Abstract
Aphids transmit viruses and are destructive crop pests1. Plants that have been attacked by aphids release volatile compounds to elicit airborne defence (AD) in neighbouring plants2,3,4,5. However, the mechanism underlying AD is unclear. Here we reveal that methyl-salicylate (MeSA), salicylic acid-binding protein-2 (SABP2), the transcription factor NAC2 and salicylic acid-carboxylmethyltransferase-1 (SAMT1) form a signalling circuit to mediate AD against aphids and viruses. Airborne MeSA is perceived and converted into salicylic acid by SABP2 in neighbouring plants. Salicylic acid then causes a signal transduction cascade to activate the NAC2–SAMT1 module for MeSA biosynthesis to induce plant anti-aphid immunity and reduce virus transmission. To counteract this, some aphid-transmitted viruses encode helicase-containing proteins to suppress AD by interacting with NAC2 to subcellularly relocalize and destabilize NAC2. As a consequence, plants become less repellent to aphids, and more suitable for aphid survival, infestation and viral transmission. Our findings uncover the mechanistic basis of AD and an aphid–virus co-evolutionary mutualism, demonstrating AD as a potential bioinspired strategy to control aphids and viruses.
Similar content being viewed by others
Main
Plants have evolved the ability to sense environmental stimuli and induce changes in metabolism to generate volatile organic compounds (VOCs) as intraplant and interplant signals2,6. VOCs, once released from VOC-emitter plants, can function as aerial cues and be perceived to elicit defences in neighbouring receiver plants, a phenomenon called AD2,3,4,5. Such plant–plant communication (PPC) and its biological and ecological importance have been observed in many species over decades2,3. Recently, UDP-arabinosyltransferase was found to be involved in airborne volatile reception in tomato distal defence against cutworms7. However, the molecular genetic framework for the VOC-mediated PPC, including AD, is largely unclear2. Moreover, with the exception of the receptor for ethylene8, the receptors that mediate the VOC-sensing system in plants have remained unidentified.
Aphids are the most destructive agricultural and horticultural pests worldwide1. Aphids are phloem-feeding and cause extensive destruction to crop production owing to their efficient transmission of numerous devastating viral pathogens1. More than 40% of plant viruses depend on aphids for transmission to infect plants, including many food, fruit and vegetable crops9. Aphid attack induces plants to emit VOCs, which are mainly composed of MeSA10,11,12,13. MeSA has been implicated in plant defence against herbivorous insects including aphids by repelling, attracting predators or reducing the survival fitness of these insects14,15,16. MeSA, which is known as a within-plant and long-distance mobile signal, is involved in inducing systemic acquired resistance (SAR) to microbial pathogens and herbivorous insects17,18. During SAR, salicylic acid (SA) accumulates in pathogen-infected cells and is converted into MeSA by SAMT119; MeSA then travels to distal tissues through the phloem and is subsequently reconverted into SA by SABP2 for SAR in systemic leaves20. Although MeSA’s function as a within-plant signal for SAR is known, how MeSA acts as an interplant communication signal to activate anti-aphid defence in neighbouring receiver plants is a long-standing and unresolved question2,10,16. For example, it is unclear whether plants possess receptor systems to sense and perceive airborne MeSA3. The mechanism for priming MeSA production during aphid infestation is also not well understood. Furthermore, it remains to be elucidated whether and how aphids and viruses can regulate AD, although MeSA has been reported to mediate AD against tobacco mosaic virus (TMV, a Tobamovirus in the family Virgaviridae)4. Here we took advantage of a pathosystem comprising aphid, virus, and VOC emitter and receiver plants to dissect AD. We established the framework for MeSA mediating AD to suppress insect infestation and virus transmission at the molecular and genetic levels, and identified an odorant-binding protein (OBP)-like receptor for sensing and perceiving airborne MeSA. We also found a viral counterdefence strategy to suppress plant AD, suggesting an undefined ecological and co-evolutionary mutualism between aphids and aphid-transmissible viruses. Moreover, our findings lay the groundwork to provide a detailed mechanism of VOC-triggered PPC and will pave the way for future studies on PPC and plant adaptation to the environment.
Plant antiviral defence requires NAC2
Following our work on cucumber mosaic virus (CMV), a Cucumovirus in the family Bromoviridae21, we investigated the role of CMV 1a protein (hereafter, CMV1a) in viral pathogenesis. By conducting immuno-pulldown coupled with mass spectrometry (MS) analysis, we identified the Nicotiana benthamiana transcription factor NAC2 as a CMV1a interactor (Extended Data Fig. 1a). NACs constitute a large plant-specific transcription factor family, some of which participate in the regulation of within-plant immunity22. We searched the N. benthamiana genome database and found two NAC2 homologues, NAC2.1 and NAC2.2, that share 96.7% nucleotide identity and should represent two alleles of one gene from different ancestry of allotetraploid N. benthamiana. We focused mainly on NAC2.1 (hereafter, NAC2) for further analyses. The CMV1a–NAC2 interaction was verified by co-immunoprecipitation (co-IP), bimolecular florescence (BiFC) and luciferase complementation imaging (LCI) assays (Extended Data Fig. 1b–d). We evaluated whether NAC2 affects CMV infection in NAC2.1/NAC2.2 double-knockout (KO) mutants (nac2 plants), which were generated using CRISPR–Cas9 gene editing (Extended Data Fig. 2a) versus wild-type (WT) plants. CMV infection resulted in more severe symptoms and a higher accumulation of viral RNA and coat protein (CP) in nac2 plants than in WT plants (Extended Data Fig. 1e–g). Similar results were also obtained in nac2 versus WT plants infected with GFP-tagged potato virus Y (PVY–GFP, a Potyvirus in the family Potyviridae) or GFP-tagged TMV (TMV–GFP) (Extended Data Fig. 1h–m). These data suggest that NAC2 is essential for plant antiviral defence.
NAC2 mediates AD against aphids through MeSA
We noticed that many more apterous green peach aphids colonized on nac2 leaves than WT leaves. This led us to examine the role of NAC2 in plant attractiveness to aphids. We performed circular-dish and Y-tube olfactometer bioassays and found that nac2 plants attracted more aphids (Myzus persicae, Sulzer) than did WT plants, probably mediated by airborne signals (Extended Data Fig. 1n,o). To determine which volatile was alterably generated, we used gas chromatography coupled with MS (GC–MS) to identify volatiles emitted by aphid-attacked WT versus nac2 plants. MeSA was the only constant VOC that was differentially produced in aphid-attacked WT versus nac2 plants, and more MeSA was emitted by WT than nac2 plants after aphid infestation (Extended Data Fig. 2e–h). Consistently, MeSA is a well-documented major—or, in certain plant species, the only—aphid-inducible VOC10,11,12,13. To test whether the effect of NAC2 on plant attractiveness to aphids is attributed to MeSA emission, we measured the emission rate of aerial MeSA from aphid-attacked WT plants using GC–MS and found that aerial MeSA was emitted at approximately 34 ng h−1 (equivalent to 0.816 µg per day) per aphid-attacked WT plant (Extended Data Fig. 2f). Furthermore, we found that the aerial MeSA concentrations in chambers containing either 0.8 µg MeSA-trapped lanolin paste (MeSA/lanolin) or an aphid-attacked WT plant were similar (Extended Data Fig. 2i,j). Thus, we used 0.8 µg MeSA/lanolin per chamber, which is equal to approximately 3 nM for the following experiments. After smearing plants with MeSA/lanolin, nac2 and WT plants showed a similar attractiveness to aphids (Extended Data Fig. 1p,q). However, when treated with lanolin alone or lanolin with other volatiles such as 3,3-dimethyl-hexane, nac2 plants were more attractive than WT plants to aphids (Extended Data Fig. 1r,s). We also kept nac2 and WT plants under volatile MeSA for 24 h followed by ventilation for 2 h, and compared how gaseous MeSA affects plant attraction of aphids. Under such conditions, WT plants were more repellent to aphids (Extended Data Fig. 1t,u). However, no obvious difference in aphid repellence was observed between nac2 plants with and without volatile MeSA treatment followed by ventilation (Extended Data Fig. 1v,w). Furthermore, like that observed without MeSA treatment, nac2 plants remained more attractive to aphids compared with WT plants after volatile MeSA treatment followed by ventilation (Extended Data Fig. 1x,y).
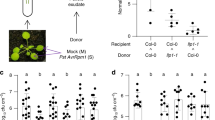
To decipher the reasons behind this phenomenon (Extended Data Fig. 1t–y), we treated nac2 or WT plants with volatile MeSA for 24 h followed by ventilation, and then quantified volatized MeSA emitted by MeSA-receiving plants (receivers). WT but not nac2 plants emitted higher levels of airborne MeSA (Fig. 1a,b). We next compared aphid attractiveness in MeSA receivers versus mock WT plants after smearing all of the plants with MeSA/lanolin, but found that they showed no difference in aphid preference (Fig. 1c,d). Moreover, after smearing both nac2 and WT receivers with MeSA/lanolin, these MeSA receivers also equally attracted aphids (Fig. 1e,f). We next used aphid-attacked plants as emitters to investigate the role of NAC2 in plant AD under a natural open-air environment (Extended Data Fig. 3a). After infestation by sap-sucking aphids, WT plants constantly emitted VOC MeSA (Fig. 1g,h). Notably, nac2 receivers released less volatile MeSA, but exhibited higher attractiveness to aphids compared with WT receivers when the emitters were attacked by aphids (Fig. 1i–l). WT receivers neighbouring aphid-attacked emitters were more repellent to aphids compared with those neighbouring mock emitters, while no significant difference in aphid repellence was found between nac2 receivers neighbouring mock- and aphid-attacked emitters (Fig. 1m,n). Furthermore, aphid-feeding in emitters reduced aphid survival in WT but not nac2 receivers after 24 h feeding on receivers (Fig. 1o,p). These results suggest that, once MeSA is perceived, MeSA biosynthesis in neighbouring receivers is regulated in an NAC2-dependent manner to mediate AD against aphids.
a,b, WT but not nac2 plants emitted a higher level of airborne MeSA after MeSA treatment followed by ventilation. GC–MS analysis (a) and quantification (b) are shown. FW, fresh weight. c,d, After smearing plants with MeSA/lanolin, WT and volatile-MeSA-treated WT plants exhibited similar attractiveness to aphids in circular-dish (c) or Y-tube olfactometer (d) bioassays. e,f, After smearing plants with MeSA/lanolin, volatile-MeSA-treated WT and nac2 plants exhibited similar attractiveness to aphids in circular-dish (e) or Y-tube olfactometer (f) bioassays. g–j, GC–MS analysis (g,i) and quantification (h,j). Aphid sap-sucking caused the attacked WT emitters to volatilize MeSA (g,h). When neighbouring AE emitters (aphid-attacked WT plants), the WT receivers (WT-R) volatilize more MeSA compared with nac2 receivers (nac2-R) (i,j). k,l, nac2-R/AE plants exhibited higher attractiveness to aphids compared with WT-R/AE plants in circular-dish (k) or Y-tube olfactometer (l) bioassays. m,n, WT-R/AE plants showed higher aphid repellence compared with WT-R/mE (WT-R plants neighbouring non-aphid attacked mock emitter) plants (m), whereas there was no significant difference in aphid repellence between nac2-R/mE and nac2-R/AE plants (n) in Y-tube olfactometer bioassays. o,p, The proportion of living aphids on WT-R (o) or nac2-R (p) plants neighbouring AE or mE emitters. For c-f, k-n, the values inside each bar show the number of choice-making aphids. Statistical analysis was performed using two-sided Student’s t-tests (b, h and j), one-way analysis of variance (ANOVA) with the least significant difference (LSD) test (the letters A and B represent statistically different groups; P < 0.05) (o and p) and χ2 tests (d.f. = 1) (c–f and k–n); all P values are shown in figure. NS, not significant. Data are mean ± s.d. n = 3 (b, h and j) and n = 4 (o and p) biologically independent samples. Experiments were repeated at least three times with similar results.
NAC2 activates SAMT1 transcription
We next set up to dissect the molecular and genetic link between NAC2 and MeSA biosynthesis. As a transcription factor (Extended Data Fig. 3b), NAC2 was found to localize in the nucleus (Extended Data Fig. 3c). We then performed RNA-sequencing (RNA-seq) and comparative transcriptome analysis in WT and nac2 plants with or without aphid feeding, and identified numerous potential NAC2-regulated differentially expressed genes (Extended Data Fig. 4 and Supplementary Tables 1 and 2), of which SAMT1 was of particular interest (Extended Data Fig. 4e). SAMT1 RNA levels were lower in nac2 plants than in WT plants, regardless of whether these plants were attacked by aphids or not (Supplementary Tables 1 and 2). Knowing that SAMT1 converts SA into MeSA19, we quantified SAMT1 transcripts in nac2, HA-NAC2-overexpressing and WT leaf tissues to assess whether NAC2 transcriptionally modulates SAMT1 expression. A list of the primers used in quantitative PCR with reverse transcription (RT–qPCR) assays is provided in Supplementary Table 3. SAMT1 mRNA levels were substantially reduced in nac2 plants but increased in HA–NAC2-overexpressing plants compared with in WT plants (Extended Data Fig. 3d,e). Moreover, a luciferase reporter assay showed that NAC2 enhanced the transcription of reporter genes under the control of the SAMT1 promoter in vivo (Extended Data Fig. 3f). Chromatin immunoprecipitation followed by qPCR (ChIP–qPCR), yeast-one-hybrid and electrophoretic mobility shift assay (EMSA) analyses all demonstrated that NAC2 bound to the SAMT1 promoter at the putative NAC transcription-factor-binding site23 and activated reporter gene transcription (Extended Data Fig. 3g–i). Furthermore, transient overexpression of NAC2 increased MeSA production in plants (Extended Data Fig. 3j). These results suggest that a NAC2–SAMT1 module involves the regulation of MeSA biosynthesis in plants.
SA-activated NAC2–SAMT1 module elicits AD
To investigate whether NAC2 affects MeSA production in receivers through activating SAMT1 transcription, we evaluated NAC2 and SAMT1 mRNA levels in nac2,NahG (N. benthamiana expressing bacterial NahG, which encodes the SA-degrading enzyme salicylate hydroxylase) and WT plants with or without airborne treatment of volatized MeSA followed by ventilation. Exogenous MeSA substantially increased the levels of SAMT1 and NAC2 mRNAs in WT plants, but did not affect the levels of SAMT1 mRNA in nac2 plants (Fig. 2a,b). Plants can sense MeSA as a within-plant signal and convert intracellular MeSA into SA to elicit intracellular defence17,20. Considering that MeSA treatment did not substantially change the expression of NAC2 and SAMT1 or alter aphid repellence in NahG plants (Fig. 2a–c), we hypothesized that SA might be the cue to initiate NAC2 expression. To test this hypothesis, we first examined the effect of aphid attack on NAC2 expression and found that NAC2 transcription was upregulated in aphid-attacked plants (Fig. 2d). We next examined the effect of aphid attack on SA and MeSA production as well as SAMT1 expression in WT versus nac2 plants. In the absence of aphids, lower levels of MeSA accumulated in nac2 plants compared with in WT plants (Fig. 2f). After aphid infestation, SA levels increased in both WT and nac2 plants to similar extents; however, increases in cellular MeSA and SAMT1 expression were found only in WT plants and not in nac2 plants (Fig. 2e–g). Similar results were also found in WT receivers and nac2 receivers neighbouring aphid-attacked plants (Fig. 2h–j). These results suggest that NAC2 is required for aphid-directed induction or aphid-mediated volatile priming of SAMT1 expression and MeSA production. Moreover, externally applied SA upregulated NAC2 and SAMT1 expression in WT plants, but did not significantly alter the level of SAMT1 mRNA in nac2 plants (Fig. 2k,l). Exogenous SA also induced stronger MeSA volatilization and aphid repellence in WT plants compared with in nac2 plants (Fig. 2m–p), and increased the levels of NAC2 and SAMT1 transcripts in SAMT1-KO plants (samt1 plants; Fig. 2q and Extended Data Fig. 2b,c). samt1 plants were also more attractive to aphids, and externally applied SA did not induce aphid repellence in samt1 plants (Fig. 2r–t). Furthermore, volatile MeSA production was compromised in samt1 plants that were exposed to aphid attack (Fig. 3a,b). In further aphid behaviour experiments, we found that WT receivers neighbouring aphid-attacked WT emitters were more repellent to aphids compared with WT receivers neighbouring mock WT emitters, while no significant difference in aphid repellence was found in WT receivers neighbouring nac2 or samt1 emitters regardless of whether these emitters were exposed to aphids or not (Fig. 3c–e). These results further confirm the role of MeSA as a PPC signal for interplant AD. Collectively, our data demonstrate that SA can activate NAC2-SAMT1 transcription to increase MeSA volatilization in both emitter and receiver plants.
a,b, NAC2 or its deletion mutant (a) or SAMT1 (b) mRNA levels in WT, nac2 and NahG plants with or without treatment with volatile MeSA. c, Treatment with volatile MeSA induced aphid repellence in WT but not NahG plants in Y-tube olfactometer bioassays. d, NAC2 expression was enhanced in aphid-attacked WT plants. e–g, Free SA (e), MeSA (f) and SAMT1 expression (g) in WT or nac2 plants after aphid attacks. h–j, Free SA (h), MeSA (i) and SAMT1 expression (j) in WT-R or nac2-R plants neighbouring AE or mE emitters. k,l, NAC2 (k) or SAMT1 (l) expression in nac2 and WT plants with or without SA treatment. m,n, GC–MS analysis (m) and quantification (n) of volatized MeSA in WT or nac2 plants treated with SA for 24 h. o,p, SA treatment induced aphid repellence in WT (o) but not nac2 (p) plants in Y-tube olfactometer bioassays. q, NAC2 or SAMT1 expression in samt1 plants with or without SA treatment. r, samt1 plants exhibited higher attractiveness to aphids compared with WT plants in Y-tube olfactometer bioassays. s,t, SA treatment induced aphid repellence in WT (s) but not samt1 (t) plants in Y-tube olfactometer bioassays. Statistical analysis was performed using two-sided Student’s t-tests (a, b, d, g, j–l, n and q), one-way ANOVA with Tukey’s multiple-comparison test (the letters A–C represent statistically different groups; P < 0.05) (e, f, h and i) and χ2 tests (d.f. = 1) (c, o, p, r–t); exact P values are shown in the figure (a–d, g, j–l and n–t) or provided in the Source Data (e, f, h and i). Data are mean ± s.d. n = 3 (a, b, d–h, j–l, n and q) and n = 4 (i) biologically independent samples. Experiments were repeated at least three times with similar results.
a,b, GC–MS analysis (a) and quantification (b). Volatile MeSA production was compromised in samt1 plants that were exposed to aphid attack. c–e, WT-R/AE plants were more repellent to aphids compared with WT-R/mE plants (c), while no significant difference in aphid repellence was observed in WT-R plants neighbouring nac2 (nac2-AE; d) or samt1 (samt1-AE; e) emitters that were or were not exposed to aphids in Y-tube olfactometer bioassays. f, The SA-binding activity of purified SABP2 protein. g, SA competition binding assays with 0, 3 and 15 nM MeSA. The binding activity of [3H]SA to SABP2 in the presence of 3 nM or 15 nM MeSA was calculated against the base level. h, sabp2 plants showed no abnormal developmental phenotypes. i, Volatile MeSA treatment induced aphid repellence in WT plants, but not in sabp2 plants in Y-tube olfactometer bioassays. j, Treatment with volatile MeSA increased the amount of free SA in WT plants but not in sabp2 plants. k, WT-R plants, but not sabp2 receivers (sabp2-R), neighbouring WT-mE mock emitters exhibited higher attractiveness compared with those neighbouring WT-AE plants as determined using Y-tube olfactometer bioassays. l,m, GC–MS analysis (l) and quantification (m). No difference in the amount of volatized MeSA was observed between WT and sabp2 plants after SA treatment. n,o, MeSA treatment rescued the virus-hypersusceptibility phenotypes of nac2 or samt1 but not sabp2 plants, whereas SA treatment rescued the virus-hypersusceptibility phenotypes of nac2, samt1 and sabp2 plants. Statistical analysis was performed using two-sided Student’s t-tests (b, g, j and m–o) and χ2 tests (d.f. = 1) (c–e, i and k); all P values are shown in the figure. Data are mean ± s.d. n = 3 (b, f and m–o) and n = 4 (g and j) biologically independent samples. Experiments were repeated at least three times with similar results.
As an SA-binding protein, SABP2 can also bind to MeSA and is essential for the conversion of intracellular MeSA into SA17,20. Thus, SABP2 may act as an OBP-like receptor that perceives and converts the volatized MeSA generated from emitters into SA to trigger NAC2-mediated aphid resistance in receivers. To test this idea, we first confirmed that SABP2 bound to SA (Fig. 3f), and then examined whether MeSA could influence the specific SABP2 SA-binding activity in competition binding assays (Fig. 3g). We set the SABP2-[3H]SA (50 Ci mmol−1) binding capacity in the absence of competitor MeSA as 100%. However, under the same experimental conditions, the binding activity of [3H]SA by SABP2 reduced to approximately 74% and 46% in the presence of 3 nM and 15 nM MeSA, respectively (Fig. 3g). Thus, 3 nM MeSA was sufficient to compete with [3H]SA for binding to SABP2, suggesting that MeSA can bind to SABP2 at a physiological concentration. We next generated SABP2-KO lines (sabp2 plants) and tested aphid repellence in sabp2 versus WT receivers that were treated with volatized MeSA followed by ventilation. Treatment with volatile MeSA increased aphid repellence and SA biosynthesis in WT but not sabp2 receivers (Fig. 3h–j and Extended Data Fig. 2d). Furthermore, aphid feeding in WT emitters increased aphid repellence in WT but not sabp2 receivers (Fig. 3k). Moreover, after external applications of SA, no difference in the volatized MeSA amount was found between WT and sabp2 plants, indicating that SABP2 is not required for MeSA emission (Fig. 3l,m). These results demonstrate that SABP2 is indeed an OBP-like receptor that perceives and converts airborne MeSA into SA to elicit NAC2-mediated aphid repellence in receiver plants.
SAMT1 is required for plant antiviral defence24. To test whether SAMT1 is a component in NAC2-mediated plant antiviral defence. We knocked-down (KD) NAC2 in samt1 plants to generate nac2/samt1 double-mutant plants using tobacco rattle virus (TRV)-based virus-induced gene silencing. Like nac2 mutants, NAC2-KD plants (in the WT N. benthamiana background) showed normal growth and were susceptible to CMV and PVY (Extended Data Figs. 1e–j and 5a–e,h–l), suggesting that NAC2-KD by virus-induced gene silencing mimics NAC2-KO. However, nac2/samt1 and samt1 plants showed similar degrees of CMV or PVY infection (Extended Data Fig. 5a–e,h–l). Furthermore, CMV infection enhanced the intracellular MeSA level in plants (Extended Data Fig. 5f); NAC2-KD, samt1 and nac2/samt1 plants produced similar amounts of MeSA, albeit lower amounts compared with WT plants, in the systemic leaves during virus infection (Extended Data Fig. 5g). Moreover, we investigated whether and how MeSA is responsible for NAC2-mediated plant antiviral defence by treating nac2, sabp2, samt1 and WT plants with external application of either MeSA or SA. Compared with WT plants, nac2, samt1 and sabp2 plants were more susceptible to CMV and PVY (Fig. 3n,o). However, exogenous treatment of MeSA rescued the virus-hypersusceptibility phenotypes of nac2 or samt1 plants but not sabp2 plants (Fig. 3n,o), probably due to MeSA conversion into SA in WT, nac2 and samt1 plants but not in sabp2 plants. External spray of SA rescued the virus-hypersusceptibility phenotypes of nac2, samt1 and sabp2 plants (Fig. 3n,o), consistent with the fact that SA can suppress infection of plants by many viruses including CMV, potato virus X and TMV25.
Taken together, our results demonstrate that SA can trigger the NAC2–SAMT1 module to increase in vivo MeSA production by activating the NAC2-driven transcription of SAMT1. SA converted from perceived MeSA by SABP2 is the cue to elicit AD against aphids and viruses in receiver plants.
CMV1a destabilizes NAC2 to suppress AD
Aphid-induced PPC and AD may have an impact on viral transmission by aphids. To investigate this, we fed CMV-bearing aphids on WT-R/AE plants (WT receivers (WT-R) neighbouring WT emitters damaged by virus-free aphids (AE)) or WT-R/mE plants (WT-R plants neighbouring non-aphid damaged WT mock emitters (mE)) and examined the aphid feeding behaviours on these plants using electrical penetration graph (EPG) analysis. EPG analysis showed that the number of short intracellular punctures (potential drop (pd)) of individual aphids, which is responsible for non-persistent aphid transmission of CMV, was lower in WT-R/AE plants than in WT-R/mE plants (Fig. 4a), suggesting that AD reduced aphid transmission of CMV in WT-R/AE plants. Furthermore, WT-R/AE or WT-R/mE plants were exposed to feeding by viruliferous aphids for 2 h, then all of the aphids were removed from the plants and the viral accumulation in these plants was tested 24 h later. We found that WT-R/AE plants accumulated less CMV than WT-R/mE plants, evidenced by a reduction in viral RNA and CP (Fig. 4b,c). Moreover, fewer viruliferous aphids survived in WT-R/AE plants compared with in WT-R/mE plants that were exposed to feeding by viruliferous aphids for 24 h (Fig. 4d). These results suggest that PPC elicits AD against aphids (including repellent and survival) and reduces their ability to transmit CMV. We also tested the effect of using ACE emitters (plants attacked by CMV-carrying aphids) on AD, and found that the pd number of individual aphids, the CMV accumulation and the proportion of living aphids were similar in WT-R/ACE and WT-R/mE plants (Fig. 4e–h). These data suggest that CMV infection can suppress interplant aphid-induced AD. However, the short probing frequency of aphids differs in WT versus CMV-infected plants26. By contrast, no difference in short-probe frequency was observed in aphid-fed WT versus CMVΔ2b-infected plants26. Nevertheless, to ensure that the effect of CMV infection on aphid-induced AD was not caused by the difference in probing frequency between the virus-free and viruliferous aphids, we conducted EPG experiments and found that the pd number was lower for virus-free aphids compared with CMV-carrying aphids, but similar for virus-free and CMVΔ2b-carrying aphids (Fig. 4i), consistent with a previous report26. We next exposed WT-R/mE, WT-R/AE, WT-R/ACE and WT-R/ACΔE (WT receivers neighbouring WT emitters that were attacked by aphids carrying CMVΔ2b (ACΔE)) plants to feeding by viruliferous aphids for 2 h, and tested the viral accumulation as well as the aphid survival in these plants. We found that AD against aphids and viruses was attenuated in WT-R plants that were positioned near to either ACE or ACΔE emitters (Fig. 4j–l). We next investigated whether virus infection affects MeSA production in plants. GC–MS analysis revealed that plants that were exposed to feeding by CMV-carrying aphids or CMVΔ2b-carrying aphids emitted less MeSA compared with plants that were exposed to feeding by virus-free aphids (Fig. 4m,n). Taken together, these data suggest that CMV infection can suppress aphid-induced AD for the benefit of aphid survival and virus transmission/infection, probably through CMV-mediated interference of MeSA production.
a, EPG analysis showed that the pd number in aphid feeding was less in WT-R/AE plants compared with in WT-R/mE plants when the receivers were exposed to feeding by CMV-carrying aphids. n = 21 individual aphids. b–d, The levels of CMV RNA (b) and CP (c) and the proportion of living aphids (d) in WT-R/mE and WT-R/AE plants. e, The pd number was similar between WT-R/mE plants and WT-R/ACE plants (WT-R receivers neighbouring emitters that had been attacked by CMV-carrying aphids) when the receivers were exposed to feeding by CMV-carrying aphids. n = 20 individual aphids. f–h, The levels of CMV RNA (f) and CP (g) and the proportion of living aphids (h) is similar between WT-R/mE and WT-R/ACE plants. i, The pd number of virus-free aphids was lower than that of CMV-carrying aphids, but similar to that of CMVΔ2b-carrying aphids. n = 19 individual aphids. j–l, The levels of CMV RNA (j) and CP (k) and the proportion of living aphids (l) in WT-R/mE, WT-R/AE, WT-R/ACE or WT-R/ACΔE (WT-R receivers neighbouring WT emitters attacked by CMVΔ2b-carrying aphids) plants. m,n, GC–MS analysis (m) and quantification (n) of volatized MeSA in WT plants exposed to feeding by virus-free aphids, CMV-carrying aphids or CMVΔ2b-carrying aphids for 3 days. Statistical analysis was performed using two-sided Student’s t-tests (a, b, e, f, i, j and n), one-way ANOVA with LSD test (d and h), one-way ANOVA with Tukey’s multiple-comparison test (l) (the letters A and B represent statistically different groups; P < 0.05); exact P values are shown in the figure (a, b, d–f, h–j and n) and in the Source Data (l). n = 3 biologically independent samples (b, d, f, h, j, l and n). Data are mean ± s.d. (b, d, f, h, j, l and n). For the box plots in a, e and i, the centre line represents the median, the box limits show the bottom and top quartiles, and the whiskers show the highest and lowest data points. Experiments were repeated at least three times with similar results.
The CMV1a–NAC2 interplay (Extended Data Fig. 1a–d) suggests that CMV1a might be involved in CMV-mediated suppression of AD. To test this, we first generated transgenic N. benthamiana expressing CMV1a (Extended Data Fig. 6a,b) and evaluated the effect of CMV1a on plant attractiveness to aphids and AD. Our circular-dish and Y-tube olfactometer bioassays showed that CMV1a expression caused higher plant attractiveness to aphids (Extended Data Fig. 6c,d). EPG assays revealed that the pd number of individual aphids was higher in CMV1a versus WT plants (Extended Data Fig. 6e). Moreover, when neighbouring 1a-AE emitters (CMV1a plants attacked by virus-free aphids), the AD of WT-R plants against aphids and viruses was suppressed (Extended Data Fig. 6f–h). Furthermore, WT-R plants neighbouring WT-mE mock emitters (non-aphid-attacked WT plants) exhibited higher attractiveness to aphids compared with WT-R plants adjacent to WT-AE emitters (WT plants attacked by virus-free aphids). However, WT-R plants exhibited similar attractiveness to aphids when neighbouring 1a-mE (transgenic CMV1a plants that were not attacked by aphids) or 1a-AE (transgenic CMV1a plants that were attacked by virus-free aphids) emitters in Y-tube olfactometer bioassays (Extended Data Fig. 6i,j). Moreover, WT-R plants exhibited higher attractiveness to aphids when neighbouring 1a-AE versus WT-AE emitters (Extended Data Fig. 6k). These data imply that CMV1a is involved in CMV-mediated suppression of AD.
To understand the importance of the CMV1a–NAC2 interaction in CMV-mediated AD suppression, we identified a key amino acid in CMV1a that is responsible for its interaction with NAC2. CMV1a consists of an N-terminal methyltransferase and a C-terminal ATP-dependent helicase domain (HD). We found that the CMV1a HD is responsible for the CMV1a–NAC2 interaction as determined using LCI (Extended Data Fig. 6l). We also modelled the structure of the CMV1a–NAC2 complex using AlphaFold-Multimer27 and observed that the glycine residue at position 983 in CMV1a has the nearest physical proximity to NAC2, predicting that this residue might be essential for CMV1a to interact with NAC2 (Extended Data Fig. 6m,n). Indeed, a G983D mutation in CMV1a HD or full-length CMV1a substantially impaired the CMV1a–NAC2 interaction in co-IP or BiFC assays (Extended Data Fig. 6o–q).
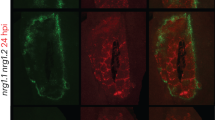
We next investigated the subcellular localization of the CMV1a–NAC2 interaction using BiFC and found that CMV1a interacted with NAC2 in both the nucleus and cytoplasm (Extended Data Fig. 1c), which is different from NAC2 localization without CMV1a co-expression (Extended Data Fig. 3c), suggesting that CMV1a can relocate some NAC2 from nucleus to cytoplasm. Consistently, CMV1a–MYC, but neither cLUC–MYC nor CMV1a(G983D)–MYC, partially caused cytoplasmic localization of RFP–NAC2 and less RFP fluorescence in the nucleus (Extended Data Fig. 7a). Notably, CMV1a–MYC did not alter RFP nuclear localization, indicating that CMV1a–MYC-directed NAC2 relocalization depends on the NAC2–CMV1a interaction (Extended Data Fig. 7b). Furthermore, we investigated the stability of cytoplasmic NAC2 using nuclear export signal (NES)-tagged NAC2. We found that NES–NAC2 was localized in the cytoplasm and was subjected to higher 26S-proteasome-system-mediated degradation compared with WT NAC2 (Extended Data Fig. 7c,d). Moreover, transient CMV1a expression enhanced NAC2 degradation by the 26S proteasome system but did not affect RFP stability (Extended Data Fig. 7e–i), whereas CMV1a(G983D) did not cause NAC2 degradation (Extended Data Fig. 7f–i).
Further transient expression assays showed that CMV1a but not CMV1a(G983D) suppressed NAC2-mediated activation of the SAMT1 promoter (Extended Data Fig. 7j). Furthermore, Y-tube olfactometer bioassays and GC–MS analysis in CMV1a(G983D) or CMV1a transgenic versus WT plants showed that CMV1a(G983D) impaired CMV1a-mediated plant attractiveness to aphids and suppression of MeSA volatilization (Extended Data Fig. 7k–m). Moreover, when neighbouring 1a(G983D)-mE (non-aphid-attacked transgenic CMV1a(G983D) plants) or 1a(G983D)-AE (transgenic CMV1a(G983D) plants attacked with virus-free aphids) emitters, WT-R plants near the 1a(G983D)-mE plants exhibited higher attractiveness to aphids compared with WT-R plants adjacent to 1a(G983D)-AE plants (Extended Data Fig. 7n). These Y-tube olfactometer bioassays provide additional evidence confirming that the amino acid residue Gly983 is essential for CMV1a to suppress the interplant AD.
Taken together, our data suggest that CMV1a interferes with AD by affecting the subcellular localization and stability of NAC2 through its direct interaction with NAC2 to impair NAC2-driven SAMT1 transcription and MeSA production.
Some aphid-borne viruses suppress AD
CMV1a—which has methyltransferase and helicase activity and forms part of the viral replicase complex—interacts with NAC2 through its HD (Extended Data Figs. 1a–d and 6l–q). We aligned the HDs of viral replicases and other HD-containing proteins of multiple plant viruses transmitted by aphids or other vectors. Notably, the HDs from many aphid-transmitted viruses, including Potyvirus, Cucumovirus, Luteovirus and Alfamovirus, contain a conserved glycine residue at the position corresponding to CMV1a Gly983 (Extended Data Figs. 8 and 9a). We further confirmed that PVY infection affected plant MeSA volatilization after aphid attack. We tested the effect of using AE emitters on AD against PVY transmission, and found that WT-R/AE plants accumulated less PVY compared with WT-R/mE plants, evidenced by the reduction in viral RNA and CP (Extended Data Fig. 9b,c). Moreover, GC–MS analysis revealed that plants that were exposed to feeding by PVY-carrying aphids emitted less MeSA compared with plants that were exposed to feeding by virus-free aphids (Extended Data Fig. 9d,e). We also tested the effect of APE emitters (emitter plants attacked by PVY-carrying aphids) on AD, and found that the PVY accumulation and the proportion of living aphids were similar between WT-R/APE and WT-R/mE plants (Extended Data Fig. 9f–h). We also showed that two aphid-transmitted viruses, CMV and PVY, enable NAC2 to relocate from the nucleus to cytoplasm (Extended Data Fig. 9i). Similarly, PVY CI, but not CI(G347D) or the 126KD protein of the non-aphid transmitted virus TMV, interacted with NAC2 (Extended Data Fig. 9j,k) and partially impaired the nuclear localization of NAC2 (Extended Data Fig. 9l). These results reveal that some aphid-transmitted viruses have evolved to deploy HD-containing proteins as a general strategy to interfere with plant AD.
Discussion
Insect-attacked plants emit VOCs as interplant signals to elicit AD in non-attacked neighbouring receiver plants11,28,29,30. However, it is unclear how the receiver plants perceive VOCs from neighbouring stressed emitter plants and activate signal transduction to elicit AD3. Here we reveal the molecular genetic mechanism underlying AD against aphids in plants.
In contrast to chewing herbivore attacks, phloem-feeding insects such as aphids avoid or suppress the jasmonate-based defence response and, instead, they elicit the SA response and increase the SA level in attacked plants31,32. As outlined in our model (Extended Data Fig. 10a), NAC2 can be activated by an upregulated SA level and directly bind to the SAMT1 promoter, activating its transcription (Fig. 2k and Extended Data Fig. 3d–i). SAMT1 then catalyses SA to form MeSA19,24. Consistently, MeSA is a predominant VOC induced by aphid attack in numerous plant species such as bean, chilli and peach, but it is very low or absent in the VOC emissions induced by beetle attack or artificial wounding10,11,12,13,33. The OBP-like receptor for perceiving airborne MeSA in receiver plants is unclear3. SABP2 is thought to be a potential receptor for airborne MeSA because it binds to MeSA3,17,20. However, SABP2 has only been reported to operate in the same plants exposed to stress (that is, within-plant signalling)3. Here we show that SABP2 is an OBP-like receptor for perceiving airborne MeSA from emitter plants and converting MeSA into SA in receiver plants (between-plant signalling) (Fig. 3f–j). Thus, plants may possess an OBP-like receptor-mediated sense system to perceive diverse airborne VOCs3. As MeSA can mediate resistance to many insects, including aphids, through attracting their predators or parasitoids in multiple plant species34,35,36,37,38, MeSA-perceived plants are also likely to attract aphid parasitoids or predators through volatile MeSA. MeSA-perceived plants repel aphids and reduce aphid survival fitness (Figs. 1m,o and 4d and Extended Data Fig. 1t,u). Thus, NAC2–SAMT1-mediated airborne PPC represents a multifunctional strategy to prevent aphid epidemic infestation.
Arabidopsis SA methyltransferase BSMT1 is strongly induced by jasmonate and biotic-stress-induced MeSA production, and is therefore severely compromised in jasmonate pathway mutants, whereas SA signalling mutants such as npr1 or pad4 do not show compromised MeSA emission39. This is different to the regulation of NbSAMT1 by SA described here. Nevertheless, our finding of SAMT1 induction by exogenous SA, intracellular SA after aphid attack or aphid-mediated volatile priming through NAC2 in N. benthamiana (Fig. 2a–l and Extended Data Fig. 3d–j) is consistent with the fact that SAMT1 can be induced by SA or insect feeding in Atropa belladonna and rice40,41 as well as in Nicotiana alata and Nicotiana sylvestris42. It is possible that diverse species may have evolved distinctive mechanisms to regulate BSMT1 versus SAMT1 expression. MeSA biosynthesis can also respond to environmental biotic and abiotic stresses in a plant-species-specific manner.
MeSA has been extensively studied as a long-distance mobile signal and has been shown to be involved in the induction of SAR to viruses and aphids, and to be herbivore-induced within plants11,12,14,17. Here we show that the NAC2–SAMT1 module participates in intraplant and interplant defence against CMV, PVY and TMV (Extended Data Figs. 1e–m and 5a–e,h–l). MeSA-mediated AD induces aphid antixenosis and suppresses virus transmission by aphids in neighbouring receiver plants (Fig. 4a–d). Thus, the NAC2–SAMT1 module is important not only for plant intra/intercellular antiviral defence but also for AD against aphids and viral transmission among plants. We have therefore unravelled an NAC2-dependent role of MeSA in PPC and AD.
Our molecular insights into AD against aphids and viruses raise questions about whether and how viruses counteract AD and affect aphid survival in receiver plants. In some cases, virus infection benefits its insect vector in virus-infected plants. For examples, TYLCCNV-encoded βC1 protein suppresses the jasmonate pathway or reduces terpene synthase and alters glucosinolate profiles to increase whitefly Bemisia tabaci attraction and performance in virus-infected plants43,44. CMV2b targets the plant jasmonate pathway or alters emission of VOC (not MeSA) to induce the attractiveness of virus-infected plants to aphids45,46. Moreover, CMV1a, which influences viral replicase complex assembly and viral systemic movement, prevents CMV2b-induced anti-aphid resistance (antibiosis), probably through limiting the CMV2b–Argonaute1 interaction47. CMV2a also stimulates feeding deterrence against aphids48. Furthermore, NIa-Pro encoded by PVY or TuMV increases ethylene production and inhibits callose accumulation to benefit insects in virus-infected plants49,50. However, reports about the mechanism underlying pathogen-induced regulation of MeSA-mediated plant AD2 are lacking. We reveal that some aphid-transmitted viruses can suppress MeSA emission in aphid-attacked plants to impair AD against virus infection and aphid infestation in adjacent plants (Fig. 4e–n and Extended Data Fig. 9b–h). This is achieved by the action of the helicase-domain-containing protein that suppresses AD through its interaction with NAC2 to partially change the subcellular localization of NAC2 from nucleus to cytoplasm, and to promote the NAC2 degradation through the 26S proteasome to block SAMT1 transcription and subsequent MeSA biosynthesis (Extended Data Fig. 10b). Thus, some aphid-transmitted viruses have evolved to use helicase-domain-containing proteins as a common means to suppress the transcriptional function of NAC2 and counteract AD in plants. These findings further suggest that not only can viruses use aphids as vector for transmission, but also aphids can in turn exploit viruses to facilitate their survival and ability to thrive in plants. This unexpected countermeasure reveals an undefined ecological and co-evolutionary mutualism between aphids and aphid-transmissible viruses (Extended Data Fig. 10).
Methods
Plant materials and growth conditions
N. benthamiana was used as the WT plant. NahG lines were transgenic N. benthamiana plants expressing bacterial NahG, which encodes the SA-degrading enzyme salicylate hydroxylase51. Transgenic CMV1a and CMV1aG983D lines are N. benthamiana lines overexpressing CMV1a or CMV1a(G983D) with a C-terminal fusion to a MYC tag. The nac2, samt1 and sabp2 plants were NbNAC2.1/2.2-, NbSAMT1- and NbSABP2-edited KO mutant N. benthamiana lines, respectively. All seeds were surface-sterilized and cultivated on MS medium (Murashige and Skoog medium, Sigma-Aldrich), grown at 26 °C under a 16 h–8 h light–dark photoperiod. Seedlings aged 10 days were then planted in soil and grown under the same photoperiod.
Generation of transgenic plants
The MYC-tagged full-length CMV1a and CMV1aG983D genes under the CaMV 35S promoter were cloned into a pCambia1300-based binary vector using the ligation-independent cloning strategy. All of the constructs were verified by DNA sequencing and transformed into Agrobacterium GV2260. To generate transgenic plants, leaf discs were plated onto MS medium in a 100 mm Petri dish (704001, NEST Biotechnology) with 0.1 mg ml−1 NAA (N8010, Solarbio), 1.0 mg ml−1 6-BA (IB0100, Solarbio), 0.2 mg ml−1 timentin and 0.025 mg ml−1 hygromycin B (400052, Merck Millipore) for callus, shoot and root regeneration. Western blot assays were used to select the T2 generation transgenic lines of 35S:CMV1a-MYC and 35S:CMV1aG983D-MYC plants, and selected transgenic plants were then used for aphid attraction bioassays and GC–MS analysis. The Cas9-edited knockout lines were generated as described previously52. In brief, the fragment containing single sgRNA (NAC2_LacZ/AtU3d-sgRNA, SABP2_LacZ/AtU3d-sgRNA) or two sgRNAs (SAMT1_LacZ/AtU3d-sgRNA1-At3b-sgRNA2) were cloned into pYLCRISPR/Cas9-DH to generate pYLCRISPR/Cas9-DH-NAC2, pYLCRISPR/Cas9-DH-SABP2 and pYLCRISPR/Cas9-DH-SAMT1. The sgRNA sequences are listed in Supplementary Table 3. The editing target site of the transgenic plant DNA sequence was characterized using PCR sequencing to select homozygous transgenic lines. PCR amplification was conducted using the 2×T5 Super PCR Mix (TSE005, Beijing Tsingke Biotech). Progenies of homozygous nac2, sabp2 and samt1 lines showed no developmental phenotype.
MeSA supplement treatment using the lanolin smear assay
Pure MeSA chemical (0.8 µg; M6752, Sigma-Aldrich) was dissolved in lanolin paste (S5106, Selleck). Plants for MeSA supplement treatment were smeared with MeSA-containing lanolin on the stems or leaf vein. MeSA-containing lanolin would release quantitative volatile MeSA continuously and constantly53,54.
MeSA supplement treatment using the airborne assay
Pure MeSA (0.8 µg) was dissolved in lanolin paste. Lanolin containing 0.8 µg MeSA was then smeared onto the surface of filter paper and the filter paper was placed in a 2 l volume gas-tight glass chamber. Plants for airborne MeSA supplement treatment were placed into these chambers separately to perceive volatile MeSA (each chamber contains one plant) for 24 h, and then transferred from chambers to open-air area for ventilation for 2 h before further experiments as described previously55.
Exogeneous SA supply treatment
Solutions of 50 µM SA (S5922, Sigma-Aldrich) were used for plant treatments. Seedlings (aged 4 weeks) were cultured in pots in a greenhouse and sprayed with the SA solutions.
Aphid choice bioassay using the Y-tube olfactometer assay
Wingless M. persicae aphids were isolated from N. benthamiana plants and starved for 2 h before testing. Plants grown in soil for 4 weeks were used. The experimental procedure and equipment were used as described previously46. For pairwise comparison of aphid attractiveness, plants in different groups were separately placed into two chambers, which were connected by the two arms of a glass Y-tube. The glass Y-tube was 1.0 cm in inner diameter and 6 cm in arm length. A humidified continuous air flow purified by activated charcoal flowed through the chambers at 200 ml min−1, then flowed into the arms of the Y-tube.
In each experiment, an individual aphid was placed at the end of the Y-tube stem. Aphids walked upwind towards the arms and made its choice. Once the aphid entered one arm and walked up to 3 cm away from the Y-junction, their choice was marked. To avoid any unknown asymmetry in the set-up, the plant-containing chambers were switched after every 10 tests, and the Y-tube and plants were replaced after 20 tests. The used Y-tubes were cleaned by ethanol and double-distilled H2O before reuse. The same number of aphids was used for each group in each experiment. All of the experiments were independently repeated at least three times.
Aphid choice bioassay using the circular-dish assay
Choice test for M. persicae was performed with detached leaves from 4- to 5-week-old N. benthamiana in Petri dishes (15 cm diameter) according to a previously described procedure56. Two leaves with a similar leaf position and size from experimental group plants and control group plants were detached and placed into Petri dishes, with their petioles inserted in moistened cotton swabs. Twenty wingless adult aphids were released at the midpoint between pairs of leaves, and the aphids on each leaf were counted at 24 h after their release in the Petri dishes. The same number of aphids was used for each group in each experiment. All of the experiments were independently repeated at least three times.
Interplant communication assays
Experimental plants were used as either emitters or receivers. The emitter and receiver plants were placed onto two trays at a distance of 30 cm from each other in an open-air experimental set-up for interplant communication assays. Each emitter was exposed to feeding by fifty virus-free or viruliferous M. persicae aphids or no aphids, then the emitters and receivers were incubated in same cage made by gauze. After 3 days, the receiver plants were taken out for further experiments. Viruliferous M. persicae aphids were obtained from aphids that were fed on CMV, CMVΔ2b or PVY-GFP-infected plants for 1 h before the preacquisition starvation period (2 h).
Aphid survival and virus-transmission analysis
To measure the effect of plant airborne defence on aphid fitness in neighbouring receiver plants, the experimental group receiver plants and mock receiver plants were exposed to feeding by the same number of wingless adult M. persicae. After 24 h, the numbers of alive aphids on each host plant was counted. To measure the influence of airborne defence on aphid-mediated virus-transmission in neighbour receiver plants, the experimental group receiver plants and control receiver plants were exposed to feeding by the same number of viruliferous adult aphids (carrying CMV or PVY) for 2 h as described previously57, then aphids were removed and, 24 h later, aphid-sucked leaves were taken to measure viral infection. All of the experiments were conducted at least three times.
EPG analysis of aphid feeding behaviour
The Giga-8 EPG amplifier (Wageningen University) was used to study the feeding behaviour of M. persicae on plants as described previously58. The amplifier with eight channels simultaneously recorded eight individual aphids on separate plants for 2 h in accordance with the inoculation access period (IAP) in viral transmission experiment. The EPG waveforms of short intracellular punctures (pd) were analysed during 2 h of aphid feeding with the software ‘Stylet + a’.
Measurement of plant endogenous free SA amount
Free SA was extracted and measured as described previously59. In brief, 0.2 g leaf tissue was homogenized in 1 ml of ethyl acetate containing 10 ng of SA-d6, and vortexed for 10 min followed by centrifuging at 18,000g at 4 °C for 20 min. The supernatants were collected and evaporated to dry by a vacuum concentrator at 30 °C. A total of 200 μl of MeOH:H2O (70:30, v/v) was added to resuspend the residues by centrifuging at 18,000g for 10 min. The collection of supernatants was analysed using a high-performance liquid chromatography–tandem MS system (QTRAP 4500, AB Sciex).
Measurement of plant endogenous MeSA
MeSA was extracted and measured by Suzhou Michy Biomedical Technology. In brief, frozen leaf tissues (150 mg) were homogenized with 200 µl of extraction buffer (water:1-propanol:HCl, 1:2:0.005). Then, 500 µl of methylene chloride was added to the homogenate and the sample was shaken thoroughly and then centrifuged at 18,800g for phase separation. Aliquots of the supernatants were separated using a GC system.
Volatile MeSA measurement by GC–MS analysis
VOCs emitted by plants were collected by a static-headspace sampling device with a solid-phase microextraction (SPME) fibre (PDMS/DVB, 65 μm) coated with polydimethylsiloxane/divinylbenzene (57345-U, Supelco) as described previously13. Each sample was enclosed in a 1.5 l glass container for 120 min, then the SPME fibre was extended into the headspace to collect volatiles for 30 min. After collection of volatile substances, the SPME fibre was inserted directly into a thermal desorption gas chromatograph–mass spectrometer (Thermo Fisher Scientific). The volatiles bound to the fibre were desorbed for 2 min. A Thermo Scientific Trace 1300 gas chromatograph equipped with a Thermo Scientific Q-Exactive Orbitrap mass spectrometer (GC-Orbitrap-MS, Thermo Fisher Scientific) was used for HRMS quantitative analysis. The split-splitless injection port was held at 250 °C for desorpting volatiles in split mode at a split ratio of 1:10. The oven temperature program was set as follows: 60 °C held for 2 min, then increased to 280 °C at 20 °C min−1 and held for 2 min.
MS analysis
Protein extracts were immunoprecipitated by MYC-Trap beads (M20012L, Abmart) and the mixture was incubated at 4 °C for 3 h. The precipitated samples were washed five times with co-IP buffer, and the immunoprecipitates were then separated by 10% SDS–PAGE. The gel was silver stained and gel-embedded protein samples were digested and analysed using LC–MS/MS.
Yeast-one-hybrid assay
The full-length cDNA sequence of NAC2 was fused in frame with the GAL4 activation domain in pGADT7-Rec2 (Clontech). The fusion construct was co-transformed with the reporter vector (pHIS2-cis/promoter of SAMT1) into Y187 yeast cells (Clontech). The empty vector pGADT7-Rec2 and the pHIS2-cis/35 S promoter were co-transformed as the negative controls. Co-transformed yeast strain Y187 was spotted onto selective medium (minus His, Leu and Trp) containing 0.5, 1 or 2 mM 3-AT. Images were taken after incubation for 3 days at 28 °C. All experiments were repeated at least three times.
BiFC and firefly luciferase LCI assays
BiFC and LCI assays were conducted as described previously60. For the BiFC assay, the full-length cDNA sequences of NAC2, CMV1a, CMV1aG983D, PVYCI, PVYCIG347D and TMV126KD were cloned into the binary cYFP or nYFP vector by ligation-independent cloning technique. Primer pairs for making these constructs were made by Beijing Tsingke Biotech and are listed in Supplementary Table 3. These gene sequences were PCR-amplified using M5 Magic High-Fidelity DNA Polymerase (MF740, Mei5Bio) from plant cDNA or virus vector. The Mut Express II Fast Mutagenesis Kit V2 (C214-01, Vazyme) was used to construct mutants. PCR products were purified using the HiPure Gel Pure DNA Mini Kit (D2111-02, Magen). The YFP fluorescence signal for each combination was detected by an inverted confocal microscope (Leica SP8) at 48 h after infiltration. Confocal images were analysed using Leica LAS X (3.3). Anti-GFP (ab290, Abcam) or anti-rabbit (HA1001, HUABIO) antibodies were used at a 1:3,000 dilution for immunoblot analysis. For LCI assays, the full-length cDNA sequences of CMV1a, CMV1a-H, CMV1a-M and NAC2 were cloned into the nLUC and cLUC vectors by ligation-independent cloning assay. The luciferase substrate (luciferin) was smeared onto the surface of the leaves and the luciferase activity was captured using the PlantView100 assay system (BLT Photon Technology). All of the experiments were independently repeated at least three times.
Co-IP assay
The constructs were transiently co-expressed with the indicated combinations in N. benthamiana leaves. At 48 h after infiltration, the leaves were taken for protein extraction. Protein extracts were immunoprecipitated using RFP-Trap beads (rtma, ChromoTek) in co-IP buffer (50 mM Tris-HCl, pH 7.5, 150 mM NaCl, 10% glycerol, 0.5 mM EDTA, 0.5% NP-40, 5 mM DTT, 1 mM PMSF and protease inhibitor cocktail (Roche)). The mixture was incubated for 3 h at 4 °C. The immunoprecipitates were then denatured by 2× protein loading buffer containing β-mercaptoethanol (JS0150, Hong Kong JSENB International Trading) and separated on a 10% SDS–PAGE gel (M00664, GenScript) followed by immunoblot analysis. Anti-MYC (M20002M, Abmart), anti-mouse (AS003, ABclonal), anti-RFP (5f8-100, ChromoTek) or anti-rat (AS028, ABclonal) antibodies were used at a 1:3,000 dilution for immunoblot analysis. The StarSignal Chemiluminescent Assay Kit (E171-01, GenStar) was used for signal detection. All of the experiments were repeated three times and generated similar results.
RNA-seq analysis
RNAs extracted from 5-week-old WT and nac2 plants (without any treatment) were used for RNA-seq analysis. To examine the expression profiles of the WT and nac2 plants in response to aphid attack, RNAs extracted from 3-week-old WT and nac2 plants (2 days after aphid infestation) were used for RNA-seq analysis. RNA-seq was performed by Biomarker Technologies (www.biomarker.com.cn). Raw data were processed using the Illumina NovaSeq 6000 sequencer (2 × 150 bp read length) and aligned to N. benthamiana reference genome (https://solgenomics.net/). Differentially expressed genes were identified according to the transcripts per million reads (TPM) (log2[fold change] ≥ 1 or ≤ −1 and P ≤ 0.05). Furthermore, Gene Ontology (GO, http://www.geneontology.org) and Kyoto Encyclopedia of Genes and Genomes (KEGG, http://www.genome.jp/kegg/) analyses were performed to identify differentially expressed genes that were significantly enriched in GO terms and KEGG pathways.
EMSA
GST and GST–NAC2 proteins were purified by Glutathione Sepharose 4B (GE Healthcare) affinity chromatography and eluted by 10 mM GSH followed by size-exclusion chromatography (Superdex 200 10/300, GE Healthcare). Oligonucleotide probes were labelled with the biotin by RUIBIO Technology. Double-distilled H2O was purified using the Milli Q (Merck) water-purification system. Chemiluminescent EMSA Kit (GS009, Beyotime) was used for EMSA. The detailed protocol was described previously21. EMSA experiments were repeated at least three times.
RT–qPCR
Total RNA was extracted using TransZol reagent (TransGen Biotech) and retained in a 1.5 ml sterile centrifugal tube (HRK-150-TP, Beijing Huaruikang Technology). Then, 5 μg of RNA was reverse-transcribed into cDNA using the TransScript One-Step gDNA Removal and cDNA Synthesis SuperMix (AT311-02, TransGen Biotech). RT–qPCR was performed using the Hieff qPCR SYBR Green Master Mix (11201ES08, YEASEN). Data were normalized to ACTIN expression using the cycle threshold (CT) \({2}^{-\Delta \Delta {C}_{{\rm{t}}}}\) method as described previously61 and analysed using Prism9 (GraphPad). All of the experiments were repeated at least three times. Primers used for RT–qPCR are listed in Supplementary Table 3. Data are represented as mean ± s.d.
Virus infection and analysis of viral RNA and protein
For testing the effect of NAC2 on virus infection, nac2 and WT N. benthamiana plants were grown in soil for 3–4 weeks and infected with CMV62, PVY-GFP63 or TMV-GFP through agro-infiltration. Viral symptoms were photographed at 6 days after inoculation for CMV and TMV-GFP, and 7 days after inoculation for PVY-GFP. For performing virus infection in silenced plants, 20-day-old WT or samt1 plants were first agro-infiltrated with TRV-NAC2 or TRV, and the top leaves were then infected with CMV or PVY-GFP at 14 days after treatment with virus-induced gene silencing. Viral symptoms were photographically recorded at 7 days after inoculation for CMV and PVY-GFP. Anti-CMV (CMV21-A, Alpha Diagnostic International), anti-PVY (POTY11-A, Alpha Diagnostic International), anti-TMV (TMV11-A, Alpha Diagnostic International), anti-rabbit (HA1001, HUABIO), or anti-goat (BE0103, EASYBIO) antibodies were used at a 1:3,000 dilution for immunoblot analysis.
For testing whether and how MeSA is responsible for NAC2-mediated plant antiviral defence, nac2, samt1, sabp2 and WT N. benthamiana plants were grown in soil for 3–4 weeks, and plants were sprayed with control or solutions of 500 µM SA or MeSA daily for 3 days before infection with CMV or PVY–GFP as described previously64,65. Systemic leaves were taken for viral infection analysis at 6 or 7 days after inoculation.
ChIP–qPCR analysis
ChIP–qPCR was performed according to the published protocol with minor modifications66. N. benthamiana leaves over-expressing HA-NAC2 or HA-nLUC were collected and chromatin that was isolated from 2 g of frozen leaf tissue was sonicated using a Bioruptor sonicator for 6 min. The SimpleChIP Plus Kit (9005 S, CST) was used to perform the experiment. Enrichment of promoter DNA was measured using the percentage input method by RT–qPCR analysis as described previously67. Amplification of the ACTIN2 promoter sequence was used as the negative control. Primers for the ChIP–qPCR assays are listed in Supplementary Table 3. ChIP–qPCR experiments were repeated three times, all showing similar results.
Semi in vivo and in vivo protein degradation
For chemical treatments, MG132 (C2211, Sigma-Aldrich), cycloheximide (CHX) (HY-12320, MedChemExpress) and ATP (IA1310, Solarbio) were used. MG132 and CHX were dissolved in DMSO and used at a concentration of 100 μM. ATP was used at the concentration of 10 mM.
For semi in vivo protein degradation analysis, N. benthamiana leaves overexpressing CMV1a-MYC, CMV1a(G983D)–MYC, RFP–NAC2 or cLUC–MYC were collected at 2 days after infiltration. The proteins were extracted using extraction buffer (50 mM Tris-MES pH 8.0, 0.5 M sucrose, 1 mM MgCl2, 10 mM EDTA, 5 mM DTT, protease inhibitor cocktail) as described previously68. For analysis of NAC2 degradation by the 26S proteasome, the plant extract containing RFP–NAC2 was mixed with chemicals containing CHX, then the extract was split equally into two tubes, into one tube was added into ATP, and into the other tube was added an equal volume of extraction buffer. The two tubes were incubated in the Eppendorf Thermomixer at 25 °C, and extracts were removed from the two tubes as samples at different timepoints. For analysis of CMV1a-mediated promotion of NAC2 protein degradation through the ubiquitin-proteasome pathway, the RFP–NAC2 extract was mixed with CMV1a–MYC, CMV1a(G983D)–MYC or cLUC–MYC extracts at a ratio of 1:1 before incubation. ATP and CHX together with MG132 or DMSO were added to the prepared leaf extracts. The samples were collected at different timepoints.
For in vivo analysis of NAC2 or NES–NAC2 protein degradation by the 26S proteasome, RFP–NAC2 or RFP–NES–NAC2 were transiently expressed in N. benthamiana leaves for 60 h. The agro-inoculated leaves were then treated with MG132 or an equal volume of DMSO control solution for 12 h before sampling. For in vivo analysis of NAC2 protein degradation promoted by CMV1a, RFP–NAC2 was co-expressed with CMV1a–MYC, CMV1a(G983D)–MYC or cLUC–MYC for 60 h before MG132 treatment for 12 h.
SA–MeSA competition binding assay
Analysis of [3H]SA binding to SABP2 was performed as described previously with modifications20,69. The GST–SABP2 proteins were expressed in Escherichia coli and purified. The size-exclusion columns were prepared by adding 0.13 g of Sephadex G-25 (GE healthcare) to Qiagen shredder columns. The columns were pre-equilibrated with reaction buffer (50 mM Tris-HCl, 150 mM NaCl, pH 7.5) containing 0.1% Tween-20 overnight at 4 °C, and excess buffer was removed by centrifuging at 735g for 2 min. The GST–SABP2 proteins were incubated with 10 nM, 20 nM, 50 nM and 100 nM [3H]SA (50 Ci mmol−1) in reaction buffer on ice for 1 h, and then loaded onto the columns and centrifuged immediately as described above. The flow-through was collected and the radioactivity was measured using 2 ml Ultima Gold AB Cocktails (PerkinElmer) with a scintillation counter (MicroBeta2; PerkinElmer). For SA–MeSA competition binding experiments, the GST–SABP2 proteins were incubated with 50 nM [3H]SA in the presence of 0 nM, 3 nM or 15 nM MeSA on ice for 1 h. The binding activity of [3H]SA (50 Ci mmol−1) by SABP2 in the absence of competitor MeSA was set to 100%, and the binding activity of [3H]SA by SABP2 with 3 nM or 15 nM MeSA was calculated along with the comparison. GraphPad Prism 9.0 was used to construct the nonlinear binding model of the Michaelis–Menten equation.
Alignment of the helicase domain
Sequences of viral helicase-domain-containing proteins were retrieved from UniProt (https://www.uniprot.org/). Multiple-sequence alignments of proteins were performed in Jalview using Mafft (L-INS-i method) (http://www.jalview.org/About).
Statistical analysis
Statistical significance was determined using two-sided Student’s t-tests or one-way ANOVA with Tukey’s multiple-comparison test for multiple groups (≥3) of data. For aphid attraction bioassays, data were statistically analysed using the χ2 tests. For analysing the aphid survival portion in two groups, data were statistically analysed using one-way ANOVA with the LSD test. Statistical analysis was performed using GraphPad Prism 9.0 or IBM SPSS Statistics 26. Detailed statistical analyses are explained in the figure legends, and P values are indicated in the figures and Source Data. The chromatography (total ion chromatogram) data of each sample were exported using The Xcalibur v.4.1. Graphics were drawn using GraphPad Prism v.9.0. For quantification analysis of blots, the intensities of bands were quantified using ImageJ.
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data and materials needed to replicate the work are available. NAC2.1 (Niben101Scf01481g02006), NAC2.2 (Niben101Scf07152g04032), SAMT1 (Niben101Scf05122g00005) and SABP2 (Niben101Scf00034g00012) are available from the N. benthamiana genome database (https://solgenomics.net/). RNA-seq raw data have been deposited at the National Center for Biotechnology Information (NCBI) under BioProject accessions PRJNA851626 (WT), PRJNA851854 (nac2), PRJNA955195 (WT-aphid) and PRJNA955395 (nac2-aphid). Uncropped gel and immunoblotting images are provided in Supplementary Fig. 1. Source data are provided with this paper.
References
Van Emden, H. F. & Harrington, R. Aphids as Crop Pests 2nd Edn (CABI, 2017).
Karban, R. Plant communication. Ann. Rev. Ecol. Evol. Syst. 52, 1–24 (2021).
Loreto, F. & D’Auria, S. How do plants sense volatiles sent by other plants? Trends Plant Sci. 27, 29–38 (2022).
Shulaev, V., Silverman, P. & Raskin, I. Airborne signalling by methyl salicylate in plant pathogen resistance. Nature 386, 738–738 (1997).
Wenig, M. et al. Systemic acquired resistance networks amplify airborne defense cues. Nat. Commun. 10, 3813 (2019).
Pickett, J. A. & Khan, Z. R. Plant volatile-mediated signalling and its application in agriculture: successes and challenges. N. Phytol. 212, 856–870 (2016).
Sugimoto, K. et al. Identification of a tomato UDP-arabinosyltransferase for airborne volatile reception. Nat. Commun. 14, 677 (2023).
Bleecker, A. B. & Schaller, G. E. The mechanism of ethylene perception. Plant Physiol. 111, 653–660 (1996).
Fereres, A. & Moreno, A. Behavioural aspects influencing plant virus transmission by homopteran insects. Virus Res. 141, 158–168 (2009).
Babikova, Z. et al. Underground signals carried through common mycelial networks warn neighbouring plants of aphid attack. Ecol. Lett. 16, 835–843 (2013).
Moreira, X., Nell, C. S., Katsanis, A., Rasmann, S. & Mooney, K. A. Herbivore specificity and the chemical basis of plant-plant communication in Baccharis salicifolia (Asteraceae). N. Phytol. 220, 703–713 (2018).
Staudt, M. et al. Volatile organic compound emissions induced by the aphid Myzus persicae differ among resistant and susceptible peach cultivars and a wild relative. Tree Physiol. 30, 1320–1334 (2010).
Saad, K. A., Mohamad Roff, M. N., Hallett, R. H. & Idris, A. B. Aphid-induced defences in chilli affect preferences of the whitefly, Bemisia tabaci (Hemiptera: Aleyrodidae). Sci. Rep. 5, 13697 (2015).
Dong, Y. J. & Hwang, S. Y. Cucumber Plants baited with methyl salicylate accelerates Scymnus (Pullus) sodalis (Coleoptera: Coccinellidae) visiting to reduce cotton aphid (Hemiptera: Aphididae) infestation. J. Econ. Entomol. 110, 2092–2099 (2017).
Mallinger, R. E., Hogg, D. B. & Gratton, C. Methyl salicylate attracts natural enemies and reduces populations of soybean aphids (Hemiptera: Aphididae) in soybean agroecosystems. J. Econ. Entomol. 104, 115–124 (2011).
Ninkovic, V., Glinwood, R., Unlu, A. G., Ganji, S. & Unelius, C. R. Effects of methyl salicylate on host plant acceptance and feeding by the aphid Rhopalosiphum padi. Front. Plant Sci. 12, 710268 (2021).
Park, S. W. Methyl salicylate is a critical mobile signal for plant systemic acquired resistance. Science 321, 342–342 (2008).
Vlot, A. C., Dempsey, D. A. & Klessig, D. F. Salicylic acid, a multifaceted hormone to combat disease. Ann. Rev. Phytopathol. 47, 177–206 (2009).
Dudareva, N., Raguso, R. A., Wang, J. H., Ross, E. J. & Pichersky, E. Floral scent production in Clarkia breweri—III. Enzymatic synthesis and emission of benzenoid esters. Plant Physiol. 116, 599–604 (1998).
Forouhar, F. et al. Structural and biochemical studies identify tobacco SABP2 as a methyl salicylate esterase and implicate it in plant innate immunity. Proc. Natl Acad. Sci. USA 102, 1773–1778 (2005).
Wang, Y. J. et al. A calmodulin-binding transcription factor links calcium signaling to antiviral RNAi defense in plants. Cell Host Microbe 29, 1393–1406 (2021).
Olsen, A. N., Ernst, H. A., Lo Leggio, L. & Skriver, K. NAC transcription factors: structurally distinct, functionally diverse. Trends Plant Sci. 10, 79–87 (2005).
De Clercq, I. et al. The membrane-bound NAC transcription factor ANAC013 functions in mitochondrial retrograde regulation of the oxidative stress response in Arabidopsis. Plant Cell 25, 3472–3490 (2013).
Zhu, F. et al. Salicylic acid and jasmonic acid are essential for systemic resistance against tobacco mosaic virus in Nicotiana benthamiana. Mol. Plant Microbe Interact. 27, 567–577 (2014).
Naylor, M., Murphy, A. M., Berry, J. O. & Carr, J. P. Salicylic acid can induce resistance to plant virus movement. Mol. Plant Microbe Interact. 11, 860–868 (1998).
Guo, H. J. et al. Aphid-borne viral spread is enhanced by virus-induced accumulation of plant reactive oxygen species. Plant Physiol. 179, 143–155 (2019).
Evans, R. et al. Protein complex prediction with AlphaFold-Multimer. Preprint at bioRxiv https://doi.org/10.1101/2021.10.04.463034 (2021).
Arimura, G. et al. Herbivory-induced volatiles elicit defence genes in lima bean leaves. Nature 406, 512–515 (2000).
Engelberth, J., Alborn, H. T., Schmelz, E. A. & Tumlinson, J. H. Airborne signals prime plants against insect herbivore attack. Proc. Natl Acad. Sci. USA 101, 1781–1785 (2004).
Karban, R., Yang, L. H. & Edwards, K. F. Volatile communication between plants that affects herbivory: a meta-analysis. Ecol. Lett. 17, 44–52 (2014).
Donovan, M. P., Nabity, P. D. & DeLucia, E. H. Salicylic acid-mediated reductions in yield in Nicotiana attenuata challenged by aphid herbivory. Arthropod Plant Interact. 7, 45–52 (2013).
Cao, H. H., Liu, H. R., Zhang, Z. F. & Liu, T. X. The green peach aphid Myzus persicae perform better on pre-infested Chinese cabbage Brassica pekinensis by enhancing host plant nutritional quality. Sci. Rep. 6, 21954 (2016).
Blande, J. D., Korjus, M. & Holopainen, J. K. Foliar methyl salicylate emissions indicate prolonged aphid infestation on silver birch and black alder. Tree Physiol. 30, 404–416 (2010).
van Poecke, R. M. P. & Dicke, M. Indirect defence of plants against herbivores: using Arabidopsis thaliana as a model plant. Plant Biol. 6, 387–401 (2004).
James, D. G. Field evaluation of herbivore-induced plant volatiles as attractants for beneficial insects: methyl salicylate and the green lacewing, Chrysopa nigricornis. J. Chem. Ecol. 29, 1601–1609 (2003).
Woods, J. L., James, D. G., Lee, J. C. & Gent, D. H. Evaluation of airborne methyl salicylate for improved conservation biological control of two-spotted spider mite and hop aphid in Oregon hop yards. Exp. Appl. Acarol. 55, 401–416 (2011).
Rowen, E., Gutensohn, M., Dudareva, N. & Kaplan, I. Carnivore attractant or plant elicitor? Multifunctional roles of methyl salicylate lures in tomato defense. J. Chem. Ecol. 43, 573–585 (2017).
Liu, J. et al. Herbivore-Induced rice volatiles attract and affect the predation ability of the wolf spiders, Pirata subpiraticus and Pardosa pseudoannulata. Insects 13, 90 (2022).
Attaran, E., Zeier, T. E., Griebel, T. & Zeier, J. Methyl salicylate production and jasmonate signaling are not essential for systemic acquired resistance in Arabidopsis. Plant Cell 21, 954–971 (2009).
Kwon, S. et al. Biotic and abiotic stresses induce AbSAMT1, encoding S-adenosyl-l-methionine: salicylic acid carboxyl methyltransferase, in Atropa belladonna. Plant Biotechnol. 26, 207–215 (2009).
Xu, R., Song, F. & Zheng, Z. OsBISAMT1, a gene encoding S-adenosyl-l-methionine: salicylic acid carboxyl methyltransferase, is differentially expressed in rice defense responses. Mol. Biol. Rep. 33, 223–231 (2006).
Hippauf, F. et al. Enzymatic, expression and structural divergences among carboxyl O-methyltransferases after gene duplication and speciation in Nicotiana. Plant Mol. Biol. 72, 311–330 (2010).
Li, R. et al. Virulence factors of geminivirus interact with MYC2 to subvert plant resistance and promote vector performance. Plant Cell 26, 4991–5008 (2014).
Zhao, P. Z. et al. Viruses mobilize plant immunity to deter nonvector insect herbivores. Sci. Adv. 5, eaav9801 (2019).
Tungadi, T. et al. Cucumber mosaic virus and its 2b protein alter emission of host volatile organic compounds but not aphid vector settling in tobacco. Virol. J. 14, 91 (2017).
Wu, D. W. et al. Viral effector protein manipulates host hormone signaling to attract insect vectors. Cell Res. 27, 402–415 (2017).
Westwood, J. H. et al. A trio of viral proteins tunes aphid-plant interactions in Arabidopsis thaliana. PLoS ONE 8, e83066 (2013).
Rhee, S. J., Watt, L. G., Bravo, A. C., Murphy, A. M. & Carr, J. P. Effects of the cucumber mosaic virus 2a protein on aphid-plant interactions in Arabidopsis thaliana. Mol. Plant Pathol. 21, 1248–1254 (2020).
Casteel, C. L. et al. The NIa-Pro protein of turnip mosaic virus improves growth and reproduction of the aphid vector, Myzus persicae (green peach aphid). Plant J. 77, 653–663 (2014).
Bak, A., Cheung, A. L., Yang, C. L., Whitham, S. A. & Casteel, C. L. A viral protease relocalizes in the presence of the vector to promote vector performance. Nat. Commun. 8, 14493 (2017).
Ying, X. B. et al. RNA-dependent RNA polymerase 1 from Nicotiana tabacum suppresses RNA silencing and enhances viral infection in Nicotiana benthamiana. Plant Cell 22, 1358–1372 (2010).
Ma, X. et al. A robust CRISPR/Cas9 system for convenient, high-efficiency multiplex genome editing in monocot and dicot plants. Mol. Plant 8, 1274–1284 (2015).
Kessler, A. & Baldwin, I. T. Defensive function of herbivore-induced plant volatile emissions in nature. Science 291, 2141–2144 (2001).
Kost, C. & Heil, M. Herbivore-induced plant volatiles induce an indirect defence in neighbouring plants. J. Ecol. 94, 619–628 (2006).
Ozawa, R., Arimura, G., Takabayashi, J., Shimoda, T. & Nishioka, T. Involvement of jasmonate- and salicylate-related signaling pathways for the production of specific herbivore-induced volatiles in plants. Plant Cell Physiol. 41, 391–398 (2000).
Feng, H. et al. Acylsugars protect Nicotiana benthamiana against insect herbivory and desiccation. Plant Mol. Biol. 109, 505–522 (2021).
Fernandez-Calvino, L., Lopez-Abella, D., Lopez-Moya, J. J. & Fereres, A. Comparison of potato virus Y and plum pox virus transmission by two aphid species in relation to their probing behavior. Phytoparasitica 34, 315–324 (2006).
Verdier, M., Chesnais, Q., Pirolles, E., Blanc, S. & Drucker, M. The cauliflower mosaic virus transmission helper protein P2 modifies directly the probing behavior of the aphid vector Myzus persicae to facilitate transmission. PLoS Pathog. 19, e1011161 (2023).
Xu, H. X. et al. A salivary effector enables whitefly to feed on host plants by eliciting salicylic acid-signaling pathway. Proc. Natl Acad. Sci. USA 116, 490–495 (2019).
Wang, Y. et al. Geminiviral V2 protein suppresses transcriptional gene silencing through interaction with AGO4. J. Virol. 93, e01675-18 (2019).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the \({2}^{-\Delta \Delta {C}_{{\rm{T}}}}\) method. Methods 25, 402–408 (2001).
Wang, X. B. et al. The 21-nucleotide, but not 22-nucleotide, viral secondary small interfering RNAs direct potent antiviral defense by two cooperative argonautes in Arabidopsis thaliana. Plant Cell 23, 1625–1638 (2011).
Geng, C. et al. Tobacco vein banding mosaic virus 6K2 protein hijacks NbPsbO1 for virus replication. Sci. Rep. 7, 43455 (2017).
Mayers, C. N., Lee, K. C., Moore, C. A., Wong, S. M. & Carr, J. P. Salicylic acid-induced resistance to cucumber mosaic virus in squash and Arabidopsis thaliana: contrasting mechanisms of induction and antiviral action. Mol. Plant Microbe Interact. 18, 428–434 (2005).
Bi, H. H., Zeng, R. S., Su, L. M., An, M. & Luo, S. M. Rice allelopathy induced by methyl jasmonate and methyl salicylate. J. Chem. Ecol. 33, 1089–1103 (2007).
Saleh, A., Alvarez-Venegas, R. & Avramova, Z. An efficient chromatin immunoprecipitation (ChIP) protocol for studying histone modifications in Arabidopsis plants. Nat. Protoc. 3, 1018–1025 (2008).
Yamaguchi, N. et al. PROTOCOLS: chromatin immunoprecipitation from Arabidopsis tissues. Arabidopsis Book 12, e0170 (2014).
Liu, L. et al. An efficient system to detect protein ubiquitination by agroinfiltration in Nicotiana benthamiana. Plant J. 61, 893–903 (2010).
Slaymaker, D. H. et al. The tobacco salicylic acid-binding protein 3 (SABP3) is the chloroplast carbonic anhydrase, which exhibits antioxidant activity and plays a role in the hypersensitive defense response. Proc. Natl Acad. Sci. USA 99, 11640–11645 (2002).
Acknowledgements
We thank L. Kang for his suggestions; X. Li and X. Wang for providing the PVY-GFP and CMV (CMV-Fny and CMVΔ2b, the latter is a CMV 2b gene deletion mutant) infectious clones, respectively; H. Guo for providing the seeds of NahG N. benthamiana; P. Zhao and J. Ye for assistance in measurement of SA amount; D. Li for her assistance in the radiolabelling experiments; and the staff at the Facility Center of Metabolomics and Lipidomics of China National Center for Protein Sciences for GC–MS analysis. This work was supported in part by the National Key R&D Program of China (2022YFD1400800 and 2021YFD1400400), the National Natural Science Foundation of China (32130086, 31920103013 and 31872636), the Young Elite Scientists Sponsorship Program by CAST (2022QNRC001) and the Shuimu Tsinghua Scholar Program.
Author information
Authors and Affiliations
Contributions
Y.L. conceived the research. Q.G., Yunjing Wang and Y.L. designed all of the experiments. Q.G. and Yunjing Wang performed all of the aphid choice bioassays, SA–MeSA competition binding experiments, semi in vivo and in vivo protein degradation experiments, ChIP–qPCR, EMSA, GC–MS analysis experiments, virus infection and analysis of viral RNA and protein experiments, and aphid survival as well as virus-transmission analysis. Yunjing Wang, L.H. and F.H. performed BiFC, yeast-one-hybrid and LCI experiments. Yunjing Wang, D.Z., Yan Wang and X.W. generated transgenic plants. Q.G., L.L. and F.C. performed EPG analysis. Yunjing Wang, M.H. and H.D. performed MS analysis. Q.G., Yunjing Wang, Y.H. and Y.L. analysed data and wrote the paper with input from all of the authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature thanks Richard Karban, Jürgen Zeier and the other, anonymous, reviewers for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Fig. 1 NAC2 interacts with CMV1a and is required for plant antiviral defence and MeSA mediated aphid repellence.
a, Representative LC-MS/MS spectrum of peptides in NAC2 protein. A peptide (AGIAQDAFVLCR) is shown. b-d, Confirmation of the CMV1a-NAC2 interaction. In Co-immunoprecipitation (Co-IP) assay (b), cLUC-MYC or CMV1a-MYC was co-expressed with RFP-NAC2 in Nb leaves and analysed 2 days post-infiltration (dpi). Both Bimolecular fluorescence complementation (BiFC) (c) and firefly luciferase complementation imaging (LCI) assays (d) further confirmed CMV1a-NAC2 interaction in Nb leaves. Scale bar = 25 μm. e-g, Viral symptoms in WT and nac2 plants infected with CMV at 6 dpi (e), and the relative accumulation of CMV RNA (f) or CP (g) in CMV-infected systemic leaves of WT or nac2 plants. h-j, Viral symptoms in WT and nac2 plants infected with PVY-GFP at 7 dpi (h), and the accumulation of PVY RNA (i) or CP (j) in PVY-GFP infected systemic leaves of WT or nac2 plants. k-m, Viral symptoms in WT and nac2 plants infected with TMV-GFP at 6 dpi (k), and the relative accumulation of TMV RNA (l) or CP (m) in TMV-GFP infected systemic leaves of WT or nac2 plants. n, o, nac2 plants exhibited higher attractiveness to aphids than WT plants in circular-dish (n) or Y-tube olfactometer bioassays (o). Numerals shown inside each bar present number of choice-making aphids. p, q, nac2 plants smeared with MeSA containing lanolin exhibited similar attractiveness to aphids with WT plants under same treatment in circular-dish bioassay (p) or Y-tube olfactometer bioassays (q). r, s, nac2 plants smeared with lanolin alone or with 3,3-dimethyl-hexane containing lanolin exhibited higher attractiveness to aphids than WT plants under same treatment in circular-dish (r) or Y-tube olfactometer bioassays (s). t, u, Volatile MeSA treatment caused WT plants more repellent to aphids in circular-dish bioassay (t) or Y-tube olfactometer bioassays (u). v, w, No significant difference in aphid repellence between nac2 plants with and without volatile MeSA treatment in circular-dish (v) or Y-tube olfactometer (w) bioassays. x, y, After volatile MeSA treatment followed by ventilation, nac2 plants showed higher attractiveness to aphids than WT plants in circular-dish (x) or Y-tube olfactometer (y) bioassays. f, i, l, Two-sided Student’s t-test, n = 3 biologically independent samples. Data are shown as mean ± s.d. n-y, χ2 test (d.f. = 1). All P values are shown in figure. Experiments were repeated at least three times with similar results.
Extended Data Fig. 2 GC-MS analysis of VOCs emitted from WT and nac2 plants.
a-d, Direct sequencing of PCR product containing targeted sites in CRISPR/Cas9-edited knockout nac2, samt1, and sabp2 homozygous plants. The rectangular area indicates the start positions at or from which the mutations occurred. It is worth mentioning that there is only one SAMT1 or SABP2 copy in N.benthamiana genome although Nb is allotetraploid. e, GC-MS analysis of VOCs emitted from WT and nac2 plants after 48 h aphid feeding. Wet weight per plant was 1.10 g on average. Identifications based on retention indices and GC-MS: (1) oxalic acid, allyl hexyl ester; (2) bicyclo[3.1.0]hex-2-ene, 2-methyl-5-(1-methylethyl)-; (3) benzene, 1,2,3-trimethyl-; (4) 6-methyl heptanoate; (5) butyl pyruvate; (6) benzene, 1,2,3,5-tetramethyl-; (7) methyl salicylate; (8) tridecane, 4-methyl-; (9) 3,3-dimethyl-hexane; (10) Cyclohexasiloxane, dodecamethyl-; (11) bicyclo[3.1.1]heptane, 6,6-dimethyl-3-methylene-; (12) heptane, 2,2,3,3,5,6,6-heptamethyl-; (13) heptane, 2,3,6-trimethyl-; (14) 2,2,4-Trimethyl-1,3-pentanediol diisobutyrate; (15) phenol, 2,6-bis(1,1-dimethylethyl)−4-(1-methylpropyl)-. f, The mean amounts of volatile MeSA collected from the aphid-attacked plants in GC-MS analysis. g, h, Mass spectrum of putative MeSA produced by plants (g) and the authentic MeSA standard (h). i, j, Comparison of the aerial MeSA concentrations in chambers containing either MeSA dissolved lanolin paste or aphid attacked WT plants by GC-MS analysis. Experiments were repeated at least three times with similar results.
Extended Data Fig. 3 NAC2 binds to SAMT1 promoter to activate its transcription.
a, Emitter and receiver plants were placed on two trays at a distance of 30 cm from each other in an interplant communication assay set-up in open-air environment. Each emitter was fed with fifty virus-free or viruliferous M. persicae aphids or no aphids before the emitters and receivers were incubated in same cage (100 cm × 70 cm × 70 cm) made by gauze. After 3 days, the receiver plants were taken out for further experiments. b, NbNAC2 and AtNAC2 share similar conserved motifs. c, RFP-tagged NAC2 showed nuclear localization, scale bar = 20 μm. d, e, RT-qPCR quantification of relative SAMT1 mRNA levels in nac2 and WT plants (d) or leaves transiently over-expressing HA-NAC2 and HA-nLUC (e). f, Transient expression assays. NAC2 activates luciferase reporter gene transcription under the control of the SAMT1 promoter (SAMT1pro) in Nb leaves. Photograph was taken at 48 h post-infiltration (hpi). g, In planta ChIP-qPCR. HA-NAC2, but not HA-nLUC, specifically binds to the SAMT1 promoter DNA. Chromatin from plants expressing HA-nLUC or HA-NAC2 were immunoprecipitated and amplified with promoter-specific primers. h, Yeast one-hybrid assay. Yeast cells were co-transformed with an effector vector containing the SAMT1pro cloned into pHis2 vector and a prey vector encoding NAC2 cloned into pGADT7. i, In vitro EMSA. Hot probe is the biotin-labelled NAC2-binding motif DNA of SAMT1pro, cold probe or cold mutant probe is the unlabelled NAC2-binding motif DNA of SAMT1pro or its mutant DNA. j, Transient over-expression of NAC2 increases MeSA production in plants, the samples were collected at 48 hpi. d, e, g, j, Two-sided Student’s t-test, n = 3 biologically independent samples. Data are shown as mean ± s.d.; n.s., no statistical significance. All P values are shown in figure. Experiments were repeated at least three times with similar results.
Extended Data Fig. 4 RNA-seq transcriptome analysis of WT and nac2 plants.
a, Comparison of WT and nac2 plants (without any treatment) RNA-seq sequences on the reference genome. The Phred quality score Q20 (99% base call accuracy) and Q30 (99.9% base call accuracy) were used to measure the quality of RNA sequencing. b, Hierarchical clustered heat map of 90 differential expressed genes (DEGs, 24-up genes and 66-down genes) based on the log2 (fold change) in transcript levels of WT and nac2 plants. c, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway analysis of 90 DEGs in (b), the module with the most enriched genes is “metabolism” (green colour, marked by red font). d, Gene ontology (GO) analysis of 90 DEGs in (b), the “biological process” module (green colour) with the most enriched genes is “metabolism process” (marked by red font). e, Phylogenetic analysis of SAMT1 proteins from various plant species. NtSAMT1, Nicotiana tabacum SAMT1 (GenBank ID: FJ015052); SlSAMT1, Solanum lycopersicum SAMT1 (GenBank ID: NM_001247880); AtBSMT1, Arabidopsis thaliana BSMT1 (Tair ID: AT3G11480); AtGAMT1 (Tair ID: AT4G26420); AtIAMT1 (Tair ID: AT5G55250). f, Comparison of WT (aphid) and nac2 (aphid) plants (under aphid attack) RNA-seq sequences on the reference genome. g, Hierarchical clustered heat map of 157 DEGs (100-up genes and 57-down genes) based on the log2(fold change) in transcript levels of WT (aphid) and nac2 (aphid) plants. h, KEGG pathway analysis of 157 DEGs in (g), the module with the most enriched genes is “metabolism” (green colour, marked by red font). i, GO analysis of 157 DEGs in (g), the “biological process” module (green colour) with the most enriched genes is “metabolism process” (marked by red font). All DEGs were identified according to the transcripts per million reads (TPM) (log2 (foldchange) ≥ 1 or ≤ −1 and P ≤ 0.05).
Extended Data Fig. 5 NAC2 regulates SAMT1 to fulfill anti-CMV and PVY function.
a-e, Analysis of antiviral role of NAC2 and SAMT1 during CMV infection in nac2, samt1, or nac2/samt1 double mutants. Viral symptoms (a), relative NAC2 mRNA levels in NAC2-silenced WT (b) or NAC2-silenced samt1 plants (c), the accumulation of CMV RNA (d) or CP protein (e), and plant endogenous MeSA amount (f, g) of WT, nac2, samt1, or nac2/samt1 double mutant plants infected with CMV. h-l, Analysis of antiviral role of NAC2 and SAMT1 during PVY-GFP infection in nac2, samt1, or nac2/samt1 double mutants. Viral symptom (h), relative NAC2 mRNA levels in NAC2-silenced WT (i) or NAC2-silenced samt1 plants (j), PVY RNA accumulation (k) or PVY CP (l) of nac2, samt1, or nac2/samt1 double mutant plants infected with PVY-GFP. Plants in panels (a and h) were photographed at 7 days post-inoculation (dpi). NAC2-knockdown (KD) was triggered by the virus-induced gene silencing (VIGS) vector TRV-NAC2 to mimic nac2 mutant while TRV was used as negative control in these experiments. b-d, f, g, i-k, Two-sided Student’s t-test, n = 3 biologically independent samples. Data are shown as mean ± s.d. All P values are shown in figure. Experiments were repeated at least three times with similar results.
Extended Data Fig. 6 CMV1a suppresses MeSA-mediated AD, and CMV1aG983D impairs its interaction with NAC2.
a, Transgenic CMV1a-MYC or CMV1aG983D-MYC expression Nb plant showed normal growth and development. b, CMV1a-MYC and CMV1aG983D-MYC were detected in transgenic plants by western blot. Rubisco was used as loading control. c, d, Transgenic CMV1a plants exhibited higher attractiveness to aphids than WT plants in circular-dish (c) or Y-tube olfactometer (d) bioassays. e, EPG analysis showed that the number of pd of individual aphids was more in CMV1a plants than WT plants. n = 19 individual aphids. f-h, Accumulation of CMV RNA (f), CP (g) or proportion of living aphids (h) in CMV-carrying aphid-attacked leaves of WT receivers (WT-R) with non-aphid-attacked WT plants as mock emitters (WT-mE), virus-free aphid-attacked WT plants as emitters (WT-AE), non-aphid-attacked transgenic CMV1a plants as mock emitters (1a-mE), or virus-free aphid-attacked transgenic CMV1a plants as emitters (1a-AE). i, j, WT-R (WT-mE) plants exhibited higher attractiveness than WT-R (WT-AE) plants (i), but WT-R plants showed similar attractiveness to aphids when non-aphid-attacked or virus-free aphid-attacked CMV1a plants were used as emitters (1a-mE or 1a-AE) (j) in Y-tube olfactometer bioassays. k, WT-R plants nearby virus-free aphid-attacked CMV1a plants as emitters (1a-AE) exhibited higher attractiveness to WT-R plants close to virus-free aphid-attacked WT plants as emitters (WT-AE). l, LCI assay to show that CMV1a helicase domain (1a-H), but not methyltransferase domain (1a-M), interacts with NAC2. m, AlphaFold-Multimer predicted CMV1a-H-NAC2 interacting complex (parameters: “MMseqs2” and “AlphaFold2-Multimer-v2” pattern). Colours are given based on AlphaFold-Multimer-calculated prediction score: pLDDT. Protein structures with scores over 90 are represented in blue (very high confidence of prediction); scores between 70 and 90 in light blue (high confidence); scores between 50 and 70 in yellow (low), and anything below 50 in orange (very low confidence of prediction). n, Red and blue indicate the CMV1a-H and NAC2, respectively. The stick model represents the potential interacting site between 1a-H and NAC2, this region is predicted with a high confidence score. o, Co-IP assay to show that G983D mutation in CMV1a helicase domain impairs the 1a helicase domain-NAC2 interaction. p, q, Co-IP assay (p) and BiFC assay (q) to show that CMV1aG983D failed to interact with NAC2 in Nb. Scale bar = 25 μm. e, f, Two-sided Student’s t-test. h, One-way ANOVA with Tukey’s multiple comparisons test; letters A-B represent statistically different groups (P < 0.05). f, h, n = 3 biologically independent samples. Data are shown as mean ± s.d.; n.s., no statistical significance. c, d, i-k, χ2 test (d.f. = 1). P values are shown in c-f, i-k; adjusted P values for h are shown in the Source Data. In box plot (e), the centre line represents the median, box edges delimit bottom and top quartiles and whiskers show the highest and lowest data points. Experiments were repeated at least three times with similar results.
Extended Data Fig. 7 CMV1a re-localizes and degrades NAC2 by 26S-proteasome system, and CMV1aG983D possesses a weakened aphid repellence.
a, CMV1a but not CMV1aG983D partially changed NAC2 localization from nucleus to cytoplasm. b, CMV1a did not alter subcellular localization of RFP. c, d, Nuclear exit signal-tagged RFP-NAC2 (NES-NAC2) changed NAC2 localization to cytoplasm (c) and enhanced 26S-proteasome system-dependent degradation (d). Scale bar = 25 μm in panels (a-c). e, Immunoblot assay of RFP protein levels. f, Immunoblots to show cLUC-MYC, CMV1a-MYC, and CMV1aG983D-MYC protein levels with anti-MYC antibody. g, In vivo assay showing effects of MG132 and the CMV1a-NAC2 interaction on NAC2 protein stability. 100 μM MG132 or an equal volume of DMSO (negative control) was infiltrated into leaves transiently co-expressing RFP-NAC2 with CMV1a-MYC, CMV1aG983D-MYC, or cLUC-MYC for 12 h before harvesting. h, Semi-in vivo assay to show that NAC2 protein stability is ATP-dependent. NAC2 protein levels were analysed with anti-RFP antibody at different times following 100 μM CHX treatment in the presence or absence of 10 mM ATP. i, Semi-in vivo assay to show that MG132 inhibits CMV1a-promoted NAC2 degradation. RFP-NAC2, cLUC-MYC, CMV1a-MYC, or CMV1aG983D-MYC was transiently expressed in Nb leaves and extracted respectively. NAC2 degradation was performed as below: The RFP-NAC2 protein extract was mixed with the cLUC-MYC, CMV1a-MYC, or CMV1aG983D-MYC extracts in a 1:1 volume of 100 μM CHX and 10 mM ATP, in the presence of 100 μM MG132 or an equal volume of control DMSO. j, Effect of CMV1a on expression of luciferase reporter gene driven by the SAMT1 promoter (SAMT1pro). Transient expression assays in Nb leaves to show that CMV1a but not CMV1aG983D suppressed NAC2-mediated activation of the SAMT1 promoter. Photographs were taken at 48 hpi. k, Transgenic plants expressing CMV1a, but not CMV1aG983D, exhibited higher attractiveness to aphids than WT plants in Y-tube olfactometer bioassays. l, m, GC-MS analysis to show that transgenic plants expressing CMV1a, but not CMV1aG983D, emitted less volatized MeSA than WT plants once they were fed with virus-free aphids for 3 days. n, WT-R (WT-mE) plants exhibited higher attractiveness than WT-R (WT-AE) plants, WT-R plants showed similar attractiveness to aphids when non-aphid-attacked or virus-free aphid-attacked CMV1a-expressinig plants were used as emitters (1a-mE or 1a-AE), whilst WT-R (1aG983D-mE) plants exhibited higher attractiveness than WT-R (1aG983D-AE) plants in Y-tube olfactometer bioassays. m, Two-sided Student’s t-test, n = 3 biologically independent samples. Data are shown as mean ± s.d.; n.s., no statistical significance. k, n, χ2 test (d.f. = 1). All P values are shown in figure. Experiments were repeated at least three times with similar results.
Extended Data Fig. 8 Alignment of helicase domain from aphid and non-aphid transmitted viruses.
BCMNV, Bean common mosaic necrosis virus (Uniprot ID: Q65399); BYMV, Bean yellow mosaic virus (Uniprot ID: P17765); MDMV, Maize dwarf mosaic virus (Uniprot ID: J7II85); PPV, Plum pox potyvirus (Uniprot ID: P13529); PRSV, Papaya ringspot virus (Uniprot ID: A0A1L2DBW1); PVY, Potato virus Y (Uniprot ID: A0A5J6BDG4); PeMV, Pepper mottle virus (Uniprot ID: Q01500); PVMV, Pepper veinal mottle virus (Uniprot ID: A0A6J4A295); SCMV, Sugarcane mosaic virus (Uniprot ID: A0A0K0Y0R3); SMV, Soybean mosaic virus (Uniprot ID: Q90069); TEV, Tobacco etch virus (Uniprot ID: P04517); TuMV, Turnip mosaic virus (Uniprot ID: Q9ICI2); TVBMV, Tobacco vein banding mosaic virus (Uniprot ID: F5A3N8); ZYMV, Zucchini yellow mosaic virus (Uniprot ID: P18479); CLV, Carnation latent virus (Uniprot ID: A0A858Z687); CMV, Cucumber mosaic virus (Uniprot ID: P17769); PSV, Peanut stunt virus (Uniprot ID: P28726); TAV, Tomato aspermy virus (Uniprot ID: P28931); BYDV, Barley yellow dwarf virus (Uniprot ID: P29044); SbDV, Soybean dwarf virus (Uniprot ID: A0A6M8PRM6); AMV, Alfalfa mosaic virus (Uniprot ID: P03589); PEBV, Pea early browning virus (Uniprot ID: Q9WJD8); TRV, Tobacco rattle virus (Uniprot ID: Q9J942); PVX, Potato virus X (Uniprot ID: A0A7H1C8Y4); TMV, Tobacco mosaic virus (Uniprot ID: P03586); TYMV, Turnip yellow mosaic virus (Uniprot ID: P10358).
Extended Data Fig. 9 PVY suppresses plant AD by CI-NAC2 interaction.
a, Alignment of helicase domain from multiple aphid- and non-aphid- transmitted viruses. Asterisk (*) indicates that the Glycine (G) amino acid residue is conserved among listed aphid-transmitted viruses. b, c, Accumulation of PVY RNA (b) or CP (c) in PVY-carrying aphid-attacked leaves of WT receivers (WT-R) with virus-free aphid-attacked WT plants as emitters (AE) or with non-aphid-attacked WT plants as mock emitters (mE) when these receiver plants were fed with PVY-containing aphids. d, e, GC–MS analysis of volatized MeSA in WT plants fed with virus-free aphids or PVY-carrying aphids for 3 days. f-h, Accumulation of PVY RNA (f), CP (g) or proportion of living aphids (h) in WT-R plants with PVY-containing aphid-attacked WT plants as emitters (APE) or WT-R (mE) plants when these receiver plants were fed with PVY-containing aphids. i, CMV and PVY infection changed NAC2 localization partially to cytoplasm. Scale bar = 25 μm. j, BiFC assay showing that PVY CI but not its mutant PVY CIG347D or TMV 126KD, interacted with NAC2. Scale bar = 25 μm. k, Immunoblot assay of protein levels in BiFC assay. l, PVY CI but not its mutant PVY CIG347D or TMV 126KD, changed NAC2 localization partially to cytoplasm, scale bar = 25 μm. b, e, f, Two-sided Student’s t-test. h, One-way ANOVA with LSD. b, e, f, h, n = 3 biologically independent samples. Data are shown as mean ± s.d.; n.s., no statistical significance. The same letter A represents no statistical difference between samples (P > 0.05). All P values are shown in figure. Experiments were repeated at least three times with similar results.
Extended Data Fig. 10 Arms race among emitter and receiver plants, aphids, and viruses.
a, AD defends plants against aphids and viruses. When emitter plants are attacked by aphids, they can sense the aphid sap-sucking action and stimulate biosynthesis of SA that activates the NAC2-SAMT1 module to produce volatile MeSA, neighbouring receiver plants perceive and convert volatile MeSA into SA by SABP2, which acts as the cue to trigger NAC2-SAMT1 module and elicit defence against aphids and viruses. b, Virus and aphid counterdefence. When emitter plants were attacked by viruliferous aphids, some aphid-transmitted viruses utilized their helicase-contained viral protein (for example, CMV1a and PVY CI) to subcellularly re-localize and destabilize NAC2, leading to suppression of NAC2-mediated plant airborne defence to facilitate aphid propagation and virus transmission. The graphical model was created with BioRender.com. In summary, we have exploited interplays among aphid, virus, VOC emitter, and receiver plants in a complexed pathosystem to dissect PPC and AD at genetic and molecular levels. Our study on deciphering AD also lays the groundbreaking work to empower VOCs as a novel bioinspired tool in defence of plants including agricultural and horticultural crops against insect infestation and virus epidemics.
Supplementary information
Supplementary Figure 1
All uncropped blots and gel images.
Supplementary Table 1
Differentially expressed genes in WT and nac2 plants.
Supplementary Table 2
Differentially expressed genes in WT (aphid) and nac2 (aphid) plants.
Supplementary Table 3
Primers used in this study.
Source data
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gong, Q., Wang, Y., He, L. et al. Molecular basis of methyl-salicylate-mediated plant airborne defence. Nature 622, 139–148 (2023). https://doi.org/10.1038/s41586-023-06533-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41586-023-06533-3
This article is cited by
-
The evolutionary trajectories of specialized metabolites towards antiviral defense system in plants
Molecular Horticulture (2024)
-
Interplant communication: an emerging battlefield in plant-aphid-virus interactions
Science China Life Sciences (2024)
-
Lipid peroxidation and stress-induced signalling molecules in systemic resistance mediated by azelaic acid/AZELAIC ACID INDUCED1: signal initiation and propagation
Physiology and Molecular Biology of Plants (2024)
-
A tug-of-war to control plant emission of an airborne alarm signal
Stress Biology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.