Abstract
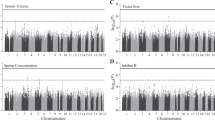
Non-obstructive azoospermia (NOA) is one of the most severe forms of male infertility. Its pathophysiology is largely unknown, and few genetic influences have been defined. To identify common variants contributing to NOA in Han Chinese men, we performed a three-stage genome-wide association study of 2,927 individuals with NOA and 5,734 controls. The combined analyses identified significant (P < 5.0 × 10−8) associations between NOA risk and common variants near PRMT6 (rs12097821 at 1p13.3: odds ratio (OR) = 1.25, P = 5.7 × 10−10), PEX10 (rs2477686 at 1p36.32: OR = 1.39, P = 5.7 × 10−12) and SOX5 (rs10842262 at 12p12.1: OR = 1.23, P = 2.3 × 10−9). These findings implicate genetic variants at 1p13.3, 1p36.32 and 12p12.1 in the etiology of NOA in Han Chinese men.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Hirsh, A. Male subfertility. Br. Med. J. 327, 669–672 (2003).
Maduro, M.R. & Lamb, D.J. Understanding new genetics of male infertility. J. Urol. 168, 2197–2205 (2002).
Bhasin, S., de Kretser, D.M. & Baker, H.W. Clinical review 64: pathophysiology and natural history of male infertility. J. Clin. Endocrinol. Metab. 79, 1525–1529 (1994).
Wu, B. et al. A frequent Y chromosome b2/b3 subdeletion shows strongly association with male infertility in Han-Chinese population. Hum. Reprod. 22, 1107–1113 (2007).
Ferlin, A. et al. Male infertility: role of genetic background. Reprod. Biomed. Online 14, 734–745 (2007).
Huynh, T., Mollard, R. & Trounson, A. Selected genetic factors associated with male infertility. Hum. Reprod. Update 8, 183–198 (2002).
O'Flynn O'Brien, K.L., Varghese, A.C. & Agarwal, A. The genetic causes of male factor infertility: a review. Fertil. Steril. 93, 1–12 (2010).
Dohle, G.R. et al. Genetic risk factors in infertile men with severe oligozoospermia and azoospermia. Hum. Reprod. 17, 13–16 (2002).
Bashamboo, A. et al. Human male infertility associated with mutations in NR5A1 encoding steroidogenic factor 1. Am. J. Hum. Genet. 87, 505–512 (2010).
Matzuk, M.M. & Lamb, D.J. The biology of infertility: research advances and clinical challenges. Nat. Med. 14, 1197–1213 (2008).
Okada, H. et al. Genome-wide expression of azoospermia testes demonstrates a specific profile and implicates ART3 in genetic susceptibility. PLoS Genet. 4, e26 (2008).
Aston, K.I. & Carrell, D.T. Genome-wide study of single-nucleotide polymorphisms associated with azoospermia and severe oligozoospermia. J. Androl. 30, 711–725 (2009).
Aston, K.I., Krausz, C., Laface, I., Ruiz-Castané, E. & Carrell, D.T. Evaluation of 172 candidate polymorphisms for association with oligozoospermia or azoospermia in a large cohort of men of European descent. Hum. Reprod. 25, 1383–1397 (2010).
Gonsalvez, G.B., Rajendra, T.K., Tian, L. & Matera, A.G. The Sm-protein methyltransferase, dart5, is essential for germ-cell specification and maintenance. Curr. Biol. 16, 1077–1089 (2006).
Anne, J., Ollo, R., Ephrussi, A. & Mechler, B.M. Arginine methyltransferase Capsuleen is essential for methylation of spliceosomal Sm proteins and germ cell formation in Drosophila. Development 134, 137–146 (2007).
Ancelin, K. et al. Blimp1 associates with Prmt5 and directs histone arginine methylation in mouse germ cells. Nat. Cell Biol. 8, 623–630 (2006).
Chen, W., Cao, M., Yang, Y., Nagahama, Y. & Zhao, H. Expression pattern of prmt5 in adult fish and embryos of medaka, Oryzias latipes. Fish Physiol. Biochem. 35, 325–332 (2009).
El-Andaloussi, N. et al. Arginine methylation regulates DNA polymerase β. Mol. Cell 22, 51–62 (2006).
Sobol, R.W. et al. Requirement of mammalian DNA polymerase-β in base-excision repair. Nature 379, 183–186 (1996).
Olsen, A.K. et al. Highly efficient base excision repair (BER) in human and rat male germ cells. Nucleic Acids Res. 29, 1781–1790 (2001).
Plug, A.W., Clairmont, C.A., Sapi, E., Ashley, T. & Sweasy, J.B. Evidence for a role for DNA polymerase β in mammalian meiosis. Proc. Natl. Acad. Sci. USA 94, 1327–1331 (1997).
Chen, H., Liu, Z. & Huang, X. Drosophila models of peroxisomal biogenesis disorder: peroxins are required for spermatogenesis and very-long-chain fatty acid metabolism. Hum. Mol. Genet. 19, 494–505 (2010).
Carpentier, M. et al. Reduced fertility in male mice deficient in the zinc metallopeptidase NL1. Mol. Cell. Biol. 24, 4428–4437 (2004).
Wunderle, V.M., Critcher, R., Ashworth, A. & Goodfellow, P.N. Cloning and characterization of SOX5, a new member of the human SOX gene family. Genomics 36, 354–358 (1996).
Denny, P., Swift, S., Connor, F. & Ashworth, A. An SRY-related gene expressed during spermatogenesis in the mouse encodes a sequence-specific DNA-binding protein. EMBO J. 11, 3705–3712 (1992).
Budde, L.M., Wu, C., Tilman, C., Douglas, I. & Ghosh, S. Regulation of IκBβ expression in testis. Mol. Biol. Cell 13, 4179–4194 (2002).
Fröjdman, K., Harley, V.R. & Pelliniemi, L.J. Sox9 protein in rat Sertoli cells is age and stage dependent. Histochem. Cell Biol. 113, 31–36 (2000).
O'Bryan, M.K. et al. Sox8 is a critical regulator of adult Sertoli cell function and male fertility. Dev. Biol. 316, 359–370 (2008).
Zafarana, G. et al. Coamplification of DAD-R, SOX5, and EKI1 in human testicular seminomas, with specific overexpression of DAD-R, correlates with reduced levels of apoptosis and earlier clinical manifestation. Cancer Res. 62, 1822–1831 (2002).
Takenaka, K. et al. Polymorphism in Sirpa modulates engraftment of human hematopoietic stem cells. Nat. Immunol. 8, 1313–1323 (2007).
Lu, C. et al. The b2/b3 subdeletion shows higher risk of spermatogenic failure and higher frequency of complete AZFc deletion than the gr/gr subdeletion in a Chinese population. Hum. Mol. Genet. 18, 1122–1130 (2009).
Dimas, A.S. et al. Common regulatory variation impacts gene expression in a cell type–dependent manner. Science 325, 1246–1250 (2009).
Hu, Z. et al. A genome-wide association study identifies two new lung cancer susceptibility loci at 13q12.12 and 22q12.2 in Han Chinese. Nat. Genet. 43, 792–796 (2011).
Purdue, M.P. et al. Genome-wide association study of renal cell carcinoma identifies two susceptibility loci on 2p21 and 11q13.3. Nat. Genet. 43, 60–65 (2011).
Price, A.L. et al. Principal components analysis corrects for stratification in genome-wide association studies. Nat. Genet. 38, 904–909 (2006).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
Barrett, J.C. et al. Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21, 263–265 (2005).
Dixon, A.L. et al. A genome-wide association study of global gene expression. Nat. Genet. 39, 1202–1207 (2007).
Stranger, B.E. et al. Population genomics of human gene expression. Nat. Genet. 39, 1217–1224 (2007).
Veyrieras, J.B. et al. High-resolution mapping of expression-QTLs yields insight into human gene regulation. PLoS Genet. 4, e1000214 (2008).
Pickrell, J.K. et al. Understanding mechanisms underlying human gene expression variation with RNA sequencing. Nature 464, 768–772 (2010).
Montgomery, S.B. et al. Transcriptome genetics using second generation sequencing in a Caucasian population. Nature 464, 773–777 (2010).
Dimas, A.S. et al. Common regulatory variation impacts gene expression in a cell type–dependent manner. Science 325, 1246–1250 (2009).
Zeller, T. et al. Genetics and beyond—the transcriptome of human monocytes and disease susceptibility. PLoS ONE 5, e10693 (2010).
Dimas, A.S. et al. Common regulatory variation impacts gene expression in a cell type–dependent manner. Science 325, 1246–1250 (2009).
Schadt, E.E. et al. Mapping the genetic architecture of gene expression in human liver. PLoS Biol. 6, e107 (2008).
Myers, A.J. et al. A survey of genetic human cortical gene expression. Nat. Genet. 39, 1494–1499 (2007).
Acknowledgements
The authors wish to thank all the study participants, research staff and students who participated in this work. We thank Q. Wei (MD Anderson Cancer Center) for editing the manuscript. This work was funded by the National Key Basic Research Program Grant (2011CB944304) and the China National High-Tech Research and Development Program Grant (2009AA022705) and partly by a Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Contributions
J.S., X.W. and H.S. directed the study, obtained financial support and were responsible for study design, interpretation of results and manuscript writing. Z.H. performed overall project management along with Y.X. and X.G., performed statistical analyses along with J.D. and drafted the initial manuscript. Y.J., F.L. and H.L. were responsible for sample processing and managed the genotyping data. X.Y., Y.W., J.L., B.Y., C.L. and Z.Z. were responsible for subject recruitment and sample preparation for the Nanjing samples. H.L., Y.G. and C.X. were responsible for subject recruitment and sample preparation for the Wuhan samples. H.H. and Z.L. were responsible for subject recruitment and sample preparation for the Shanghai samples. All authors approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1 and 2 and Supplementary Tables 1–7. (PDF 965 kb)
Rights and permissions
About this article
Cite this article
Hu, Z., Xia, Y., Guo, X. et al. A genome-wide association study in Chinese men identifies three risk loci for non-obstructive azoospermia. Nat Genet 44, 183–186 (2012). https://doi.org/10.1038/ng.1040
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.1040
This article is cited by
-
Genome-wide assessment and mapping of inbreeding depression identifies candidate genes associated with semen traits in Holstein bulls
BMC Genomics (2023)
-
A novel tumor suppressor encoded by a 1p36.3 lncRNA functions as a phosphoinositide-binding protein repressing AKT phosphorylation/activation and promoting autophagy
Cell Death & Differentiation (2023)
-
Phase-separated CCER1 coordinates the histone-to-protamine transition and male fertility
Nature Communications (2023)
-
Immune and spermatogenesis-related loci are involved in the development of extreme patterns of male infertility
Communications Biology (2022)
-
Replicating a GWAS: two novel candidate markers for oligospermia in Greek population
Molecular Biology Reports (2021)