Abstract
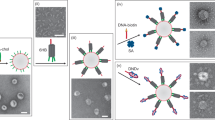
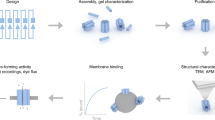
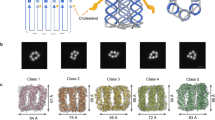
The diverse structure and regulated deformation of lipid bilayer membranes are among a cell's most fascinating features. Artificial membrane-bound vesicles, known as liposomes, are versatile tools for modelling biological membranes and delivering foreign objects to cells. To fully mimic the complexity of cell membranes and optimize the efficiency of delivery vesicles, controlling liposome shape (both statically and dynamically) is of utmost importance. Here we report the assembly, arrangement and remodelling of liposomes with designer geometry: all of which are exquisitely controlled by a set of modular, reconfigurable DNA nanocages. Tubular and toroid shapes, among others, are transcribed from DNA cages to liposomes with high fidelity, giving rise to membrane curvatures present in cells yet previously difficult to construct in vitro. Moreover, the conformational changes of DNA cages drive membrane fusion and bending with predictable outcomes, opening up opportunities for the systematic study of membrane mechanics.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Change history
22 November 2017
In the original version of this Article, the name of co-author Marc C. Llaguno was coded erroneously, resulting in it being incorrect when exported to citation databases. This has now been corrected, though no changes will be visible.
References
McMahon, H. T. & Gallop, J. L. Membrane curvature and mechanisms of dynamic cell membrane remodelling. Nature 438, 590–596 (2005).
Yoo, J. W., Irvine, D. J., Discher, D. E. & Mitragotri, S. Bio-inspired, bioengineered and biomimetic drug delivery carriers. Nat. Rev. Drug Discov. 10, 521–535 (2011).
Beales, P. A., Ciani, B. & Cleasby, A. J. Nature's lessons in design: nanomachines to scaffold, remodel and shape membrane compartments. Phys. Chem. Chem. Phys. 17, 15489–15507 (2015).
Schubert, R. Liposome preparation by detergent removal. Methods Enzymol. 367, 46–70 (2003).
Zhou, Y. & Shimizu, T. Lipid nanotubes: a unique template to create diverse one-dimensional nanostructures. Chem. Mater. 20, 625–633 (2008).
Frolov, V. A., Shnyrova, A. V. & Zimmerberg, J. Lipid polymorphisms and membrane shape. Cold Spring Harb. Perspect. Biol. 3, a004747 (2011).
Jesorka, A. et al. Generation of phospholipid vesicle-nanotube networks and transport of molecules therein. Nat. Protoc. 6, 791–805 (2011).
van Swaay, D. & deMello, A. Microfluidic methods for forming liposomes. Lab Chip 13, 752–767 (2013).
Carugo, D., Bottaro, E., Owen, J., Stride, E. & Nastruzzi, C. Liposome production by microfluidics: potential and limiting factors. Sci. Rep. 6, 25876 (2016).
Wu, F. & Dekker, C. Nanofabricated structures and microfluidic devices for bacteria: from techniques to biology. Chem. Soc. Rev. 45, 268–280 (2016).
Fourcade, B., Mutz, M. & Bensimon, D. Experimental and theoretical study of toroidal vesicles. Phys. Rev. Lett. 68, 2551–2554 (1992).
Genc, R., Ortiz, M. & O'Sullivan, C. K. Curvature-tuned preparation of nanoliposomes. Langmuir 25, 12604–12613 (2009).
Zidovska, A. et al. Block liposomes from curvature-stabilizing lipids: connected nanotubes, -rods, or -spheres. Langmuir 25, 2979–2985 (2009).
Tan, G. et al. Highly aspherical silica nanoshells by templating tubular liposomes. Soft Matter 5, 3006–3009 (2009).
Xu, P. et al. Undulating tubular liposomes through incorporation of a synthetic skin ceramide into phospholipid bilayers. Langmuir 25, 10422–10425 (2009).
Shih, W. M. & Lin, C. Knitting complex weaves with DNA origami. Curr. Opin. Struct. Biol. 20, 276–282 (2010).
Jabbari, H., Aminpour, M. & Montemagno, C. Computational approaches to nucleic acid origami. ACS Comb. Sci. 17, 535–547 (2015).
Pfeifer, W. & Saccà, B. From nano to macro through hierarchical self-assembly: the DNA paradigm. ChemBioChem 17, 1063–1080 (2016).
Douglas, S. M. et al. Rapid prototyping of 3D DNA-origami shapes with caDNAno. Nucleic Acids Res. 37, 5001–5006 (2009).
Dietz, H., Douglas, S. M. & Shih, W. M. Folding DNA into twisted and curved nanoscale shapes. Science 325, 725–730 (2009).
Gerling, T., Wagenbauer, K. F., Neuner, A. M. & Dietz, H. Dynamic DNA devices and assemblies formed by shape-complementary, non-base pairing 3D components. Science 347, 1446–1452 (2015).
Marras, A. E., Zhou, L., Su, H. J. & Castro, C. E. Programmable motion of DNA origami mechanisms. Proc. Natl Acad. Sci. USA 112, 713–718 (2015).
Yang, Y. et al. Self-assembly of size-controlled liposomes on DNA nanotemplates. Nat. Chem. 8, 476–483 (2016).
Perrault, S. D. & Shih, W. M. Virus-inspired membrane encapsulation of DNA nanostructures to achieve in vivo stability. ACS Nano 8, 5132–5140 (2014).
Lin, C., Perrault, S. D., Kwak, M., Graf, F. & Shih, W. M. Purification of DNA-origami nanostructures by rate-zonal centrifugation. Nucleic Acids Res. 41, e40 (2013).
Roux, A. et al. A minimal system allowing tubulation with molecular motors pulling on giant liposomes. Proc. Natl Acad. Sci. USA 99, 5394–5399 (2002).
Diz-Muñoz, A., Fletcher, D. A. & Weiner, O. D. Use the force: membrane tension as an organizer of cell shape and motility. Trends Cell Biol. 23, 47–53 (2013).
Marsden, H. R., Tomatsu, I. & Kros, A. Model systems for membrane fusion. Chem. Soc. Rev. 40, 1572–1585 (2011).
Ma, M. & Bong, D. Controlled fusion of synthetic lipid membrane vesicles. Acc. Chem. Res. 46, 2988–2997 (2013).
Zhang, D. Y. & Seelig, G. Dynamic DNA nanotechnology using strand-displacement reactions. Nat. Chem. 3, 103–113 (2011).
Bosco, A., Camunas-Soler, J. & Ritort, F. Elastic properties and secondary structure formation of single-stranded DNA at monovalent and divalent salt conditions. Nucleic Acids Res. 42, 2064–2074 (2014).
Cohen, F. S. & Melikyan, G. B. The energetics of membrane fusion from binding, through hemifusion, pore formation, and pore enlargement. J. Membr. Biol. 199, 1–14 (2004).
François-Martin, C., Rothman, J. E. & Pincet, F. Low energy cost for optimal speed and control of membrane fusion. Proc. Natl Acad. Sci. USA 114, 1238–1241 (2017).
Yan, H., Park, S. H., Finkelstein, G., Reif, J. H. & LaBean, T. H. DNA-templated self-assembly of protein arrays and highly conductive nanowires. Science 301, 1882–1884 (2003).
Udomprasert, A. et al. Amyloid fibrils nucleated and organized by DNA origami constructions. Nat. Nanotechnol. 9, 537–541 (2014).
Sun, W. et al. Casting inorganic structures with DNA molds. Science 346, 1258361 (2014).
Jones, M. R., Seeman, N. C. & Mirkin, C. A. Nanomaterials. programmable materials and the nature of the DNA bond. Science 347, 1260901 (2015).
Dong, Y. et al. Cuboid vesicles formed by frame-guided assembly on DNA origami scaffolds. Angew. Chem. Int. Ed. 56, 1586–1589 (2017).
Rajendran, A., Endo, M., Katsuda, Y., Hidaka, K. & Sugiyama, H. Photo-cross-linking-assisted thermal stability of DNA origami structures and its application for higher-temperature self-assembly. J. Am. Chem. Soc. 133, 14488–14491 (2011).
Feigenson, G. W. Phase boundaries and biological membranes. Annu. Rev. Biophys. Biomol. Struct. 36, 63–77 (2007).
Howorka, S. Changing of the guard. Science 352, 890–891 (2016).
McMahon, H. T., Kozlov, M. M. & Martens, S. Membrane curvature in synaptic vesicle fusion and beyond. Cell 140, 601–605 (2010).
Xu, W. et al. A programmable DNA origami platform to organize SNAREs for membrane fusion. J. Am. Chem. Soc. 138, 4439–4447 (2016).
Ferguson, S. M. & De Camilli, P. Dynamin, a membrane-remodelling GTPase. Nat. Rev. Mol. Cell Biol. 13, 75–88 (2012).
Morlot, S. & Roux, A. Mechanics of dynamin-mediated membrane fission. Annu. Rev. Biophys. 42, 629–649 (2013).
McCullough, J., Colf, L. A. & Sundquist, W. I. Membrane fission reactions of the mammalian ESCRT pathway. Annu. Rev. Biochem. 82, 663–692 (2013).
Henne, W. M., Stenmark, H. & Emr, S. D. Molecular mechanisms of the membrane sculpting ESCRT pathway. Cold Spring Harb. Perspect. Biol. 5, a016766 (2013).
Fan, W., Nassiri, A. & Zhong, Q. Autophagosome targeting and membrane curvature sensing by Barkor/Atg14(L). Proc. Natl Acad. Sci. USA 108, 7769–7774 (2011).
Nath, S. et al. Lipidation of the LC3/GABARAP family of autophagy proteins relies on a membrane-curvature-sensing domain in Atg3. Nat. Cell Biol. 16, 415–424 (2014).
Zadeh, J. N. et al. NUPACK: analysis and design of nucleic acid systems. J. Comput. Chem. 32, 170–173 (2011).
Douglas, S. M. et al. Self-assembly of DNA into nanoscale three-dimensional shapes. Nature 459, 414–418 (2009).
Rasband, W. S. ImageJ (US National Institutes of Health, 1997); https://imagej.nih.gov/ij/
Kremer, J. R., Mastronarde, D. N. & McIntosh, J. R. Computer visualization of three-dimensional image data using IMOD. J. Struct. Biol. 116, 71–76 (1996).
Acknowledgements
The authors thank J. T. Powell for proofreading the manuscript and D. K. Toomre and T. J. Melia for discussion. This work is supported by a National Institutes of Health (NIH) Director's New Innovator Award (DP2-GM114830), an NIH grant (R21-GM109466), and a Yale University faculty startup fund to C.L. and an European Research Council (ERC) funded grant under the European Union's Horizon 2020 research and innovation programme (grant agreement no. 669612) to F.P.
Author information
Authors and Affiliations
Contributions
Z.Z. initiated the project, designed and carried out most of the experiments, analysed the data, and prepared most of the manuscript. Y.Y. performed cryo-EM study and prepared the manuscript. F.P. modelled energy input for membrane fusion and prepared the manuscript. M.C.L. performed tomography study, analysed the data, and prepared the manuscript. C.L. initiated the project, designed and supervised the study, interpreted the data, and prepared the manuscript. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information
Supplementary information (PDF 53058 kb)
Rights and permissions
About this article
Cite this article
Zhang, Z., Yang, Y., Pincet, F. et al. Placing and shaping liposomes with reconfigurable DNA nanocages. Nature Chem 9, 653–659 (2017). https://doi.org/10.1038/nchem.2802
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchem.2802
This article is cited by
-
Structure and dynamics of an archetypal DNA nanoarchitecture revealed via cryo-EM and molecular dynamics simulations
Nature Communications (2023)
-
Digital nanoreactors to control absolute stoichiometry and spatiotemporal behavior of DNA receptors within lipid bilayers
Nature Communications (2023)
-
Creating complex protocells and prototissues using simple DNA building blocks
Nature Communications (2023)
-
Functionalization and higher-order organization of liposomes with DNA nanostructures
Nature Communications (2023)
-
Programmable multispecific DNA-origami-based T-cell engagers
Nature Nanotechnology (2023)