Abstract
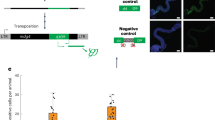
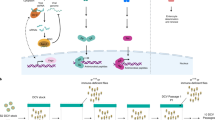
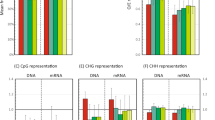
How persistent viral infections are established and maintained is widely debated and remains poorly understood. We found here that the persistence of RNA viruses in Drosophila melanogaster was achieved through the combined action of cellular reverse-transcriptase activity and the RNA-mediated interference (RNAi) pathway. Fragments of diverse RNA viruses were reverse-transcribed early during infection, which resulted in DNA forms embedded in retrotransposon sequences. Those virus-retrotransposon DNA chimeras produced transcripts processed by the RNAi machinery, which in turn inhibited viral replication. Conversely, inhibition of reverse transcription hindered the appearance of chimeric DNA and prevented persistence. Our results identify a cooperative function for retrotransposons and antiviral RNAi in the control of lethal acute infection for the establishment of viral persistence.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Emerman, M. & Malik, H.S. Paleovirology–modern consequences of ancient viruses. PLoS Biol. 8, e1000301 (2010).
Goldstone, D.C. et al. Structural and functional analysis of prehistoric lentiviruses uncovers an ancient molecular interface. Cell Host Microbe 8, 248–259 (2010).
Goic, B. & Saleh, M.C. Living with the enemy: viral persistent infections from a friendly viewpoint. Curr. Opin. Microbiol. 15, 531–537 (2012).
Flegel, T.W. Update on viral accommodation, a model for host-viral interaction in shrimp and other arthropods. Dev. Comp. Immunol. 31, 217–231 (2007).
Roekring, S., Flegel, T.W., Malasit, P. & Kittayapong, P. Challenging successive mosquito generations with a densonucleosis virus yields progressive survival improvement but persistent, innocuous infections. Dev. Comp. Immunol. 30, 878–892 (2006).
Weaver, S.C. & Reisen, W.K. Present and future arboviral threats. Antiviral Res. 85, 328–345 (2010).
Rechavi, O., Minevich, G. & Hobert, O. Transgenerational inheritance of an acquired small RNA-based antiviral response in C. elegans. Cell 147, 1248–1256 (2011).
Dasgupta, R. et al. Replication of flock house virus in three genera of medically important insects. J. Med. Entomol. 44, 102–110 (2007).
Dasgupta, R., Selling, B. & Rueckert, R. Flock house virus: a simple model for studying persistent infection in cultured Drosophila cells. Arch. Virol. (suppl. 9), 121–132 (1994).
Li, H., Li, W.X. & Ding, S.W. Induction and suppression of RNA silencing by an animal virus. Science 296, 1319–1321 (2002).
Jovel, J. & Schneemann, A. Molecular characterization of Drosophila cells persistently infected with Flock House virus. Virology 419, 43–53 (2011).
Ebner, P.D., Kim, S.K. & O'Callaghan, D.J. Biological and genotypic properties of defective interfering particles of equine herpesvirus 1 that mediate persistent infection. Virology 381, 98–105 (2008).
Tsai, K.N., Tsang, S.F., Huang, C.H. & Chang, R.Y. Defective interfering RNAs of Japanese encephalitis virus found in mosquito cells and correlation with persistent infection. Virus Res. 124, 139–150 (2007).
Simon, A.E., Roossinck, M.J. & Havelda, Z. Plant virus satellite and defective interfering RNAs: new paradigms for a new century. Annu. Rev. Phytopathol. 42, 415–437 (2004).
Huang, A.S. & Baltimore, D. Defective viral particles and viral disease processes. Nature 226, 325–327 (1970).
Marriott, A.C. & Dimmock, N.J. Defective interfering viruses and their potential as antiviral agents. Rev. Med. Virol. 20, 51–62 (2010).
Vodovar, N., Goic, B., Blanc, H. & Saleh, M.C. In silico reconstruction of viral genomes from small RNAs improves virus-derived small interfering RNA profiling. J. Virol. 85, 11016–11021 (2011).
Flynt, A., Liu, N., Martin, R. & Lai, E.C. Dicing of viral replication intermediates during silencing of latent Drosophila viruses. Proc. Natl. Acad. Sci. USA 106, 5270–5275 (2009).
Galiana-Arnoux, D., Dostert, C., Schneemann, A., Hoffmann, J.A. & Imler, J.L. Essential function in vivo for Dicer-2 in host defense against RNA viruses in Drosophila. Nat. Immunol. 7, 590–597 (2006).
van Rij, R.P. et al. The RNA silencing endonuclease Argonaute 2 mediates specific antiviral immunity in Drosophila melanogaster. Genes Dev. 20, 2985–2995 (2006).
Wang, X.H. et al. RNA interference directs innate immunity against viruses in adult Drosophila. Science 312, 452–454 (2006).
Belyi, V.A., Levine, A.J. & Skalka, A.M. Unexpected inheritance: multiple integrations of ancient bornavirus and ebolavirus/marburgvirus sequences in vertebrate genomes. PLoS Pathog. 6, e1001030 (2010).
Crochu, S. et al. Sequences of flavivirus-related RNA viruses persist in DNA form integrated in the genome of Aedes spp. mosquitoes. J. Gen. Virol. 85, 1971–1980 (2004).
Geuking, M.B. et al. Recombination of retrotransposon and exogenous RNA virus results in nonretroviral cDNA integration. Science 323, 393–396 (2009).
Klenerman, P., Hengartner, H. & Zinkernagel, R.M. A non-retroviral RNA virus persists in DNA form. Nature 390, 298–301 (1997).
Maori, E., Tanne, E. & Sela, I. Reciprocal sequence exchange between non-retro viruses and hosts leading to the appearance of new host phenotypes. Virology 362, 342–349 (2007).
Chiba, S. et al. Widespread endogenization of genome sequences of non-retroviral RNA viruses into plant genomes. PLoS Pathog. 7, e1002146 (2011).
Horie, M. et al. Endogenous non-retroviral RNA virus elements in mammalian genomes. Nature 463, 84–87 (2010).
Katzourakis, A. & Gifford, R.J. Endogenous viral elements in animal genomes. PLoS Genet. 6, e1001191 (2011).
Liu, H. et al. Widespread horizontal gene transfer from double-stranded RNA viruses to eukaryotic nuclear genomes. J. Virol. 84, 11876–11887 (2010).
Li, Y. & Ball, L.A. Nonhomologous RNA recombination during negative-strand synthesis of flock house virus RNA. J. Virol. 67, 3854–3860 (1993).
Zhong, W., Dasgupta, R. & Rueckert, R. Evidence that the packaging signal for nodaviral RNA2 is a bulged stem-loop. Proc. Natl. Acad. Sci. USA 89, 11146–11150 (1992).
Wu, Q. et al. Virus discovery by deep sequencing and assembly of virus-derived small silencing RNAs. Proc. Natl. Acad. Sci. USA 107, 1606–1611 (2010).
Eickbush, T.H. & Jamburuthugoda, V.K. The diversity of retrotransposons and the properties of their reverse transcriptases. Virus Res. 134, 221–234 (2008).
Terzian, C., Pelisson, A. & Bucheton, A. Evolution and phylogeny of insect endogenous retroviruses. BMC Evol. Biol. 1, 3 (2001).
Wu, Y. & Marsh, J.W. Selective transcription and modulation of resting T cell activity by preintegrated HIV DNA. Science 293, 1503–1506 (2001).
Saleh, M.C. et al. Antiviral immunity in Drosophila requires systemic RNA interference spread. Nature 458, 346–350 (2009).
Teixeira, L., Ferreira, A. & Ashburner, M. The bacterial symbiont Wolbachia induces resistance to RNA viral infections in Drosophila melanogaster. PLoS Biol. 6, e2 (2008).
Bian, G., Xu, Y., Lu, P., Xie, Y. & Xi, Z. The endosymbiotic bacterium Wolbachia induces resistance to dengue virus in Aedes aegypti. PLoS Pathog. 6, e1000833 (2010).
Flegel, T.W. Hypothesis for heritable, anti-viral immunity in crustaceans and insects. Biol. Direct 4, 32 (2009).
Killip, M.J., Young, D.F., Precious, B.L., Goodbourn, S. & Randall, R.E. Activation of the β interferon promoter by paramyxoviruses in the absence of virus protein synthesis. J. Gen. Virol. 93, 299–307 (2012).
Reed, L.J. & Muench, H. A simple method of estimating fifty per cent endpoints. Am. J. Hyg. 27, 493–497 (1938).
Cherry, S. & Perrimon, N. Entry is a rate-limiting step for viral infection in a Drosophila melanogaster model of pathogenesis. Nat. Immunol. 5, 81–87 (2004).
Gausson, V. & Saleh, M.C. Viral small RNA cloning and sequencing. Methods Mol. Biol. 721, 107–122 (2011).
Vagin, V.V. et al. A distinct small RNA pathway silences selfish genetic elements in the germline. Science 313, 320–324 (2006).
Acknowledgements
We thank members of the Saleh, Antoniewski and Vignuzzi laboratories for discussions and technical support; M. Vignuzzi, J. Chandler and R. van Rij for critical reading of the manuscript; A. Schneemann (The Scripps Research Institute) for antibody to FHV; P. Vargas for assistance with flow cytometry; and A. Pelisson for advice on retrotransposition. Supported by the French Agence Nationale de la Recherche (ANR-09-JCJC-0045-01 to M.-C.S.), the European Research Council (FP7/2007-2013 ERC 242703 to M.-C.S. and ERC 243312 to G.C.), Institut National de la Santé et de la Recherche Médicale and the Institut National du Cancer (G.C.), the French Ministry of Research (C.M.), La Fondation ARC pour la Recherche sur le Cancer (C.M.) and the AXA Research Fund (J.A.M.).
Author information
Authors and Affiliations
Contributions
B.G. and M.-C.S. designed the experiments, discussed the interpretation of the results and wrote the manuscript; G.C. participated in interpreting the data and writing the manuscript; B.G., N.V., J.A.M. and V.G. did research; H.B. generated the genomic DNA and the small-RNA libraries; N.V. and L.F. contributed to the bioinformatics analysis; and C.M., J.V.-O. and G.C. did the in vitro analysis of the reverse-transcriptase activity and AZT inhibition.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6 and Table 1 (PDF 656 kb)
Rights and permissions
About this article
Cite this article
Goic, B., Vodovar, N., Mondotte, J. et al. RNA-mediated interference and reverse transcription control the persistence of RNA viruses in the insect model Drosophila. Nat Immunol 14, 396–403 (2013). https://doi.org/10.1038/ni.2542
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2542
This article is cited by
-
In-depth transcriptomic analysis of Anopheles gambiae hemocytes uncovers novel genes and the oenocytoid developmental lineage
BMC Genomics (2024)
-
Virome and nrEVEome diversity of Aedes albopictus mosquitoes from La Reunion Island and China
Virology Journal (2022)
-
Plant and animal small RNA communications between cells and organisms
Nature Reviews Molecular Cell Biology (2022)
-
Establishing RNAi for basic research and pest control and identification of the most efficient target genes for pest control: a brief guide
Frontiers in Zoology (2021)
-
A survey of RNA viruses in mosquitoes from Mozambique reveals novel genetic lineages of flaviviruses and phenuiviruses, as well as frequent flavivirus-like viral DNA forms in Mansonia
BMC Microbiology (2020)