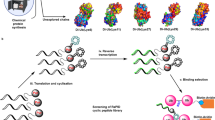
Abstract
Protein conformation and function are often inextricably linked, such that the states a protein adopts define its enzymatic activity or its affinity for various partners. Here we combine computational design with macromolecular display to isolate functional conformations of ubiquitin that tightly bind the catalytic core of the oncogenic ubiquitin-specific protease 7 (USP7) deubiquitinase. Structural and biochemical characterization of these ubiquitin variants suggest that remodeled backbone conformations and core packing poise these molecules for stronger interactions, leading to potent and specific inhibition of enzymatic activity. A ubiquitin variant expressed in human tumor cell lines binds and inhibits endogenous USP7, thereby enhancing Mdm2 proteasomal turnover and stabilizing p53. In sum, we have developed an approach to rationally target macromolecular libraries toward the remodeling of protein conformation, shown that engineering of ubiquitin conformation can greatly increase its interaction with deubiquitinases and developed powerful tools to probe the cellular role of USP7.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Change history
29 January 2013
In the version of this article initially published, the authors neglected to acknowledge an important collaborator and to include a citation of their related work. The error has been corrected in the HTML and PDF versions of the article.
References
Rizk, S.S. et al. Allosteric control of ligand-binding affinity using engineered conformation-specific effector proteins. Nat. Struct. Mol. Biol. (2011).
Gao, J., Sidhu, S.S. & Wells, J.A. Two-state selection of conformation-specific antibodies. Proc. Natl. Acad. Sci. USA 106, 3071–3076 (2009).
Lange, O.F. et al. Recognition dynamics up to microseconds revealed from an RDC-derived ubiquitin ensemble in solution. Science 320, 1471–1475 (2008).
Clague, M.J. & Urbé, S. Ubiquitin: same molecule, different degradation pathways. Cell 143, 682–685 (2010).
Pickart, C.M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Pickart, C.M. & Fushman, D. Polyubiquitin chains: polymeric protein signals. Curr. Opin. Chem. Biol. 8, 610–616 (2004).
Humphris, E.L. & Kortemme, T. Design of multi-specificity in protein interfaces. PLoS Comput. Biol. 3, e164 (2007).
Friedland, G.D., Lakomek, N.-A., Griesinger, C., Meiler, J. & Kortemme, T. A correspondence between solution-state dynamics of an individual protein and the sequence and conformational diversity of its family. PLoS Comput. Biol. 5, e1000393 (2009).
Komander, D., Clague, M.J. & Urbé, S. Breaking the chains: structure and function of the deubiquitinases. Nat. Rev. Mol. Cell Biol. 10, 550–563 (2009).
Nicholson, B. & Suresh Kumar, K.G. The multifaceted roles of USP7: new therapeutic opportunities. Cell Biochem. Biophys. 60, 61–68 (2011).
Hussain, S., Zhang, Y. & Galardy, P.J. DUBs and cancer: the role of deubiquitinating enzymes as oncogenes, non-oncogenes and tumor suppressors. Cell Cycle 8, 1688–1697 (2009).
Li, M., Brooks, C.L., Kon, N. & Gu, W. A dynamic role of HAUSP in the p53-Mdm2 pathway. Mol. Cell 13, 879–886 (2004).
Li, M. et al. Deubiquitination of p53 by HAUSP is an important pathway for p53 stabilization. Nature 416, 648–653 (2002).
Faesen, A.C. et al. Mechanism of USP7/HAUSP activation by its C-terminal ubiquitin-like domain and allosteric regulation by GMP-synthetase. Mol. Cell 44, 147–159 (2011).
Fernández-Montalván, A. et al. Biochemical characterization of USP7 reveals post-translational modification sites and structural requirements for substrate processing and subcellular localization. FEBS J. 274, 4256–4270 (2007).
Ernst, A. Science 10.1126/science.1230161 (3 January 2013).
Lowman, H.B. & Wells, J.A. Affinity maturation of human growth hormone by monovalent phage display. J. Mol. Biol. 234, 564–578 (1993).
Levin, A.M. et al. Exploiting a natural conformational switch to engineer an interleukin-2 'superkine'. Nature 484, 529–533 (2012).
Leaver-Fay, A. et al. ROSETTA3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 487, 545–574 (2011).
Hu, M. et al. Crystal structure of a UBP-family deubiquitinating enzyme in isolation and in complex with ubiquitin aldehyde. Cell 111, 1041–1054 (2002).
Yuan, J., Luo, K., Zhang, L., Cheville, J.C. & Lou, Z. USP10 regulates p53 localization and stability by deubiquitinating p53. Cell 140, 384–396 (2010).
Huang, O.W. et al. Phosphorylation-dependent activity of the deubiquitinase DUBA. Nat. Struct. Mol. Biol. 19, 171–175 (2012).
Reyes-Turcu, F.E. et al. The ubiquitin binding domain ZnF UBP recognizes the C-terminal diglycine motif of unanchored ubiquitin. Cell 124, 1197–1208 (2006).
Renatus, M. et al. Structural basis of ubiquitin recognition by the deubiquitinating protease USP2. Structure 14, 1293–1302 (2006).
Ganesan, R. et al. Unraveling the allosteric mechanism of serine protease inhibition by an antibody. Structure 17, 1614–1624 (2009).
Ultsch, M.H., Somers, W., Kossiakoff, A.A. & de Vos, A.M. The crystal structure of affinity-matured human growth hormone at 2-Å resolution. J. Mol. Biol. 236, 286–299 (1994).
Lazar, G.A., Johnson, E.C., Desjarlais, J.R. & Handel, T.M. Rotamer strain as a determinant of protein structural specificity. Protein Sci. 8, 2598–2610 (1999).
The Practice of Medicinal Chemistry, 2nd edn. (ed. Wermuth, C.G.) (Academic Press, 2003).
Giebel, L.B. et al. Screening of cyclic peptide phage libraries identifies ligands that bind streptavidin with high affinities. Biochemistry 34, 15430–15435 (1995).
O'Neil, K.T. et al. Identification of novel peptide antagonists for GPIIb/IIIa from a conformationally constrained phage peptide library. Proteins 14, 509–515 (1992).
Tzeng, S.-R. & Kalodimos, C.G. Dynamic activation of an allosteric regulatory protein. Nature 462, 368–372 (2009).
Tzeng, S.-R. & Kalodimos, C.G. Protein activity regulation by conformational entropy. Nature 488, 236–240 (2012).
Hammes, G.G., Benkovic, S.J. & Hammes-Schiffer, S. Flexibility, diversity, and cooperativity: pillars of enzyme catalysis. Biochemistry 50, 10422–10430 (2011).
Leaver-Fay, A. et al. ROSETTA3: an object-oriented software suite for the simulation and design of macromolecules. Methods Enzymol. 487, 545–574 (2011).
Dong, K.C. et al. Preparation of distinct ubiquitin chain reagents of high purity and yield. Structure 19, 1053–1063 (2011).
Tonikian, R., Zhang, Y., Boone, C. & Sidhu, S.S. Identifying specificity profiles for peptide recognition modules from phage-displayed peptide libraries. Nat. Protoc. 2, 1368–1386 (2007).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Lange, O.F., van der Spoel, D. & de Groot, B.L. Scrutinizing molecular mechanics force fields on the submicrosecond timescale with NMR data. Biophys. J. 99, 647–655 (2010).
Cai, M. et al. An efficient and cost-effective isotope labeling protocol for proteins expressed in Escherichia coli. J. Biomol. NMR 11, 97–102 (1998).
Hansen, D.F. et al. An exchange-free measure of 15N transverse relaxation: an NMR spectroscopy application to the study of a folding intermediate with pervasive chemical exchange. J. Am. Chem. Soc. 129, 11468–11479 (2007).
Bahrami, A., Assadi, A.H., Markley, J.L. & Eghbalnia, H.R. Probabilistic interaction network of evidence algorithm and its application to complete labeling of peak lists from protein NMR spectroscopy. PLoS Comput. Biol. 5, e1000307 (2009).
Findeisen, M., Brand, T. & Berger, S. A 1H-NMR thermometer suitable for cryoprobes. Magn. Reson. Chem. 45, 175–178 (2007).
Grzesiek, S. & Bax, A. The importance of not saturating water in protein NMR. Application to sensitivity enhancement and NOE measurements. J. Am. Chem. Soc. 115, 12593–12594 (1993).
Hansen, D.F., Feng, H., Zhou, Z., Bai, Y. & Kay, L.E. Selective characterization of microsecond motions in proteins by NMR relaxation. J. Am. Chem. Soc. 131, 16257–16265 (2009).
Delaglio, F. et al. NMRPipe: a multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 277–293 (1995).
Tollinger, M., Skrynnikov, N.R., Mulder, F.A., Forman-Kay, J.D. & Kay, L.E. Slow dynamics in folded and unfolded states of an SH3 domain. J. Am. Chem. Soc. 123, 11341–11352 (2001).
Mulder, F.A.A., Skrynnikov, N.R., Hon, B., Dahlquist, F.W. & Kay, L.E. Measurement of slow (μs-ms) time scale dynamics in protein side chains by 15N relaxation dispersion NMR spectroscopy: application to Asn and Gln residues in a cavity mutant of T4 lysozyme. J. Am. Chem. Soc. 123, 967–975 (2001).
Wertz, I.E. et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-κB signalling. Nature 430, 694–699 (2004).
Newton, K. et al. Using linkage-specific monoclonal antibodies to analyze cellular ubiquitylation. Methods Mol. Biol. 832, 185–196 (2012).
Matsumoto, M.L. et al. Engineering and structural characterization of a linear polyubiquitin-specific antibody. J. Mol. Biol. 10.1016/j.jmb.2011.12.053 (2011).
Matsumoto, M.L. et al. K11-linked polyubiquitination in cell cycle control revealed by a K11 linkage-specific antibody. Mol. Cell 39, 477–484 (2010).
Perkins, D.N., Pappin, D.J., Creasy, D.M. & Cottrell, J.S. Probability-based protein identification by searching sequence databases using mass spectrometry data. Electrophoresis 20, 3551–3567 (1999).
Acknowledgements
We thank the DNA synthesis and sequencing groups for invaluable assistance in creating and isolating variants. We also thank E. Dueber, B. Lazarus and W. Fairbrother for critical reading of the manuscript and I. Chen for assistance in purifying U7Ub7.
We apologize for inadvertently omitting the acknowledgment of the following important collaborators. We thank S. Sidhu and A. Ernst for advice on ubiquitin surface library design and USP7 protein production. We also thank M. Kwok, Y. Franke and K. Bowman for the USP7cd* construct. A manuscript describing the use of surface-engineered ubiquitin variants to inhibit ubiquitin-signaling enzymes has recently been published, and we have added a citation to this work.
Author information
Authors and Affiliations
Contributions
Y.Z. and L.Z. performed mutagenesis, protein expression and purification, phage display, phage ELISA, protein ELISA, biolayer interferometry and enzymatic assays. L.R. and J.M.M. crystallized proteins and solved structures. A.H.P. performed NMR analysis. P.L. and W.S. performed MS. E.H. expressed and purified proteins. I.E.W. and C.L. performed the cellular experiments. J.E.C. performed computational design and modeling and measured affinities by ITC. All authors designed, performed and analyzed experiments. All authors contributed to writing the paper, with Y.Z. and J.E.C. coordinating.
Corresponding author
Ethics declarations
Competing interests
All authors are employees of Genentech.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 21559 kb)
Rights and permissions
About this article
Cite this article
Zhang, Y., Zhou, L., Rouge, L. et al. Conformational stabilization of ubiquitin yields potent and selective inhibitors of USP7. Nat Chem Biol 9, 51–58 (2013). https://doi.org/10.1038/nchembio.1134
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1134
This article is cited by
-
Structural and functional characterization of USP47 reveals a hot spot for inhibitor design
Communications Biology (2023)
-
USP7 inhibition induces apoptosis in glioblastoma by enhancing ubiquitination of ARF4
Cancer Cell International (2021)
-
UCHL3 promotes ovarian cancer progression by stabilizing TRAF2 to activate the NF-κB pathway
Oncogene (2020)
-
Understanding molecular mechanisms in cell signaling through natural and artificial sequence variation
Nature Structural & Molecular Biology (2019)
-
The deubiquitylating enzyme UCHL3 regulates Ku80 retention at sites of DNA damage
Scientific Reports (2018)