Abstract
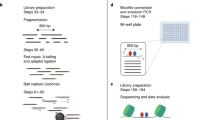
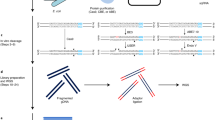
Ligation-mediated polymerase chain reaction (LM-PCR) is a genomic analysis technique for determination of (1) primary DNA nucleotide sequences (2) cytosine methylation patterns (3) DNA lesion formation and repair, and (4) in vivo protein–DNA footprints1,2,3,4. However, LM-PCR can be limited by the multiple steps required and the relatively short stretch of sequence (usually <200 bp) that can be analyzed per reaction. We report here a simplified, one-day LM-PCR protocol in which all pipetting steps can be performed by a robotic workstation and which, moreover, provides longer reads (>350 bp) and enhanced signal quality by use of nonradioactive detection and a LI-COR DNA sequencing instrument. Sensitivity comparable to radiolabeling is achieved using oligonucleotide primers that are 5′-end labeled with infrared fluorochromes. We showed that the technique could be used for sensitive and reproducible in vivo photofootprinting of the human phosphoglycerate kinase 1 (PGK1) promoter, as well as providing good Maxam–Gilbert sequence information. The methods described here should allow high-throughput, high-resolution analysis of transcription factor binding and chromatin structure, and also may be useful for sequencing gaps that are refractory to cloning.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Mueller, P.R. & Wold, B. In vivo footprinting of a muscle specific enhancer by ligation-mediated PCR. Science 246, 780–786 (1989).
Pfeifer, G.P., Steigerwald, S.D., Muller, P.R., Wold, B. & Riggs, A.D. Genomic sequencing and methylation analysis by ligation mediated PCR. Science 246, 810–813 (1989).
Pfeifer, G.P., Chen, H.H., Komura, J. & Riggs, A.D. Chromatin structure analysis by ligation-mediated PCR and terminal transferase-mediated PCR. Methods Enzymol. 304, 548–571 (1999).
Becker, M.M. & Grossmann, G. Photofootprinting DNA in vitro and vivo. Footprinting of nucleic acid–protein complexes (ed. Revzin, A.) 129–159 (Academic Press, New York, NY; 1993).
Mullis, K.B. & Faloona, F.A. Specific synthesis of DNA in vitro via a polymerase-catalyzed chain reaction. Methods Enzymol. 155, 335–350 (1987).
Machida, M., Kamio, K. & Sorensen, D. Long range and highly sensitive DNase I footprinting by an automated infrared DNA sequencer. Biotechniques 23, 300–303 (1997).
Tormanen, V.T., Swiderski, P.M., Kaplan, B.E., Pfeifer, G.P. & Riggs, A.D. Extension product capture improves genomic sequencing and DNase I footprinting by ligation-mediated PCR. Nucleic Acids Res. 20, 5487–5488 (1992).
Chou, Q., Russell, M., Birch, D.E., Raymond, J. & Bloch, W. Prevention of pre-PCR mis-priming and primer dimerization improves low-copy-number amplifications. Nucleic Acids Res. 20, 1717–1723 (1992).
Hebert, B., Bergeron, J., Potworowski, E.F. & Tijssen, P. Increased PCR sensitivity by using paraffin wax as a reaction mix overlay. Mol. Cell. Probes 7, 249–252 (1993).
Horton, R.M., Hoppe, B.L. & Conti-Tronconi, B.M. Ampligrease–hot start PCR using petroleum jelly. Biotechniques 16, 42–43 (1994).
Cairns, M.J. & Murray, V. Influence of chromatin structure on bleomycin–DNA interactions at base pair resolution in the human β-globin gene cluster. Biochemistry 35, 8753–8760 (1996).
Pfeifer, G.P., Tanguay, R.L., Steigerwald, S.D. & Riggs, A.D. In vivo footprint and methylation analysis by PCR-aided genomic sequencing: comparison of active and inactive X chromosomal DNA at the CpG island and promoter of human PGK-1. Genes Dev. 4, 1277–1287 (1990).
Pfeifer, G.P. & Riggs, A.D. Chromatin differences between active and inactive X chromosomes revealed by genomic footprinting of permeabilized cells using DNase I and ligation-mediated PCR. Genes Dev. 5, 1102–1113 (1991).
Pfeifer, G.P., Drouin, R., Riggs, A.D. & Holmquist, G.P. Binding of transcription factors creates hot spots for UV photoproducts in vivo. Mol. Cell. Biol. 12, 1798–1804 (1992).
Rodriguez, H., Drouin, R., Holmquist, G.P. & Akman, S.A. A hot spot for hydrogen peroxide-induced damage in the human hypoxia-inducible factor 1 binding site of the PGK 1 gene. Arch. Biochem. Biophys. 338, 207–212 (1997).
Faisst, S. & Meyer, S. Compilation of vertebrate-encoded transcription factors. Nucleic Acids Res. 20, 3–26 (1992).
Yamaguchi, Y., Nishio, H., Kishi, K., Ackerman, S.J. & Suda, T. C/EBPβ and GATA-1 synergistically regulate activity of the eosinophil granule major basic protein promoter: implication for C/EBPβ activity in eosinophil gene expression. Blood 94, 1429–1439 (1999).
Read, D. & Manley, J.L. Alternatively spliced transcripts of the Drosophila tramtrack gene encode zinc finger proteins with distinct DNA binding specificities. EMBO J. 11, 1035–1044 (1992).
Smale, S.T. & Baltimore, D. The “initiator” as a transcription control element. Cell 57, 103–113 (1989).
Meyers, S., Downing, J.R. & Hiebert, S.W. Identification of AML-1 and the (8;21) translocation protein (AML-1/ETO) as sequence-specific DNA-binding proteins: the runt homology domain is required for DNA binding and protein–protein interactions. Mol. Cell. Biol. 13, 6336–6345 (1993).
Kroeger, P.E. & Morimoto, R.I. Selection of new HSF1 and HSF2 DNA-binding sites reveals difference in trimer cooperativity. Mol. Cell. Biol. 14, 7592–7603 (1994).
Tornaletti, S. & Pfeifer, G.P. UV light as a footprinting agent: modulation of UV-induced DNA damage by transcription factors bound at the promoters of three human genes. J. Mol. Biol. 249, 714–728 (1995).
Hillier, L. & Green, P. OSP: A computer program for choosing PCR and DNA sequencing primers. PCR Methods Appl. 1, 124–128 (1991).
Mueller, P.R., Garrity, P.A. & Wold, B. Ligation-mediated PCR for genomic sequencing and footprinting. pp. 15.5.1–15.5.26. Current protocols in molecular biology (John Wiley & Sons, New York, NY; 1992).
Maxam, A.M. & Gilbert, W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 65, 499–560 (1980).
Acknowledgements
We thank Dr. G. Holmquist for encouragement and support, and J. Flanagan for help with editing the manuscript. Jin Zhou supplied Maxam–Gilbert HeLa DNA for these studies. This work was supported in part by grants GM50575 (to A.D.R.), AG15695 (to S.D.F.), and CA69449 (G. Holmquist) from the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Dai, SM., Chen, HH., Chang, C. et al. Ligation-mediated PCR for quantitative in vivo footprinting. Nat Biotechnol 18, 1108–1111 (2000). https://doi.org/10.1038/80323
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/80323