011).
Birkbak, N.J. et al. Paradoxical relationship between chromosomal instability and survival outcome in cancer. Cancer Res. 71, 3447–3452 (2011).
Bochtler, T. et al. Clonal heterogeneity as detected by metaphase karyotyping is an indicator of poor prognosis in acute myeloid leukemia. J. Clin. Oncol. 31, 3898–3905 (2013).
Merlo, L.M.F. et al. A comprehensive survey of clonal diversity measures in Barrett’s esophagus as biomarkers of progression to esophageal adenocarcinoma. Cancer Prev. Res. (Phila.) 3, 1388–1397 (2010).
Maley, C.C. et al. Genetic clonal diversity predicts progression to esophageal adenocarcinoma. Nat. Genet. 38, 468–473 (2006).
Cibulskis, K. et al. Sensitive detection of somatic point mutations in impure and heterogeneous cancer samples. Nat. Biotechnol. 31, 213–219 (2013).
Roth, A. et al. JointSNVMix: a probabilistic model for accurate detection of somatic mutations in normal-tumor–paired next-generation sequencing data. Bioinformatics 28, 907–913 (2012).
Sathirapongsasuti, J.F. et al. Exome sequencing–based copy-number variation and loss of heterozygosity detection: ExomeCNV. Bioinformatics 27, 2648–2654 (2011).
Andor, N., Harness, J.V., Müller, S., Mewes, H.W. & Petritsch, C. EXPANDS: expanding ploidy and allele frequency on nested subpopulations. Bioinformatics 30, 50–60 (2014).
Roth, A. et al. PyClone: statistical inference of clonal population structure in cancer. Nat. Methods 11, 396–398 (2014).
Goode, A., Gilbert, B., Harkes, J., Jukic, D. & Satyanarayanan, M. OpenSlide: a vendor-neutral software foundation for digital pathology. J. Pathol. Inform. 4, 27 (2013).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Huang, D.W., Sherman, B.T. & Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Huang, D.W., Sherman, B.T. & Lempicki, R.A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009).
Griffith, M. et al. DGIdb: mining the druggable genome. Nat. Methods 10, 1209–1210 (2013).
Swanton, C. Cancer evolution constrained by mutation order. N. Engl. J. Med. 372, 661–663 (2015).
Kandoth, C. et al. Mutational landscape and significance across 12 major cancer types. Nature 502, 333–339 (2013).
Acknowledgements
This work was supported in part by the US National Institutes of Health (NIH) (grant no. P01 CA91955 (C.C.M.), R01 CA149566 (C.C.M.), R01 CA170595 (C.C.M.), R01 CA185138 (C.C.M.), R01 CA140657 (C.C.M.), P01 HG000205 (H.P.J.), U01CA151920 (H.P.J.), U01CA17629901 (H.P.J.), R01 HG006137 (H.P.J.), R01 CA164746 (C.P.), R01 NS08061904 (C.P.) and R01 HG006137 (L.C.X.)). Additional support to C.C.M. came from the Breast Cancer Research Program Breakthrough Award (award no. BC132057), a Congressionally Directed Medical Research Program (CDMRP). Additional support to H.P.J. came from the Doris Duke Clinical Foundation Clinical Scientist Development Award, a Research Scholar Grant from the American Cancer Society (award no. RSG-13-297-01-TBG) and a Howard Hughes Medical Institute Early Career Grant. N.A. was supported by awards from the Don and Ruth Seiler Fund and the National Cancer Institute (NCI) Cancer Target Discovery and Development (CTDD) Consortium (grant no. U01CA17629901). T.A.G. was supported by the Higher Education Founding Council for England (HEFCE). We are grateful to W. Mewes for advice on the presentation of our results and for insightful discussions about their implications; S.T. Jensen for advice on statistical data analysis; and C.W. Turck and M. Oft for reviewing the manuscript. The results presented here are in part based upon data generated by TCGA Research Network. We thank Hoffmann H. (University of Bonn, Germany) for the availability of the MATLAB function 'violin' that we used to generate the violin plots for the distribution of clone numbers and clone sizes.
Author information
Authors and Affiliations
Contributions
N.A. developed analytic methods, analyzed data and wrote the manuscript. T.A.G. developed analytic methods, gave technical support and conceptual advice, and wrote the manuscript. M.J. analyzed the histopathology images and provided advice on data visualization and interpretation. L.C.X. provided advice on the choice of statistical methods and the design of the statistical analysis. C.A.A. gave technical support and conceptual advice. C.C.M. developed analytic methods, wrote the manuscript and supervised the project. H.P.J. wrote the manuscript and supervised the project. C.P. supervised the project. All authors edited the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13, Supplementary Notes 1–4, & Supplementary Tables 1–9 (PDF 12015 kb)
Rights and permissions
About this article
Cite this article
Andor, N., Graham, T., Jansen, M. et al. Pan-cancer analysis of the extent and consequences of intratumor heterogeneity. Nat Med 22, 105–113 (2016). https://doi.org/10.1038/nm.3984
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3984
This article is cited by
-
A whirl of radiomics-based biomarkers in cancer immunotherapy, why is large scale validation still lacking?
npj Precision Oncology (2024)
-
Multiregional transcriptomics identifies congruent consensus subtypes with prognostic value beyond tumor heterogeneity of colorectal cancer
Nature Communications (2024)
-
Intratumoral heterogeneity of Ki67 proliferation index outperforms conventional immunohistochemistry prognostic factors in estrogen receptor-positive HER2-negative breast cancer
Virchows Archiv (2024)
-
A pathway activity-based proteomic classifier stratifies prostate tumors into two subtypes
Clinical Proteomics (2023)
-
Mismatch repair deficiency is not sufficient to elicit tumor immunogenicity
Nature Genetics (2023)