Abstract
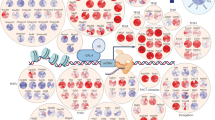
The bHLH transcription factor Hand1 is essential for placentation and cardiac morphogenesis in the developing embryo. Here we implicate Hand1 as a molecular switch that determines whether a trophoblast stem cell continues to proliferate or commits to differentiation. We identify a novel interaction of Hand1 with a protein that contains an I-mfa (inhibitor of myogenic factor) domain that anchors Hand1 in the nucleolus where it negatively regulates Hand1 activity. In the trophoblast stem-cell line Rcho-1, nucleolar sequestration of Hand1 accompanies sustained cell proliferation and renewal, whereas release of Hand1 into the nucleus leads to its activation, thus committing cells to a differentiated giant-cell fate. Site-specific phosphorylation is required for nucleolar release of Hand1, for its dimerization and biological function, and this is mediated by the non-canonical polo-like kinase Plk4 (Sak). Sak is co-expressed in Rcho-1 cells, localizes to the nucleolus during G2 and phosphorylates Hand1 as a requirement for trophoblast stem-cell commitment to a giant-cell fate. This study defines a novel cellular mechanism for regulating Hand1 that is a crucial step in the stem-cell differentiation pathway.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Rossant, J. & Cross, J. C. Placental development: lessons from mouse mutants. Nature Rev. Genet. 2, 538–548 (2001).
Cross, J. C. et al. Trophoblast functions, angiogenesis and remodeling of the maternal vasculature in the placenta. Mol. Cell Endocrinol. 187, 207–212 (2002).
Hughes, M. et al. The Hand1, Stra13 and Gcm1 transcription factors override FGF signaling to promote terminal differentiation of trophoblast stem cells. Dev. Biol. 271, 26–37 (2004).
Cross, J. C. et al. Hxt encodes a basic helix-loop-helix transcription factor that regulates trophoblast cell development. Development 121, 2513–2523 (1995).
Riley, P., Anson-Cartwright, L. & Cross, J. C. The Hand1 bHLH transcription factor is essential for placentation and cardiac morphogenesis. Nat. Genet. 18, 271–275 (1998).
Firulli, A. B., McFadden, D. G., Lin, Q., Srivastava, D. & Olson, E. N. Heart and extra-embryonic mesodermal defects in mouse embryos lacking the bHLH transcription factor Hand1. Nature Genet. 18, 266–270 (1998).
Sahgal, N., Canham, L. N., Konno, T., Wolfe, M. W. & Soares, M. J. Modulation of trophoblast stem cell and giant cell phenotypes: analyses using the Rcho-1 cell model. Differentiation 73, 452–462 (2005).
Scott, I. C., Anson-Cartwright, L., Riley, P., Reda, D. & Cross, J. C. The HAND1 basic helix-loop-helix transcription factor regulates trophoblast differentiation via multiple mechanisms. Mol. Cell Biol. 20, 530–541 (2000).
Firulli, B. A., Hadzic, D. B., McDaid, J. R. & Firulli, A. B. The basic helix-loop-helix transcription factors dHAND and eHAND exhibit dimerization characteristics that suggest complex regulation of function. J. Biol. Chem. 275, 33567–33573 (2000).
Hollenberg, S. M., Sternglanz, R., Cheng, P. F. & Weintraub, H. Identification of a new family of tissue-specific basic helix-loop-helix proteins with a two-hybrid system. Mol. Cell Biol. 15, 3813–3822 (1995).
Thebault, S., Gachon, F., Lemasson, I., Devaux, C. & Mesnard, J. M. Molecular cloning of a novel human I-mfa domain-containing protein that differently regulates human T-cell leukemia virus type I and HIV-1 expression. J. Biol. Chem. 275, 4848–4857 (2000).
Chen, C. M., Kraut, N., Groudine, M. & Weintraub, H. I-mf, a novel myogenic repressor, interacts with members of the MyoD family. Cell 86, 731–741 (1996).
Kraut, N., Snider, L., Chen, C. M., Tapscott, S. J. & Groudine, M. Requirement of the mouse I-mfa gene for placental development and skeletal patterning. EMBO J. 17, 6276–6288 (1998).
Gautier, V. W., Sheehy, N., Duffy, M., Hashimoto, K. & Hall, W. W. Direct interaction of the human I-mfa domain-containing protein, HIC, with HIV-1 Tat results in cytoplasmic sequestration and control of Tat activity. Proc. Natl Acad. Sci. USA 102, 16362–16367 (2005).
Snider, L. et al. Inhibition of Tcf3 binding by I-mfa domain proteins. Mol. Cell Biol. 21, 1866–1873 (2001).
Thebault, S., Basbous, J., Gay, B., Devaux, C. & Mesnard, J. M. Sequence requirement for the nucleolar localization of human I-mfa domain-containing protein (HIC p40). Eur. J. Cell Biol. 79, 834–838 (2000).
Thebault, S. & Mesnard, J. M. How the sequestration of a protein interferes with its mechanism of action: example of a new family of proteins characterized by a particular cysteine-rich carboxy-terminal domain involved in gene expression regulation. Curr. Prot. Pept. Sci. 2, 155–167 (2001).
Stark, L. A. & Dunlop, M. G. Nucleolar sequestration of RelA (p65) regulates NF-kappaB-driven transcription and apoptosis. Mol. Cell Biol. 25, 5985–6004 (2005).
Weber, J. D., Taylor, L. J., Roussel, M. F., Sherr, C. J. & Bar-Sagi, D. Nucleolar Arf sequesters Mdm2 and activates p53. Nature Cell Biol. 1, 20–26 (1999).
Tao, W. & Levine, A. J. P19(ARF) stabilizes p53 by blocking nucleo-cytoplasmic shuttling of Mdm2. Proc. Natl Acad. Sci. US A 96, 6937–6941 (1999).
Weber, J. D. et al. Cooperative signals governing ARF-mdm2 interaction and nucleolar localization of the complex. Mol. Cell Biol. 20, 2517–2528 (2000).
Lohrum, M. A., Ashcroft, M., Kubbutat, M. H. & Vousden, K. H. Identification of a cryptic nucleolar-localization signal in MDM2. Nature Cell Biol. 2, 179–181 (2000).
Datta, A. et al. Myc-ARF (alternate reading frame) interaction inhibits the functions of Myc. J. Biol. Chem. 279, 36698–36707 (2004).
Sahgal, N., Canham, L. N., Canham, B. & Soares, M. J. Rcho-1 trophoblast stem cells: a model system for studying trophoblast cell differentiation. Methods Mol. Med. 121, 159–178 (2006).
Hamlin, G. P., Lu, X. J., Roby, K. F. & Soares, M. J. Recapitulation of the pathway for trophoblast giant cell differentiation in vitro: stage-specific expression of members of the prolactin gene family. Endocrinology 134, 2390–2396 (1994).
Faria, T. N. & Soares, M. J. Trophoblast cell differentiation: establishment, characterization, and modulation of a rat trophoblast cell line expressing members of the placental prolactin family. Endocrinology 129, 2895–2906 (1991).
Parast, M. M., Aeder, S. & Sutherland, A. E. Trophoblast giant-cell differentiation involves changes in cytoskeleton and cell motility. Dev. Biol. 230, 43–60 (2001).
Tanaka, S., Kunath, T., Hadjantonakis, A. K., Nagy, A. & Rossant, J. Promotion of trophoblast stem cell proliferation by FGF4. Science 282, 2072–2075 (1998).
Visintin, R., Stegmeier, F. & Amon, A. The role of the polo kinase Cdc5 in controlling Cdc14 localization. Mol. Biol. Cell 14, 4486–4498 (2003).
Firulli, B. A. et al. PKA, PKC, and the protein phosphatase 2A influence HAND factor function: a mechanism for tissue-specific transcriptional regulation. Mol. Cell 12, 1225–1237 (2003).
Huang, Z., Traugh, J. A. & Bishop, J. M. Negative control of the Myc protein by the stress-responsive kinase Pak2. Mol. Cell Biol. 24, 1582–1594 (2004).
Centonze, V. E., Firulli, B. A. & Firulli, A. B. Fluorescence resonance energy transfer (FRET) as a method to calculate the dimerization strength of basic helix-loop-helix (bHLH) proteins. Biol. Proced. Online. 6, 78–82 (2004).
Jaken, S. Protein kinase C isozymes and substrates. Curr. Opin. Cell Biol. 8, 168–173 (1996).
Griffioen, G. & Thevelein, J. M. Molecular mechanisms controlling the localisation of protein kinase A. Curr. Genet. 41, 199–207 (2002).
Andersen, J. S. et al. Nucleolar proteome dynamics. Nature 433, 77–83 (2005).
Newton, A. C. Regulation of protein kinase C. Curr. Opin. Cell Biol. 9, 161–167 (1997).
MacAuley, A., Cross, J. C. & Werb, Z. Reprogramming the cell cycle for endoreduplication in rodent trophoblast cells. Mol. Biol. Cell 9, 795–807 (1998).
Fode, C., Motro, B., Yousefi, S., Heffernan, M. & Dennis, J. W. Sak, a murine protein-serine/threonine kinase that is related to the Drosophila polo kinase and involved in cell proliferation. Proc. Natl Acad. Sci. USA 91, 6388–6392 (1994).
Hudson, J. W. et al. Late mitotic failure in mice lacking Sak, a polo-like kinase. Curr. Biol. 11, 441–446 (2001).
Leung, G. C., Ho, C. S., Blasutig, I. M., Murphy, J. M. & Sicheri, F. Determination of the Plk4/Sak consensus phosphorylation motif using peptide spots arrays. FEBS Lett. 581, 77–83 (2007).
Ma, G. T. & Linzer, D. I. GATA-2 restricts prolactin-like protein A expression to secondary trophoblast giant cells in the mouse. Biol. Reprod. 63, 570–574 (2000).
Queralt, E., Lehane, C., Novak, B. & Uhlmann, F. Downregulation of PP2A(Cdc55) phosphatase by separase initiates mitotic exit in budding yeast. Cell 125, 719–732 (2006).
Pederson, T. The plurifunctional nucleolus. Nucleic Acids Res. 26, 3871–3876 (1998).
Hill, A. A. & Riley, P. R. Differential regulation of Hand1 homodimer and Hand1-E12 heterodimer activity by the cofactor FHL2. Mol. Cell Biol. 24, 9835–9847 (2004).
Kumar, R., Conklin, D. S. & Mittal, V. High-throughput selection of effective RNAi probes for gene silencing. Genome Res. 13, 2333–2340 (2003).
Kunath, T. et al. Transgenic RNA interference in ES cell-derived embryos recapitulates a genetic null phenotype. Nature Biotechnol. 21, 559–561 (2003).
Kurki, S. et al. Nucleolar protein NPM interacts with HDM2 and protects tumor suppressor protein p53 from HDM2-mediated degradation. Cancer Cell 5, 465–475 (2004).
Swallow, C. J., Ko, M. A., Siddiqui, N. U., Hudson, J. W. & Dennis, J. W. Sak/Plk4 and mitotic fidelity. Oncogene 24, 306–312 (2005).
Moorman, A. F., Houweling, A. C., de Boer, P. A. & Christoffels, V. M. Sensitive nonradioactive detection of mRNA in tissue sections: novel application of the whole-mount in situ hybridization protocol. J. Histochem. Cytochem. 49, 1–8 (2001).
Riley, P. R., Gertsenstein, M., Dawson, K. & Cross, J. C. Early exclusion of hand1-deficient cells from distinct regions of the left ventricular myocardium in chimeric mouse embryos. Dev. Biol. 227, 156–168 (2000).
Acknowledgements
We thank Anthony Firulli, Vivek Mittal and Sabine Thebault, for generously providing plasmids, Jean-Michel Mesnard for the α-HIC antibody and Michael Soares and Satoshi Tanaka for generously providing the Rcho-1 and TS cell lines respectively. This research was supported by the British Heart Foundation.
Author information
Authors and Affiliations
Contributions
D.M.J.M carried out the functional cell-based characterization studies. C.A.R. and N.S. contributed to northern and western data and histological sections from Sak-null embryos. M.D.M.F.-V. peformed the yeast two-hybrid screen. COR collected and prepared Sak-null embryos. C.J.S. and J.W.D. provided Sak-null embryos and critical appraisal of Sak-null embryo data. P.R.R. carried out initial characterization studies and Sak-null embryo analyses, devised the functional studies and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Title
Supplementary Movie 1 (MOV 1070 kb)
Supplementary Title
Supplementary Figures S1, S2, S3 and S4 (PDF 886 kb)
Rights and permissions
About this article
Cite this article
Martindill, D., Risebro, C., Smart, N. et al. Nucleolar release of Hand1 acts as a molecular switch to determine cell fate. Nat Cell Biol 9, 1131–1141 (2007). https://doi.org/10.1038/ncb1633
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1633
This article is cited by
-
Opposite Roles of the JMJD1A Interaction Partners MDFI and MDFIC in Colorectal Cancer
Scientific Reports (2020)
-
Grabbing the genome by the NADs
Chromosoma (2016)
-
A novel role for Plk4 in regulating cell spreading and motility
Oncogene (2015)
-
Drug-induced cell cycle modulation leading to cell-cycle arrest, nuclear mis-segregation, or endoreplication
BMC Cell Biology (2011)
-
A bHLH Code for Cardiac Morphogenesis
Pediatric Cardiology (2010)