Abstract
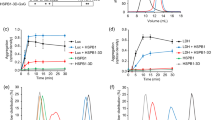
Cellular toxicity introduced by protein misfolding threatens cell fitness and viability. Failure to eliminate these polypeptides is associated with numerous aggregation diseases. Several protein quality control mechanisms degrade non-native proteins by the ubiquitin–proteasome system. Here, we use quantitative mass spectrometry to demonstrate that heat-shock triggers a large increase in the level of ubiquitylation associated with misfolding of cytosolic proteins. We discover that the Hul5 HECT ubiquitin ligase participates in this heat-shock stress response. Hul5 is required to maintain cell fitness after heat-shock and to degrade short-lived misfolded proteins. In addition, localization of Hul5 in the cytoplasm is important for its quality control function. We identify potential Hul5 substrates in heat-shock and physiological conditions to reveal that Hul5 is required for ubiquitylation of low-solubility cytosolic proteins including the Pin3 prion-like protein. These findings indicate that Hul5 is involved in a cytosolic protein quality control pathway that targets misfolded proteins for degradation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
McClellan, A. J., Tam, S., Kaganovich, D. & Frydman, J. Protein quality control: chaperones culling corrupt conformations. Nat. Cell Biol. 7, 736–741 (2005).
Wickner, S., Maurizi, M. R. & Gottesman, S. Posttranslational quality control: folding, refolding, and degrading proteins. Science 286, 1888–1893 (1999).
Balch, W. E., Morimoto, R. I., Dillin, A. & Kelly, J. W. Adapting proteostasis for disease intervention. Science 319, 916–919 (2008).
Heinemeyer, W., Kleinschmidt, J. A., Saidowsky, J., Escher, C. & Wolf, D. H. Proteinase yscE, the yeast proteasome/multicatalytic-multifunctional proteinase: mutants unravel its function in stress induced proteolysis and uncover its necessity for cell survival. EMBO J. 10, 555–562 (1991).
Rock, K. L. et al. Inhibitors of the proteasome block the degradation of most cell proteins and the generation of peptides presented on MHC class I molecules. Cell 78, 761–771 (1994).
Chau, V. et al. A multiubiquitin chain is confined to specific lysine in a targeted short-lived protein. Science 243, 1576–1583 (1989).
Vembar, S. S. & Brodsky, J. L. One step at a time: endoplasmic reticulum-associated degradation. Nat. Rev. Mol. Cell Biol. 9, 944–957 (2008).
Swanson, R., Locher, M. & Hochstrasser, M. A conserved ubiquitin ligase of the nuclear envelope/endoplasmic reticulum that functions in both ER-associated and Mat α2 repressor degradation. Genes Dev. 15, 2660–2674 (2001).
Bays, N. W., Gardner, R. G., Seelig, L. P., Joazeiro, C. A. & Hampton, R. Y. Hrd1p/Der3p is a membrane-anchored ubiquitin ligase required for ER-associated degradation. Nat. Cell Biol. 3, 24–29 (2001).
Rosenbaum, J. C. et al. Disorder targets misorder in nuclear quality control degradation: a disordered ubiquitin ligase directly recognizes its misfolded substrates. Mol. Cell 41, 93–106 (2011).
Gardner, R. G., Nelson, Z. W. & Gottschling, D. E. Degradation-mediated protein quality control in the nucleus. Cell 120, 803–815 (2005).
Fredrickson, E. K., Rosenbaum, J. C., Locke, M. N., Milac, T. I. & Gardner, R. G. Exposed hydrophobicity is a key determinant of nuclear quality control degradation. Mol. Biol. Cell 22, 2384–2395 (2011).
Connell, P. et al. The co-chaperone CHIP regulates protein triage decisions mediated by heat-shock proteins. Nat. Cell Biol. 3, 93–96 (2001).
Heck, J. W., Cheung, S. K. & Hampton, R. Y. Cytoplasmic protein quality control degradation mediated by parallel actions of the E3 ubiquitin ligases Ubr1 and San1. Proc. Natl Acad. Sci. USA 107, 1106–1111 (2010).
Prasad, R., Kawaguchi, S. & Ng, D. T. A nucleus-based quality control mechanism for cytosolic proteins. Mol. Biol. Cell 21, 2117–2127 (2010).
Nillegoda, N. B. et al. Ubr1 and ubr2 function in a quality control pathwayfor degradation of unfolded cytosolic proteins. Mol. Biol. Cell 21, 2102–2116 (2010).
Bengtson, M. H. & Joazeiro, C. A. Role of a ribosome-associated E3 ubiquitin ligase in protein quality control. Nature 467, 470–473 (2010).
Biederer, T., Volkwein, C. & Sommer, T. Degradation of subunits of the Sec61p complex, an integral component of the ER membrane, by the ubiquitin-proteasome pathway. EMBO J. 15, 2069–2076 (1996).
Hiller, M. M., Finger, A., Schweiger, M. & Wolf, D. H. ER degradation of a misfolded luminal protein by the cytosolic ubiquitin-proteasome pathway. Science 273, 1725–1728 (1996).
Seufert, W. & Jentsch, S. Ubiquitin-conjugating enzymes UBC4 and UBC5 mediate selective degradation of short-lived and abnormal proteins. EMBO J. 9, 543–550 (1990).
Mayor, T., Graumann, J., Bryan, J., MacCoss, M. J. & Deshaies, R. J. Quantitative profiling of ubiquitylated proteins reveals proteasome substrates and the substrate repertoire influenced by the Rpn10 receptor pathway. Mol. Cell. Proteomics 6, 1885–1895 (2007).
Derkatch, I. L., Bradley, M. E., Hong, J. Y. & Liebman, S. W. Prions affect the appearance of other prions: the story of [PIN(+)]. Cell 106, 171–182 (2001).
Alberti, S., Halfmann, R., King, O., Kapila, A. & Lindquist, S. A systematic survey identifies prions and illuminates sequence features of prionogenic proteins. Cell 137, 146–158 (2009).
Chernova, T. A. et al. Prion induction by the short-lived, stress-induced protein lsb2 is regulated by ubiquitination and association with the actin cytoskeleton. Mol. Cell 43, 242–252 (2011).
Huh, W. K. et al. Global analysis of protein localization in budding yeast. Nature 425, 686–691 (2003).
Xie, Y. & Varshavsky, A. UFD4 lacking the proteasome-binding region catalyses ubiquitination but is impaired in proteolysis. Nat. Cell Biol. 4, 1003–1007 (2002).
Wang, G., Yang, J. & Huibregtse, J. M. Functional domains of the Rsp5 ubiquitin-protein ligase. Mol. Cell. Biol. 19, 342–352 (1999).
Aviram, S. & Kornitzer, D. The ubiquitin ligase Hul5 promotes proteasomal processivity. Mol. Cell. Biol. 30, 985–994 (2010).
Crosas, B. et al. Ubiquitin chains are remodeled at the proteasome by opposing ubiquitin ligase and deubiquitinating activities. Cell 127, 1401–1413 (2006).
Kohlmann, S., Schafer, A. & Wolf, D. H. Ubiquitin ligase Hul5 is required for fragment-specific substrate degradation in endoplasmic reticulum-associated degradation. J. Biol. Chem. 283, 16374–16383 (2008).
Leggett, D. S. et al. Multiple associated proteins regulate proteasome structure and function. Mol. Cell 10, 495–507 (2002).
Hartl, F. U. Molecular chaperones in cellular protein folding. Nature 381, 571–579 (1996).
Young, J. C., Agashe, V. R., Siegers, K. & Hartl, F. U. Pathways of chaperone-mediated protein folding in the cytosol. Nat. Rev. Mol. Cell Biol. 5, 781–791 (2004).
Medicherla, B. & Goldberg, A. L. Heat shock and oxygen radicals stimulate ubiquitin-dependent degradation mainly of newly synthesized proteins. J. Cell Biol. 182, 663–673 (2008).
Tharun, S. et al. Yeast Sm-like proteins function in mRNA decapping and decay. Nature 404, 515–518 (2000).
Searfoss, A., Dever, T. E. & Wickner, R. Linking the 3′ poly(A) tail to the subunit joining step of translation initiation: relations of Pab1p, eukaryotic translation initiation factor 5b (Fun12p), and Ski2p-Slh1p. Mol. Cell. Biol. 21, 4900–4908 (2001).
McClellan, A. J., Scott, M. D. & Frydman, J. Folding and quality control of the VHL tumor suppressor proceed through distinct chaperone pathways. Cell 121, 739–748 (2005).
Lee, B. H. et al. Enhancement of proteasome activity by a small-molecule inhibitor of USP14. Nature 467, 179–184 (2010).
Tam, Y. Y., Fagarasanu, A., Fagarasanu, M. & Rachubinski, R. A. Pex3p initiates the formation of a preperoxisomal compartment from a subdomain of the endoplasmic reticulum in Saccharomyces cerevisiae. J. Biol. Chem. 280, 34933–34939 (2005).
Rappsilber, J., Ishihama, Y. & Mann, M. Stop and go extraction tips for matrix-assisted laser desorption/ionization, nanoelectrospray, and LC/MS sample pretreatment in proteomics. Anal. Chem. 75, 663–670 (2003).
Rappsilber, J., Mann, M. & Ishihama, Y. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat. Protoc. 2, 1896–1906 (2007).
Olsen, J. V. et al. Parts per million mass accuracy on an Orbitrap massspectrometer via lock mass injection into a C-trap. Mol. Cell. Proteomics 4, 2010–2021 (2005).
Wilde, I. B., Brack, M., Winget, J. M. & Mayor, T. Proteomic characterization of aggregating proteins after the inhibition of the ubiquitin proteasome system. J. Proteome Res. 10, 1062–1072 (2011).
Ducker, C. E. & Simpson, R. T. The organized chromatin domain of the repressed yeast a cell-specific gene STE6 contains two molecules of the corepressor Tup1p per nucleosome. EMBO J. 19, 400–409 (2000).
Miller, M. J., Xuong, N. H. & Geiduschek, E. P. A response of protein synthesis to temperature shift in the yeast Saccharomyces cerevisiae. Proc. Natl Acad. Sci. USA 76, 5222–5225 (1979).
Acknowledgements
The authors express gratitude to the following people for providing reagents: D. Finley (Harvard University, USA; HUL5 plasmids); J. Thorner (University of California, Berkeley, USA; anti-Pgk1 antibody); L. Howe (University of British Columbia, Canada; anti-H3 antibody); A. Chruscicki (University of British Columbia, Canada; Dynal-IgG); S. Jentsch (Max Planck Institute of Biochemistry, Germany; E2 double deletion strains); E. Craig (University of Wisconsin–Madison, USA; ssa1-45 strains); L. Conibear (University of British Columbia, Canada; TAP strains); R. Deshaies (California Institute of Technology, USA), C. Boone (University of Toronto, Canada) and M. Roberge (University of British Columbia, Canada) (deletion strains). We also thank P. Kaiser for comments on the manuscript, J. Gsponer for discussion, T.M. laboratory members for their encouragement and discussion and the following people for their support: L. Foster and N. Stoynov (mass spectrometry analysis); A. Chang (microscopy); P. Hieter and J. Stoepel (plate reader analysis); E. Jan (35S labelling and Odyssey). T.M. is supported by a grant from the Canada Institutes of Health Research (CIHR) and is a CIHR New Investigator. V.M. is supported by a grant from the CIHR.
Author information
Authors and Affiliations
Contributions
N.N.F. and T.M. conceived the project and designed experiments; N.N.F. carried out experiments; A.H.M.N. designed and carried out the SSA1 analysis and developed reagents; V.M. designed and carried out part of the localization analysis; N.N.F., A.H.M.N., V.M. and T.M. analysed the data; N.N.F., V.M. and T.M. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1731 kb)
Supplementary Table 1
Supplementary Information (XLSX 201 kb)
Supplementary Table 2
Supplementary Information (XLSX 63 kb)
Supplementary Table 3
Supplementary Information (XLSX 46 kb)
Rights and permissions
About this article
Cite this article
Fang, N., Ng, A., Measday, V. et al. Hul5 HECT ubiquitin ligase plays a major role in the ubiquitylation and turnover of cytosolic misfolded proteins. Nat Cell Biol 13, 1344–1352 (2011). https://doi.org/10.1038/ncb2343
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb2343
This article is cited by
-
Yeast polyubiquitin unit regulates synaptonemal complex formation and recombination during meiosis
Journal of Microbiology (2022)
-
The proteasome 19S cap and its ubiquitin receptors provide a versatile recognition platform for substrates
Nature Communications (2020)
-
CAT tails drive degradation of stalled polypeptides on and off the ribosome
Nature Structural & Molecular Biology (2019)
-
A SUMO-dependent feedback loop senses and controls the biogenesis of nuclear pore subunits
Nature Communications (2018)
-
Variable repeats in the eukaryotic polyubiquitin gene ubi4 modulate proteostasis and stress survival
Nature Communications (2017)