Abstract
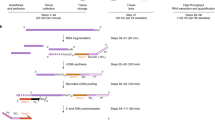
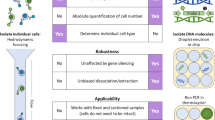
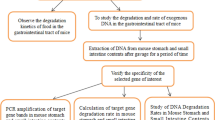
Deoxyribonucleic acid (DNA) can be extracted from different tissue sources for genotyping of mice. Methods of collecting tissue vary with respect to their perceived invasiveness, and in some cases, tissue is collected multiple times in order to verify a genotype. The authors' goal was to refine and optimize tissue collection methods with quantitative measures for subsequent genotyping. To do this, the authors used real-time quantitative polymerase chain reaction (qPCR) analysis to quantify DNA extracted from fecal pellets, hair, buccal swab samples, ear punch samples and tail biopsy samples and then compared the quantities of DNA obtained by using either previously published protocols or commercial kits to extract DNA. They found that 2-mm tail biopsy samples yielded significantly more DNA than did the other sources and that commercial extraction kits generally yielded more DNA than did published protocols. The authors also assessed the stability of DNA extracted from the various tissue sources by repeating the qPCR analysis after the samples had been frozen and stored for 44 months. Although the quantities of DNA in the stored samples had decreased, all the samples could still be used for PCR-based genotyping. The authors' work supports the collection of a single minimal biopsy sample of 2 mm of tail or ear tissue, above other sources, for highest yield of DNA for genotyping. With appropriate storage, DNA remains usable for PCR-based genotyping for years without a need for repeated animal sampling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
We are sorry, but there is no personal subscription option available for your country.
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Pinkert, C.A. Transgenic animal technology: alternatives in genotyping and phenotyping. Comp. Med. 53, 126–139 (2003).
Attal, J., Cajero-Juarez, M. & Houdebine, L.M. A simple method of DNA extraction from whole tissues and blood using glass powder for detection of transgenic animals by PCR. Transgenic Res. 4, 149–150 (1995).
Burkhart, C.A., Norris, M.D. & Haber, M. A simple method for the isolation of genomic DNA from mouse tail free of real-time PCR inhibitors. J. Bioch. Biophys. Methods 52, 145–149 (2002).
Drews, R., Drohan, W.N. & Lubon, H. Transgene detection in mouse tail digests. Biotechniques 17, 866–867 (1994).
Gaw, A., Mancini, F.P. & Ishibashi, S. Rapid genotyping of low density lipoprotein receptor knockout mice using a polymerase chain reaction technique. Lab. Anim. 29, 447–449 (1995).
Henneberger, C., Grantyn, R. & Rothe, T. Rapid genotyping of newborn gene mutant mice. J. Neurosci. Methods 100, 123–126 (2000).
Lin, C.S., Magnuson, T. & Samols, D. A rapid procedure to identify newborn transgenic mice. DNA 8, 297–299 (1989).
Petkov, P.M. et al. Development of a SNP genotyping panel for genetic monitoring of the laboratory mouse. Genomics 83, 902–911 (2004).
Petkov, P.M. et al. An efficient SNP system for mouse genome scanning and elucidating strain relationships. Genome Res. 14, 1806–1811 (2004).
Sorensen, D.B. et al. The impact of tail tip amputation and ink tattoo on C57BL/6JBomTac mice. Lab. Anim. 41, 19–29 (2007).
Zimmermann, K., Schwarz, H.P. & Turecek, P.L. Deoxyribonucleic acid preparation in polymerase chain reaction genotyping of transgenic mice. Comp. Med. 50, 314–316 (2000).
Campbell, D.B. & Hess, E.J. Rapid genotyping of mutant mice using dried blood spots for polymerase chain reaction (PCR) analysis. Brain Res. Brain Res. Protoc. 1, 117–123 (1997).
Chen, S. & Evans, G.A. A simple screening method for trangenic mice using the polymerase chain reaction. Biotechniques 8, 32–33 (1990).
Gregory, C.A., Myal, Y. & Shiu, R.P. Rapid genotyping of transgenic mice using dried blood spots from Guthrie cards for PCR analysis. Biotechniques 18, 758–760 (1995).
Hofstetter, J.R. et al. Genomic DNA from mice: a comparison of recovery methods and tissue sources. Biochem. Mol. Med. 62, 197–202 (1997).
Irwin, M.H., Moffatt, R.J. & Pinkert, C.A. Identification of transgenic mice by PCR analysis of saliva. Nat. Biotech. 14, 1146–1148 (1996).
Meldgaard, M., Bollen, P.J. & Finsen, B. Non-invasive method for sampling and extraction of mouse DNA for PCR. Lab. Anim. 38, 413–417 (2004).
Mitrecic', D., Mavric', S., Branica, B.V. & Gajovic', S. Mice genotyping using buccal swab samples: an improved method. iochem. Genet. 46, 105–112 (2008).
Lahm, H., Hoeflich, A., Rieger, N., Wanke, R. & Wolf, E. Identification of transgenic mice by direct PCR analysis of lysates of epithelial cells obtained from the inner surface of the rectum. Transgenic Res. 7, 131–134 (1998).
Broome, R.L. et al. Non-invasive transgenic mouse genotyping using stool analysis. FEBS Lett. 462, 159–160 (1999).
Ohhara, M., Kurosu, Y. & Esumi, M. Direct PCR of whole blood and hair shafts by microwave treatment. Biotechniques 17, 726–728 (1994).
Schmitteckert, E.M., Prokop, C.M. & Hedrich, H.J. DNA detection in hair of transgenic mice—a simple technique minimizing the distress on the animals. Lab. Anim. 33, 385–389 (1999).
Busler, D.E. & Li, S.W. Rapid screening of transgenic type II and type XI collagen knock-out mice with three-primer PCR. Biotechniques 21, 1002–1004 (1996).
Malumbres, M., Mangues, R., Ferrer, N., Lu, S. & Pellicer, A. Isolation of high molecular weight DNA for reliable genotyping of transgenic mice. Biotechniques 22, 1114–1149 (1997).
Ren, S., Li, M., Cai, H., Hudgins, S. & Furth, P.A. A simplified method to prepare PCR template DNA for screening of transgenic and knockout mice. Contemp. Top. Lab. Anim. Sci. 40, 27–30 (2001).
Cinelli, P., Rettich, A., Seifert, B., Bürki, K. & Arras, M. Comparative analysis and physiological impact of different tissue biopsy methodologies used for the genotyping of laboratory mice. Lab. Anim. 41, 174–184 (2007).
Wilhelm, J. & Pingoud, A. Real-time polymerase chain reaction. Chembiochem 4, 1120–1128 (2003).
Institute for Laboratory Animal Research. Guide for the Care and Use of Laboratory Animals (National Academies Press, Washington, DC, 1996).
Schunck, B., Kraft, W. & Truyen, U. A simple touch-down polymerase chain reaction for the detection of canine parvovirus and feline panleukopenia virus in feces. J. Virol. Methods 55, 427–433 (1995).
Safadi, F.F. et al. Osteopathy and resistance to vitamin D toxicity in mice null for vitamin D binding protein. J. Clin. Invest. 103, 239–251 (1999).
Freeman, B. et al. DNA from buccal swabs recruited by mail: evaluation of storage effects on long-term stability and suitability for multiplex polymerase chain reaction genotyping. Behav. Genet. 33, 67–72 (2003).
Hankenson, F.C., Garzel, L.M., Fischer, D.D., Nolan, B. & Hankenson, K.D. Evaluation of tail biopsy collection in laboratory mice (Mus musculus): vertebral ossification, DNA quantity, and acute behavioral responses. J. Am. Assoc. Lab. Anim. Sci. 47, 10–18 (2008).
Wilson, W.O., Riskin, D.K. & Mott, K.M. Ultrasonic sound as an indicator of acute pain in laboratory mice. J. Am. Assoc. Lab. Anim. Sci. 47, 8–10 (2008).
Acknowledgements
We thank Jessica Knowlton (formerly of the UM Orthopedic Research Laboratories) and Shawn Terkhorn and Alice Huang (formerly of the University of Pennsylvania) for technical assistance. We thank the American College of Laboratory Animal Medicine Foundation and the Universities of Michigan and Pennsylvania for financial support. K.D.H. was supported by National Institutes of Health grants R01 AR49862 and K01 RR RR00161.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Garzel, L., Hankenson, F., Combs, J. et al. Use of quantitative polymerase chain reaction analysis to compare quantity and stability of isolated murine DNA. Lab Anim 39, 283–289 (2010). https://doi.org/10.1038/laban0910-283
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/laban0910-283