Featured
-
-
Article
| Open Access Identification of domains in Plasmodium falciparum proteins of unknown function using DALI search on AlphaFold predictions
Identification of domains in Plasmodium falciparum proteins of unknown function using DALI search on AlphaFold predictions
- Hannah Michaela Behrens
- & Tobias Spielmann
-
Article
| Open Access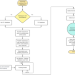 DeepReg: a deep learning hybrid model for predicting transcription factors in eukaryotic and prokaryotic genomes
DeepReg: a deep learning hybrid model for predicting transcription factors in eukaryotic and prokaryotic genomes
- Leonardo Ledesma-Dominguez
- , Erik Carbajal-Degante
- & Ernesto Perez-Rueda
-
Article
| Open Access CCDC58 is a potential biomarker for diagnosis, prognosis, immunity, and genomic heterogeneity in pan-cancer
CCDC58 is a potential biomarker for diagnosis, prognosis, immunity, and genomic heterogeneity in pan-cancer
- Kai Yang
- , Yan Ma
- & Ruohan Zhang
-
Article
| Open Access Characterization of a putative orexin receptor in Ciona intestinalis sheds light on the evolution of the orexin/hypocretin system in chordates
Characterization of a putative orexin receptor in Ciona intestinalis sheds light on the evolution of the orexin/hypocretin system in chordates
- Maiju K. Rinne
- , Lauri Urvas
- & Henri Xhaard
-
Article
| Open Access VirtuousPocketome: a computational tool for screening protein–ligand complexes to identify similar binding sites
VirtuousPocketome: a computational tool for screening protein–ligand complexes to identify similar binding sites
- Lorenzo Pallante
- , Marco Cannariato
- & Marco A. Deriu
-
Article
| Open Access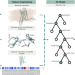 Characterizing conformational states in GPCR structures using machine learning
Characterizing conformational states in GPCR structures using machine learning
- Ilya Buyanov
- & Petr Popov
-
Article
| Open Access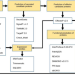 A repertoire of candidate effector proteins of the fungus Ceratocystis cacaofunesta
A repertoire of candidate effector proteins of the fungus Ceratocystis cacaofunesta
- Gabriela N. Ramos-Lizardo
- , Jonathan J. Mucherino-Muñoz
- & Ronan Xavier Corrêa
-
Article
| Open Access AiKPro: deep learning model for kinome-wide bioactivity profiling using structure-based sequence alignments and molecular 3D conformer ensemble descriptors
AiKPro: deep learning model for kinome-wide bioactivity profiling using structure-based sequence alignments and molecular 3D conformer ensemble descriptors
- Hyejin Park
- , Sujeong Hong
- & Jae-Min Shin
-
Article
| Open Access Rare variant contribution to cholestatic liver disease in a South Asian population in the United Kingdom
Rare variant contribution to cholestatic liver disease in a South Asian population in the United Kingdom
- Julia Zöllner
- , Sarah Finer
- & Peter H. Dixon
-
Article
| Open Access Coenzyme A binding sites induce proximal acylation across protein families
Coenzyme A binding sites induce proximal acylation across protein families
- Chris Carrico
- , Andrew Cruz
- & Eric Verdin
-
Article
| Open Access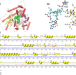 Accurate prediction by AlphaFold2 for ligand binding in a reductive dehalogenase and implications for PFAS (per- and polyfluoroalkyl substance) biodegradation
Accurate prediction by AlphaFold2 for ligand binding in a reductive dehalogenase and implications for PFAS (per- and polyfluoroalkyl substance) biodegradation
- Hao-Bo Guo
- , Vanessa A. Varaljay
- & Rajiv Berry
-
Article
| Open Access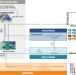 Optimizing variant-specific therapeutic SARS-CoV-2 decoys using deep-learning-guided molecular dynamics simulations
Optimizing variant-specific therapeutic SARS-CoV-2 decoys using deep-learning-guided molecular dynamics simulations
- Katharina Köchl
- , Tobias Schopper
- & Christian C. Gruber
-
Article
| Open Access Pan-kinome of Legionella expanded by a bioinformatics survey
Pan-kinome of Legionella expanded by a bioinformatics survey
- Marianna Krysińska
- , Bartosz Baranowski
- & Marcin Gradowski
-
Article
| Open Access Structural informatic study of determined and AlphaFold2 predicted molecular structures of 13 human solute carrier transporters and their water-soluble QTY variants
Structural informatic study of determined and AlphaFold2 predicted molecular structures of 13 human solute carrier transporters and their water-soluble QTY variants
- Eva Smorodina
- , Igor Diankin
- & Shuguang Zhang
-
Article
| Open Access The effect of mutations on binding interactions between the SARS-CoV-2 receptor binding domain and neutralizing antibodies B38 and CB6
The effect of mutations on binding interactions between the SARS-CoV-2 receptor binding domain and neutralizing antibodies B38 and CB6
- Jonathan E. Barnes
- , Peik K. Lund-Andersen
- & F. Marty Ytreberg
-
Article
| Open Access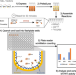 Parallel functional annotation of cancer-associated missense mutations in histone methyltransferases
Parallel functional annotation of cancer-associated missense mutations in histone methyltransferases
- Ashley J. Canning
- , Susan Viggiano
- & Michael S. Cosgrove
-
Article
| Open Access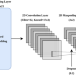 Improving protein succinylation sites prediction using embeddings from protein language model
Improving protein succinylation sites prediction using embeddings from protein language model
- Suresh Pokharel
- , Pawel Pratyush
- & Dukka B. KC
-
Article
| Open Access De novo metatranscriptomic exploration of gene function in the millipede holobiont
De novo metatranscriptomic exploration of gene function in the millipede holobiont
- Puspendu Sardar
- , Vladimír Šustr
- & Lucie Faktorová
-
Article
| Open Access ProteinGLUE multi-task benchmark suite for self-supervised protein modeling
ProteinGLUE multi-task benchmark suite for self-supervised protein modeling
- Henriette Capel
- , Robin Weiler
- & K. Anton Feenstra
-
Article
| Open Access Identification and expression of functionally conserved circadian clock genes in lichen-forming fungi
Identification and expression of functionally conserved circadian clock genes in lichen-forming fungi
- Henrique F. Valim
- , Francesco Dal Grande
- & Imke Schmitt
-
Article
| Open Access Exploring the medicinally important secondary metabolites landscape through the lens of transcriptome data in fenugreek (Trigonella foenum graecum L.)
Exploring the medicinally important secondary metabolites landscape through the lens of transcriptome data in fenugreek (Trigonella foenum graecum L.)
- Mahantesha B. N. Naika
- , Nitish Sathyanarayanan
- & Ramanathan Sowdhamini
-
Article
| Open Access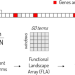 Gene function prediction in five model eukaryotes exclusively based on gene relative location through machine learning
Gene function prediction in five model eukaryotes exclusively based on gene relative location through machine learning
- Flavio Pazos Obregón
- , Diego Silvera
- & Rafael Cantera
-
Article
| Open Access AlphaFold2 models indicate that protein sequence determines both structure and dynamics
AlphaFold2 models indicate that protein sequence determines both structure and dynamics
- Hao-Bo Guo
- , Alexander Perminov
- & Rajiv Berry
-
Article
| Open Access Multi-task learning to leverage partially annotated data for PPI interface prediction
Multi-task learning to leverage partially annotated data for PPI interface prediction
- Henriette Capel
- , K. Anton Feenstra
- & Sanne Abeln
-
Article
| Open Access A novel method for assessing and measuring homophily in networks through second-order statistics
A novel method for assessing and measuring homophily in networks through second-order statistics
- Nicola Apollonio
- , Paolo G. Franciosa
- & Daniele Santoni
-
Article
| Open Access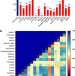 Dynamic insights into the effects of nonsynonymous polymorphisms (nsSNPs) on loss of TREM2 function
Dynamic insights into the effects of nonsynonymous polymorphisms (nsSNPs) on loss of TREM2 function
- Raju Dash
- , Yeasmin Akter Munni
- & Il Soo Moon
-
Article
| Open Access Non-covalent Fc-Fab interactions significantly alter internal dynamics of an IgG1 antibody
Non-covalent Fc-Fab interactions significantly alter internal dynamics of an IgG1 antibody
- Ramakrishnan Natesan
- & Neeraj J. Agrawal
-
Article
| Open Access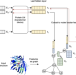 LM-GVP: an extensible sequence and structure informed deep learning framework for protein property prediction
LM-GVP: an extensible sequence and structure informed deep learning framework for protein property prediction
- Zichen Wang
- , Steven A. Combs
- & Peter M. Clark
-
Article
| Open Access LIM domain-wide comprehensive virtual mutagenesis provides structural rationale for cardiomyopathy mutations in CSRP3
LIM domain-wide comprehensive virtual mutagenesis provides structural rationale for cardiomyopathy mutations in CSRP3
- Pankaj Kumar Chauhan
- & Ramanathan Sowdhamini
-
Article
| Open Access Interactions of Co, Cu, and non-metal phthalocyanines with external structures of SARS-CoV-2 using docking and molecular dynamics
Interactions of Co, Cu, and non-metal phthalocyanines with external structures of SARS-CoV-2 using docking and molecular dynamics
- Wilson Luna Machado Alencar
- , Tiago da Silva Arouche
- & Antonio Maia de Jesus Chaves Neto
-
Article
| Open Access A deep learning model to detect novel pore-forming proteins
A deep learning model to detect novel pore-forming proteins
- Theju Jacob
- & Theodore W. Kahn
-
Article
| Open Access Extracting phylogenetic dimensions of coevolution reveals hidden functional signals
Extracting phylogenetic dimensions of coevolution reveals hidden functional signals
- Alexandre Colavin
- , Esha Atolia
- & Kerwyn Casey Huang
-
Article
| Open Access In-silico phenotype prediction by normal mode variant analysis in TUBB4A-related disease
In-silico phenotype prediction by normal mode variant analysis in TUBB4A-related disease
- Avi Fellner
- , Yael Goldberg
- & Felix Benninger
-
Article
| Open Access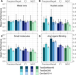 Protein embeddings and deep learning predict binding residues for various ligand classes
Protein embeddings and deep learning predict binding residues for various ligand classes
- Maria Littmann
- , Michael Heinzinger
- & Burkhard Rost
-
Article
| Open Access ACP-MHCNN: an accurate multi-headed deep-convolutional neural network to predict anticancer peptides
ACP-MHCNN: an accurate multi-headed deep-convolutional neural network to predict anticancer peptides
- Sajid Ahmed
- , Rafsanjani Muhammod
- & Abdollah Dehzangi
-
Article
| Open Access Apoplastic effector candidates of a foliar forest pathogen trigger cell death in host and non-host plants
Apoplastic effector candidates of a foliar forest pathogen trigger cell death in host and non-host plants
- Lukas Hunziker
- , Mariana Tarallo
- & Rosie E. Bradshaw
-
Article
| Open Access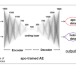 Autoencoder-based detection of the residues involved in G protein-coupled receptor signaling
Autoencoder-based detection of the residues involved in G protein-coupled receptor signaling
- Yuko Tsuchiya
- , Kei Taneishi
- & Yasushige Yonezawa
-
Article
| Open Access Theoretical and experimental study of interaction of macroheterocyclic compounds with ORF3a of SARS-CoV-2
Theoretical and experimental study of interaction of macroheterocyclic compounds with ORF3a of SARS-CoV-2
- Natalia Sh. Lebedeva
- , Yury A. Gubarev
- & Oskar I. Koifman
-
Article
| Open Access Computational identification of multiple lysine PTM sites by analyzing the instance hardness and feature importance
Computational identification of multiple lysine PTM sites by analyzing the instance hardness and feature importance
- Sabit Ahmed
- , Afrida Rahman
- & S. M. Shovan
-
Article
| Open Access Insights into the substrate binding mechanism of SULT1A1 through molecular dynamics with excited normal modes simulations
Insights into the substrate binding mechanism of SULT1A1 through molecular dynamics with excited normal modes simulations
- Balint Dudas
- , Daniel Toth
- & Maria A. Miteva
-
Article
| Open Access A deep learning based approach for prediction of Chlamydomonas reinhardtii phosphorylation sites
A deep learning based approach for prediction of Chlamydomonas reinhardtii phosphorylation sites
- Niraj Thapa
- , Meenal Chaudhari
- & Dukka B. KC
-
Article
| Open Access Critical assessment of coiled-coil predictions based on protein structure data
Critical assessment of coiled-coil predictions based on protein structure data
- Dominic Simm
- , Klas Hatje
- & Martin Kollmar
-
Article
| Open Access Predicting MHC I restricted T cell epitopes in mice with NAP-CNB, a novel online tool
Predicting MHC I restricted T cell epitopes in mice with NAP-CNB, a novel online tool
- Carlos Wert-Carvajal
- , Rubén Sánchez-García
- & Arrate Muñoz-Barrutia
-
Article
| Open Access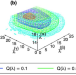 Difference of binding modes among three ligands to a receptor mSin3B corresponding to their inhibitory activities
Difference of binding modes among three ligands to a receptor mSin3B corresponding to their inhibitory activities
- Tomonori Hayami
- , Narutoshi Kamiya
- & Junichi Higo
-
Article
| Open Access An in silico deep learning approach to multi-epitope vaccine design: a SARS-CoV-2 case study
An in silico deep learning approach to multi-epitope vaccine design: a SARS-CoV-2 case study
- Zikun Yang
- , Paul Bogdan
- & Shahin Nazarian
-
Article
| Open Access Improved prediction and characterization of anticancer activities of peptides using a novel flexible scoring card method
Improved prediction and characterization of anticancer activities of peptides using a novel flexible scoring card method
- Phasit Charoenkwan
- , Wararat Chiangjong
- & Watshara Shoombuatong
-
Article
| Open Access Comparative transcriptomics and host-specific parasite gene expression profiles inform on drivers of proliferative kidney disease
Comparative transcriptomics and host-specific parasite gene expression profiles inform on drivers of proliferative kidney disease
- Marc Faber
- , Sophie Shaw
- & Jason W. Holland
-
Article
| Open Access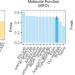 Embeddings from deep learning transfer GO annotations beyond homology
Embeddings from deep learning transfer GO annotations beyond homology
- Maria Littmann
- , Michael Heinzinger
- & Burkhard Rost
-
Article
| Open Access In silico analysis of epitope-based vaccine candidate against tuberculosis using reverse vaccinology
In silico analysis of epitope-based vaccine candidate against tuberculosis using reverse vaccinology
- Shaheen Bibi
- , Inayat Ullah
- & Shiquan Niu

 Structure-based epitope prediction and assessment of cross-reactivity of Myrmecia pilosula venom-specific IgE and recombinant Sol g proteins (Solenopsis geminata)
Structure-based epitope prediction and assessment of cross-reactivity of Myrmecia pilosula venom-specific IgE and recombinant Sol g proteins (Solenopsis geminata)