Abstract
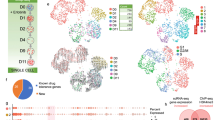
To elucidate the genomics of cellular responses to cancer treatment, we analyzed the expression of over 9,600 human genes in acute lymphoblastic leukemia cells before and after in vivo treatment with methotrexate and mercaptopurine given alone or in combination. Based on changes in gene expression, we identified 124 genes that accurately discriminated among the four treatments. Discriminating genes included those involved in apoptosis, mismatch repair, cell cycle control and stress response. Only 14% of genes that changed when these medications were given as single agents also changed when they were given together. These data indicate that lymphoid leukemia cells of different molecular subtypes share common pathways of genomic response to the same treatment, that changes in gene expression are treatment-specific and that gene expression can illuminate differences in cellular response to drug combinations versus single agents.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Pui, C.H. & Evans, W.E. Acute lymphoblastic leukemia. N. Engl. J. Med. 339, 605–615 (1998).
Evans, W.E. & Relling, M.V. Pharmacogenomics: translating functional genomics into rational therapeutics. Science 286, 487–491 (1999).
Evans, W.E. & McLeod, H.L. Pharmacogenomics—drug disposition, drug targets, and side effects. N. Engl. J. Med. 348, 538–549 (2003).
Pui, C.H., Campana, D. & Evans, W.E. Childhood acute lymphoblastic leukaemia—current status and future perspectives. Lancet Oncol. 2, 597–607 (2001).
Golub, T.R. et al. Molecular classification of cancer: class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Armstrong, S.A. et al. MLL translocations specify a distinct gene expression profile that distinguishes a unique leukemia. Nat. Genet. 30, 41–47 (2002).
Ferrando, A.A. et al. Gene expression signatures define novel oncogenic pathways in T cell acute lymphoblastic leukemia. Cancer Cell 1, 75–87 (2002).
Yeoh, E.-J. et al. Classification, subtype discovery, and prediction of outcome in pediatric acute lymphoblastic leukemia by gene expression profiling. Cancer Cell 1, 133–143 (2002).
Scherf, U. et al. A gene expression database for the molecular pharmacology of cancer. Nat. Genet. 24, 236–244 (2000).
Tsurusawa, M., Saeki, K. & Fujimoto, T. Differential induction of apoptosis on human lymphoblastic leukemia Nalm-6 and Molt-4 cells by various antitumor drugs. Int. J. Hematol. 66, 79–88 (1997).
Elion, G.B. The purine path to chemotherapy. Science 244, 41–47 (1989).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Kastan, M.B. & Lim, D.S. The many substrates and functions of ATM. Nat. Rev. Mol. Cell Biol. 1, 179–186 (2000).
Baskaran, R. et al. Ataxia telangiectasia mutant protein activates c-Abl tyrosine kinase in response to ionizing radiation. Nature 387, 516–519 (1997).
Li, J.C. & Kaminskas, E. Accumulation of DNA strand breaks and methotrexate cytotoxicity. Proc. Natl. Acad. Sci. USA 81, 5694–5698 (1984).
Lorico, A. et al. Accumulation of DNA strand breaks in cells exposed to methotrexate or N10-propargyl-5,8-dideazafolic acid. Cancer Res. 48, 2036–2041 (1988).
Nelson, W.G. & Kastan, M.B. DNA strand breaks: the DNA template alterations that trigger p53-dependent DNA damage response pathways. Mol. Cell Biol. 14, 1815–1823 (1994).
Bakkenist, C.J. & Kastan, M.B. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421, 499–506 (2003).
Wijnen, J. et al. Majority of hMLH1 mutations responsible for hereditary nonpolyposis colorectal cancer cluster at the exonic region 15–16. Am. J. Hum. Genet. 58, 300–307 (1996).
Palmirotta, R. et al. Transcripts with splicings of exons 15 and 16 of the hMLH1 gene in normal lymphocytes: implications in RNA-based mutation screening of hereditary non-polyposis colorectal cancer. Eur. J. Cancer 34, 927–930 (1998).
Gong, J.G. et al. The tyrosine kinase c-Abl regulates p73 in apoptotic response to cisplatin-induced DNA damage. Nature 399, 806–809 (1999).
Swann, P.F. et al. Role of postreplicative DNA mismatch repair in the cytotoxic action of thioguanine. Science 273, 1109–1111 (1996).
Marton, M.J. et al. Drug target validation and identification of secondary drug target effects using DNA microarrays. Nat. Med. 4, 1293–1301 (1998).
Sotiriou, C. et al. Gene expression profiles derived from fine needle aspiration correlate with response to systemic chemotherapy in breast cancer. Breast Cancer Res. 4, R3 (2002).
Bokkerink, J.P. et al. 6-Mercaptopurine: cytotoxicity and biochemical pharmacology in human malignant T-lymphoblasts. Biochem. Pharmacol. 45, 1455–1463 (1993).
Gorlick, R. et al. Intrinsic and acquired resistance to methotrexate in acute leukemia. N. Engl. J. Med. 335, 1041–1048 (1996).
Masson, E. et al. Accumulation of methotrexate polyglutamates in lymphoblasts is a determinant of antileukemic effects in vivo. A rationale for high-dose methotrexate. J. Clin. Invest. 97, 73–80 (1996).
Synold, T.W. et al. Blast cell methotrexate-polyglutamate accumulation in vivo differs by lineage, ploidy, and methotrexate dose in acute lymphoblastic leukemia. J. Clin. Invest. 94, 1996–2001 (1994).
Lipshutz, R.J., Fodor, S.P., Gingeras, T.R. & Lockhart, D.J. High density synthetic oligonucleotide arrays. Nat. Genet. 21, 20–24 (1999).
Acknowledgements
The authors gratefully acknowledge the technical support of K. Brown, C. Ding, J. Morris, D. Patel, M. Shipman and M. Wilkinson, and we thank M. Caldwell and N. Kornegay for help in preparing the manuscript and establishing our research databases. The authors also thank C. Sherr, J. Cleveland, T. Curran, B. Schulman and M. Kastan for providing critical feedback, S. Shurtleff for contributions to gene expression analysis and R. Ashmun for flow cytometric analysis. This work was supported by grants from the US National Institutes of Health to W.E.E., M.V.R. and J.R.D., by a Cancer Center Support Grant from the US National Cancer Institute, by a F.M. Kirby Clinical Research Professorship from the American Cancer Society to C.H.P., by a stipend from the Dr. Hilmer Foundation, German Science Foundation to M.H.C., and by the American Lebanese Syrian Associated Charities.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Cheok, M., Yang, W., Pui, CH. et al. Treatment-specific changes in gene expression discriminate in vivo drug response in human leukemia cells. Nat Genet 34, 85–90 (2003). https://doi.org/10.1038/ng1151
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng1151
This article is cited by
-
Early changes in gene expression profiles in AML patients during induction chemotherapy
BMC Genomics (2022)
-
Multi-view feature selection for identifying gene markers: a diversified biological data driven approach
BMC Bioinformatics (2020)
-
Integrative genomic analyses reveal mechanisms of glucocorticoid resistance in acute lymphoblastic leukemia
Nature Cancer (2020)
-
Gene expression signatures and ex vivo drug sensitivity profiles in children with acute lymphoblastic leukemia
Journal of Applied Genetics (2012)
-
Peripheral blood gene expression profiles in metabolic syndrome, coronary artery disease and type 2 diabetes
Genes & Immunity (2011)