Abstract
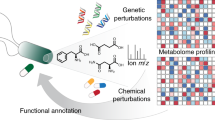
Bioactive compounds can be valuable research tools and drug leads, but it is often difficult to identify their mechanism of action or cellular target. Here we investigate the potential for integration of chemical-genetic and genetic interaction data to reveal information about the pathways and targets of inhibitory compounds. Taking advantage of the existing complete set of yeast haploid deletion mutants, we generated drug-hypersensitivity (chemical-genetic) profiles for 12 compounds. In addition to a set of compound-specific interactions, the chemical-genetic profiles identified a large group of genes required for multidrug resistance. In particular, yeast mutants lacking a functional vacuolar H+-ATPase show multidrug sensitivity, a phenomenon that may be conserved in mammalian cells. By filtering chemical-genetic profiles for the multidrug-resistant genes and then clustering the compound-specific profiles with a compendium of large-scale genetic interaction profiles, we were able to identify target pathways or proteins. This method thus provides a powerful means for inferring mechanism of action.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Winzeler, E.A. et al. Functional characterization of the S. cerevisiae genome by gene deletion and parallel analysis. Science 285, 901–906 (1999).
Hughes, T.R. et al. Functional discovery via a compendium of expression profiles. Cell 102, 109–926 (2000).
Marton, M.J. et al. Drug target validation and identification of secondary drug target effects using DNA microarrays. Nat. Med. 4, 1293–1301 (1998).
Hartwell, L.H., Szankasi, P., Roberts, C.J., Murray, A.W. & Friend, S.H. Integrating genetic approaches into the discovery of anticancer drugs. Science 278, 1064–1068 (1997).
Tong, A.H. et al. Systematic genetic analysis with ordered arrays of yeast deletion mutants. Science 294, 2364–2368 (2001).
Thomas, J.H., Neff, N.F. & Botstein, D. Isolation and characterization of mutations in the β-tubulin gene of Saccharomyces cerevisiae. Genetics 111, 715–734 (1985).
Liu, J. et al. Calcineurin is a common target of cyclophilin-cyclosporin A and FKBP-FK506 complexes. Cell 66, 807–815 (1991).
Rittberg, D.A. & Wright, J.A. Relationships between sensitivity to hydroxyurea and 4-methyl-5-amino-1-formylisoquinoline thiosemicarbazone (MAIO) and ribonucleotide reductase RNR2 mRNA levels in strains of Saccharomyces cerevisiae. Biochem. Cell. Biol. 67, 352–357 (1989).
Hsiang, Y.H., Lihou, M.G. & Liu, L.F. Arrest of replication forks by drug-stabilized topoisomerase I–DNA cleavable complexes as a mechanism of cell killing by camptothecin. Cancer Res. 49, 5077–5082 (1989).
Turi, T.G. & Loper, J.C. Multiple regulatory elements control expression of the gene encoding the Saccharomyces cerevisiae cytochrome P450, lanosterol 14 α-demethylase (ERG11). J. Biol. Chem. 267, 2046–2056 (1992).
Truan, G., Epinat, J.C., Rougeulle, C., Cullin, C. & Pompon, D. Cloning and characterization of a yeast cytochrome b5-encoding gene which suppresses ketoconazole hypersensitivity in a NADPH-P-450 reductase-deficient strain. Gene 142, 123–127 (1994).
Zheng, X.F., Florentino, D., Chen, J., Crabtree, G.R. & Schreiber, S.L. TOR kinase domains are required for two distinct functions, only one of which is inhibited by rapamycin. Cell 82, 121–130 (1995).
Kuo, S.C. & Lampen, J.O. Tunicamycin—an inhibitor of yeast glycoprotein synthesis. Biochem. Biophys. Res. Commun. 58, 287–295 (1974).
Cutler, N.S., Heitman, J. & Cardenas, M.E. STT4 is an essential phosphatidylinositol 4-kinase that is a target of wortmannin in Saccharomyces cerevisiae. J. Biol. Chem. 272, 27671–27677 (1997).
Falco, S.C. & Dumas, K.S. Genetic analysis of mutants of Saccharomyces cerevisiae resistant to the herbicide sulfometuron methyl. Genetics 109, 21–35 (1985).
Parsons, W.J., Ramkumar, V. & Stiles, G.L. Isobutylmethylxanthine stimulates adenylate cyclase by blocking the inhibitory regulatory protein, Gi. Mol. Pharmacol. 34, 37–41 (1988).
Garrett-Engele, P., Moilanen, B. & Cyert, M.S. Calcineurin, the Ca2+/calmodulin-dependent protein phosphatase, is essential in yeast mutants with cell integrity defects and in mutants that lack a functional vacuolar H+-ATPase. Mol. Cell. Biol. 15, 4103–4114 (1995).
Bauer, B.E., Wolfger, H. & Kuchler, K. Inventory and function of yeast ABC proteins: about sex, stress, pleiotropic drug and heavy metal resistance. Biochim. Biophys. Acta 1461, 217–236 (1999).
Mukhopadhyay, K., Kohli, A. & Prasad, R. Drug susceptibilities of yeast cells are affected by membrane lipid composition. Antimicrob. Agents Chemother. 46, 3695–3705 (2002).
Yoshida, S. & Anraku, Y. Characterization of staurosporine-sensitive mutants of Saccharomyces cerevisiae: vacuolar functions affect staurosporine sensitivity. Mol. Gen. Genet. 263, 877–888 (2000).
Simon, S., Roy, D. & Schindler, M. Intracellular pH and the control of multidrug resistance. Proc. Natl. Acad. Sci. USA 91, 1128–1132 (1994).
Ma, L. & Center, M.S. The gene encoding vacuolar H+-ATPase subunit C is overexpressed in multidrug-resistant HL60 cells. Biochem. Biophys. Res. Commun. 182, 675–681 (1992).
Drose, S. et al. Inhibitory effect of modified bafilomycins and concanamycins on P- and V-type adenosine triphosphatases. Biochemistry 32, 3902–3906 (1993).
Ouar, Z. et al. Inhibitors of vacuolar H+-ATPase impair the preferential accumulation of daunomycin in lysosomes and reverse the resistance to anthracyclines in drug-resistant renal epithelial cells. Biochem. J. 370, 185–193 (2003).
Dohmen, R.J., Wu, P. & Varshavsky, A. Heat-inducible degron: a method for constructing temperature-sensitive mutants. Science 263, 1273–1276 (1994).
Cyert, M.S. Genetic analysis of calmodulin and its targets in Saccharomyces cerevisiae. Annu. Rev. Genet. 35, 647–672 (2001).
Cyert, M.S. & Thorner, J. Regulatory subunit (CNB1 gene product) of yeast Ca2+/calmodulin-dependent phosphoprotein phosphatases is required for adaptation to pheromone. Mol. Cell. Biol. 12, 3460–3469 (1992).
Tanida, I., Hasegawa, A., Iida, H., Ohya, Y. & Anraku, Y. Cooperation of calcineurin and vacuolar H+-ATPase in intracellular Ca2+ homeostasis of yeast cells. J. Biol. Chem. 270, 10113–10119 (1995).
Aasland, R., Stewart, A.F. & Gibson, T. The SANT domain: a putative DNA-binding domain in the SWI-SNF and ADA complexes, the transcriptional co-repressor N-CoR and TFIIIB. Trends Biochem. Sci. 21, 87–88 (1996).
Boyer, L.A. et al. Essential role for the SANT domain in the functioning of multiple chromatin remodeling enzymes. Mol. Cell. 10, 935–942 (2002).
Bellaoui, M. et al. Elg1 forms an alternative RFC complex important for DNA replication and genome integrity. EMBO J. 22, 4304–13 (2003).
Ben-Aroya, S., Koren, A., Liefshitz, B., Steinlauf, R. & Kupiec, M. ELG1, a yeast gene required for genome stability, forms a complex related to replication factor C. Proc. Natl. Acad. Sci. USA 100, 9906–9911 (2003).
Ersfeld, K. et al. Characterization of the tubulin-tyrosine ligase. J. Cell. Biol. 120, 725–732 (1993).
Shoemaker, D.D., Lashkari, D.A., Morris, D., Mittmann, M. & Davis, R.W. Quantitative phenotypic analysis of yeast deletion mutants using a highly parallel molecular bar-coding strategy. Nat. Genet. 14, 450–456 (1996).
Giaever, G. et al. Genomic profiling of drug sensitivities via induced haploinsufficiency. Nat. Genet. 21, 278–283 (1999).
Barstead, R. Genome-wide RNAi. Curr. Opin. Chem. Biol. 5, 63–66 (2001).
Shi, Y. Mammalian RNAi for the masses. Trends Genet. 19, 9–12 (2003).
Brachmann, C.B. et al. Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14, 115–132 (1998).
Breitkreutz, B.J., Stark, C. & Tyers, M. Osprey: a network visualization system. Genome Biol. 4, R22 (2003).
Ashburner, M. et al. Gene ontology: tool for the unification of biology. The Gene Ontology Consortium. Nat. Genet. 25, 25–29 (2000).
Mewes, H.W., Albermann, K., Heumann, K., Liebl, S. & Pfeiffer, F. MIPS: a database for protein sequences, homology data and yeast genome information. Nucleic Acids Res. 25, 28–30 (1997).
Breitkreutz, B.J., Stark, C. & Tyers, M. The GRID: the General Repository for Interaction Datasets. Genome Biol. 4, R23 (2003).
Acknowledgements
We thank D. Drubin for tub2-403 and M. Cyert, J. Anderson and S. Brill for gifts of FK506, fluconazole and camptothecin, respectively. We thank H. Lu, X. Xin and V. Ghandi for assistance with drug screening and confirmations and Y. Chen and X. Cheng for assistance with SGA screening. This work was supported by grants from the Canadian Institute of Health Research (CIHR) to C.B. and T.R.H. and from the National Cancer Institute of Canada (NCIC) to G.W.B. A.B.P holds a Natural Sciences and Engineering Research Council of Canada (NSERC) graduate student fellowship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Parsons, A., Brost, R., Ding, H. et al. Integration of chemical-genetic and genetic interaction data links bioactive compounds to cellular target pathways. Nat Biotechnol 22, 62–69 (2004). https://doi.org/10.1038/nbt919
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt919
This article is cited by
-
Genome-wide identification of resistance genes and transcriptome regulation in yeast to accommodate ammonium toxicity
BMC Genomics (2022)
-
Identification of anticancer drug target genes using an outside competitive dynamics model on cancer signaling networks
Scientific Reports (2021)
-
Ccr4-Not as a mediator of environmental signaling: a jack of all trades and master of all
Current Genetics (2021)
-
CGINet: graph convolutional network-based model for identifying chemical-gene interaction in an integrated multi-relational graph
BMC Bioinformatics (2020)
-
Quantitative and multiplexed chemical-genetic phenotyping in mammalian cells with QMAP-Seq
Nature Communications (2020)