Abstract
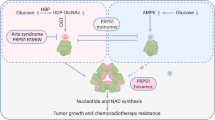
Lower glycolysis involves a series of reversible reactions, which interconvert intermediates that also feed anabolic pathways. 3-phosphoglycerate (3-PG) is an abundant lower glycolytic intermediate that feeds serine biosynthesis via the enzyme phosphoglycerate dehydrogenase, which is genomically amplified in several cancers. Phosphoglycerate mutase 1 (PGAM1) catalyzes the isomerization of 3-PG into the downstream glycolytic intermediate 2-phosphoglycerate (2-PG). PGAM1 needs to be histidine phosphorylated to become catalytically active. We show that the primary PGAM1 histidine phosphate donor is 2,3-bisphosphoglycerate (2,3-BPG), which is made from the glycolytic intermediate 1,3-bisphosphoglycerate (1,3-BPG) by bisphosphoglycerate mutase (BPGM). When BPGM is knocked out, 1,3-BPG can directly phosphorylate PGAM1. In this case, PGAM1 phosphorylation and activity are decreased, but nevertheless sufficient to maintain normal glycolytic flux and cellular growth rate. 3-PG, however, accumulates, leading to increased serine synthesis. Thus, one biological function of BPGM is controlling glycolytic intermediate levels and thereby serine biosynthetic flux.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Vander Heiden, M.G., Cantley, L.C. & Thompson, C.B. Understanding the Warburg effect: the metabolic requirements of cell proliferation. Science 324, 1029–1033 (2009).
Hitosugi, T. et al. Phosphoglycerate mutase 1 coordinates glycolysis and biosynthesis to promote tumor growth. Cancer Cell 22, 585–600 (2012).
Possemato, R. et al. Functional genomics reveal that the serine synthesis pathway is essential in breast cancer. Nature 476, 346–350 (2011).
Labuschagne, C.F., van den Broek, N.J., Mackay, G.M., Vousden, K.H. & Maddocks, O.D. Serine, but not glycine, supports one-carbon metabolism and proliferation of cancer cells. Cell Rep. 7, 1248–1258 (2014).
Mattaini, K.R. et al. An epitope tag alters phosphoglycerate dehydrogenase structure and impairs ability to support cell proliferation. Cancer Metab. 3, 5 (2015).
Samanta, D. et al. PHGDH expression is required for mitochondrial redox homeostasis, breast cancer stem cell maintenance, and lung metastasis. Cancer Res. 76, 4430–4442 (2016).
Fothergill-Gilmore, L.A. & Watson, H.C. The phosphoglycerate mutases. Adv. Enzymol. 62, 227–313 (1989).
Rose, Z.B. The enzymology of 2,3-bisphosphoglycerate. Adv. Enzymol. 51, 211–253 (1980).
Nelson, D.L., Lehninger, A.L. & Cox, M.M. Lehninger Principles of Biochemistry (W.H. Freeman, New York, 2008).
Sutherland, E.W., Posternak, T.Z. & Cori, C.F. The mechanism of action of phosphoglucomutase and phosphoglyceric acid mutase. J. Biol. Chem. 179, 501 (1949).
Rose, Z.B. The purification and properties of diphosphoglycerate mutase from human erythrocytes. J. Biol. Chem. 243, 4810–4820 (1968).
Narita, H., Yanagawa, S., Sasaki, R. & Chiba, H. Induction of 2,3-bisphosphoglycerate synthase in Friend leukemia cells. Biochem. Biophys. Res. Commun. 103, 90–96 (1981).
Joulin, V. et al. Isolation and characterization of the human 2,3-bisphosphoglycerate mutase gene. J. Biol. Chem. 263, 15785–15790 (1988).
Chiba, H. & Sasaki, R. Functions, of 2,3-bisphosphoglycerate and its metabolism. Curr. Top. Cell. Regul. 14, 75–116 (1978).
Sasaki, R., Ikura, K., Narita, H., Yanagawa, S.-i. & Chiba, H. 2,3-bisphosphoglycerate in erythroid cells. Trends Biochem. Sci. 7, 140–142 (1982).
Rigden, D.J., Walter, R.A., Phillips, S.E. & Fothergill-Gilmore, L.A. Sulphate ions observed in the 2.12 A structure of a new crystal form of S. cerevisiae phosphoglycerate mutase provide insights into understanding the catalytic mechanism. J. Mol. Biol. 286, 1507–1517 (1999).
White, M.F. & Fothergill-Gilmore, L.A. Mutase versus synthase: the phosphoglycerate mutase family studied by protein engineering. Biochem. Soc. Trans. 18, 257 (1990).
Hitosugi, T. et al. Tyr26 phosphorylation of PGAM1 provides a metabolic advantage to tumours by stabilizing the active conformation. Nat. Commun. 4, 1790 (2013).
Kee, J.M., Villani, B., Carpenter, L.R. & Muir, T.W. Development of stable phosphohistidine analogues. J. Am. Chem. Soc. 132, 14327–14329 (2010).
Kee, J.M., Oslund, R.C., Perlman, D.H. & Muir, T.W. A pan-specific antibody for direct detection of protein histidine phosphorylation. Nat. Chem. Biol. 9, 416–421 (2013).
Oslund, R.C. et al. A phosphohistidine proteomics strategy based on elucidation of a unique gas-phase phosphopeptide fragmentation mechanism. J. Am. Chem. Soc. 136, 12899–12911 (2014).
Kee, J.M., Oslund, R.C., Couvillon, A.D. & Muir, T.W. A second-generation phosphohistidine analog for production of phosphohistidine antibodies. Org. Lett. 17, 187–189 (2015).
Pritlove, D.C., Gu, M., Boyd, C.A., Randeva, H.S. & Vatish, M. Novel placental expression of 2,3-bisphosphoglycerate mutase. Placenta 27, 924–927 (2006).
Ran, F.A. et al. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 8, 2281–2308 (2013).
Johnson, C.M. & Price, N.C. Denaturation and renaturation of the monomeric phosphoglycerate mutase from Schizosaccharomyces pombe. Biochem. J. 245, 525–530 (1987).
Vander Heiden, M.G. et al. Evidence for an alternative glycolytic pathway in rapidly proliferating cells. Science 329, 1492–1499 (2010).
Moellering, R.E. & Cravatt, B.F. Functional lysine modification by an intrinsically reactive primary glycolytic metabolite. Science 341, 549–553 (2013).
Negelein, E. Synthesis, determination, analysis, and properties of 1,3-diphosphoglyceric acid. in Methods in Enzymology 3, 216–220 (1957).
Han, C.H. & Rose, Z.B. Active site phosphohistidine peptides from red cell bisphosphoglycerate synthase and yeast phosphoglycerate mutase. J. Biol. Chem. 254, 8836–8840 (1979).
Wang, Y. et al. Seeing the process of histidine phosphorylation in human bisphosphoglycerate mutase. J. Biol. Chem. 281, 39642–39648 (2006).
Fuhs, S.R. et al. Monoclonal 1- and 3-phosphohistidine antibodies: new tools to study histidine phosphorylation. Cell 162, 198–210 (2015).
Lu, W., Kimball, E. & Rabinowitz, J.D. A high-performance liquid chromatography-tandem mass spectrometry method for quantitation of nitrogen-containing intracellular metabolites. J. Am. Soc. Mass Spectrom. 17, 37–50 (2006).
Rose, Z.B. & Dube, S. Rates of phosphorylation and dephosphorylation of phosphoglycerate mutase and bisphosphoglycerate synthase. J. Biol. Chem. 251, 4817–4822 (1976).
Rose, Z.B. & Dube, S. Phosphoglycerate mutase. Kinetics and effects of salts on the mutase and bisphosphoglycerate phosphatase activities of the enzyme from chicken breast muscle. J. Biol. Chem. 253, 8583–8592 (1978).
Samanta, D. & Semenza, G.L. Serine synthesis helps hypoxic cancer stem cells regulate redox. Cancer Res. 76, 6458–6462 (2016).
Melamud, E., Vastag, L. & Rabinowitz, J.D. Metabolomic analysis and visualization engine for LC–MS data. Anal. Chem. 82, 9818–9826 (2010).
Clasquin, M.F., Melamud, E. & Rabinowitz, J.D. LC–MS data processing with MAVEN: a metabolomic analysis and visualization engine. Curr. Protoc. Bioinformatics 37, 14.11 (2011).
Shevchenko, A., Tomas, H., Havlis, J., Olsen, J.V. & Mann, M. In-gel digestion for mass spectrometric characterization of proteins and proteomes. Nat. Protoc. 1, 2856–2860 (2006).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
David, Y., Vila-Perelló, M., Verma, S. & Muir, T.W. Chemical tagging and customizing of cellular chromatin states using ultrafast trans-splicing inteins. Nat. Chem. 7, 394–402 (2015).
Brinkman, E.K., Chen, T., Amendola, M. & van Steensel, B. Easy quantitative assessment of genome editing by sequence trace decomposition. Nucleic Acids Res. 42, e168 (2014).
Sanjana, N.E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Methods 11, 783–784 (2014).
Park, J.O. et al. Metabolite concentrations, fluxes and free energies imply efficient enzyme usage. Nat. Chem. Biol. 12, 482–489 (2016).
Acknowledgements
We thank current and former members of the Muir and Rabinowitz laboratories for helpful discussions during the preparation of this manuscript and V. Suri of Raze Therapeutics for technical help. This work was funded by the US National Institutes of Health for T.W.M. (5R01GM095880) and J.D.R. (R01 CA163591 and P30DK019525), the US Department of Energy for J.D.R. (DE-SC0012461), Stand Up to Cancer for J.D.R. (SU2C-AACR-DT0509), and the Brewster Foundation and Breast Cancer Research Foundation for Y.K. R.C.O. and J.-M.K. were supported by postdoctoral fellowships from the National Institutes of Health Research Service Award (1F32CA167901) and Damon Runyon Cancer Research Foundation (DRG-2005-09), respectively.
Author information
Authors and Affiliations
Contributions
R.C.O. designed experiments, prepared materials, analyzed data, interpreted results, generated HEK 293T and HCT116 knockout cell lines, and conducted experiments for metabolomic data analysis, western blotting, cell growth measurements, and in vitro phosphorylation analysis. X.S. designed experiments, prepared materials, analyzed data, interpreted results, designed primers and performed all DNA sequencing analysis for BPGM knockout experiments, conducted all metabolomic experiments and data analysis, prepared 1,3-BPG, and derived and conducted all serine flux calculations. M.H. designed experiments, analyzed data, interpreted results, performed cell assays for metabolomic data analysis, performed western blots for BPGM rescue and PGAM1 knockdown experiments, and performed cell growth measurements of knockout cells with serine/glycine or oxygen depletion. J.-M.K. designed experiments, analyzed data, interpreted results, and performed western blot analysis of BPGM protein levels. M.E. generated MDA-MB-231 knockout cells, prepared PGAM1 knockdown cells, and designed, performed, and analyzed mouse xenograft experiments. Y.D. designed and conducted RNA-seq analysis of wild-type and knockout HEK 293T cells. B.W. designed and conducted 32P-labeling experiments, analyzed data, and interpreted results of metabolomic experiments. E.G. performed phosphorylated PGAM1 western blot analysis from multiple cell lines. D.H.P. performed LC–MS-based proteomic analysis of phosphorylated PGAM1, analyzed data, and interpreted results. T.W.M. designed experiments, analyzed data, and interpreted results. J.D.R. designed experiments, analyzed data, and interpreted results. R.C.O., X.S., M.H., T.W.M., and J.D.R. wrote the manuscript with contributions from all authors.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–28 (PDF 35287 kb)
Rights and permissions
About this article
Cite this article
Oslund, R., Su, X., Haugbro, M. et al. Bisphosphoglycerate mutase controls serine pathway flux via 3-phosphoglycerate. Nat Chem Biol 13, 1081–1087 (2017). https://doi.org/10.1038/nchembio.2453
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2453
This article is cited by
-
Bisphosphoglycerate mutase predicts myocardial dysfunction and adverse outcome in sepsis: an observational cohort study
BMC Infectious Diseases (2024)
-
Transcriptomic and metabolomic characterization of post-hatch metabolic reprogramming during hepatic development in the chicken
BMC Genomics (2021)
-
Serine synthesis through PHGDH coordinates nucleotide levels by maintaining central carbon metabolism
Nature Communications (2018)