Featured
-
-
Article
| Open Access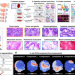 Spatially resolved multi-omics highlights cell-specific metabolic remodeling and interactions in gastric cancer
Spatially resolved multi-omics highlights cell-specific metabolic remodeling and interactions in gastric cancer
The spatial signature of metabolic remodeling in tumours remains to be explored. Here, the integration of mass spectrometry imaging-based spatial metabolomics and lipidomics with microarray-based spatial transcriptomics allows the visualisation of metabolic heterogeneity in gastric cancer.
- Chenglong Sun
- , Anqiang Wang
- & Jiuming He
-
Article
| Open Access Designing reliable and accurate isotope-tracer experiments for CO2 photoreduction
Designing reliable and accurate isotope-tracer experiments for CO2 photoreduction
Current verification of CO2 photoreduction products using isotope-tracer methods is insufficient and can provide false-positive results. Here, rigorous protocols and case studies are presented to avoid pitfalls in isotope-tracer experiments.
- Shengyao Wang
- , Bo Jiang
- & Jinhua Ye
-
Article
| Open Access Single-cell metabolic fingerprints discover a cluster of circulating tumor cells with distinct metastatic potential
Single-cell metabolic fingerprints discover a cluster of circulating tumor cells with distinct metastatic potential
Circulating tumor cells (CTCs) are the tumor cells responsible for metastasis. Here, the authors develop a home-built single-cell quantitative mass spectrometric platform and discover a cluster of CTCs with distinct metastatic potential using a molecular typing system based on the single-cell metabolic fingerprints.
- Wenjun Zhang
- , Feifei Xu
- & Yun Chen
-
Article
| Open Access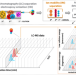 PeakDecoder enables machine learning-based metabolite annotation and accurate profiling in multidimensional mass spectrometry measurements
PeakDecoder enables machine learning-based metabolite annotation and accurate profiling in multidimensional mass spectrometry measurements
Alternative algorithms exploiting advantages of multidimensional mass spectrometry in untargeted metabolomics are needed. Here, the authors develop and demonstrate PeakDecoder for confident and accurate metabolite profiling in 116 microbial sample runs and using a library built from 64 standards.
- Aivett Bilbao
- , Nathalie Munoz
- & Kristin E. Burnum-Johnson
-
Article
| Open Access PepQuery2 democratizes public MS proteomics data for rapid peptide searching
PepQuery2 democratizes public MS proteomics data for rapid peptide searching
Billions of MS/MS spectra are available in public proteomics data repositories, but their usage has been limited to informatics experts. Here, the authors provide a solution to democratize these data for rapid peptide searching and demonstrate utilities in a wide range of biological applications
- Bo Wen
- & Bing Zhang
-
Article
| Open Access Workflow enabling deepscale immunopeptidome, proteome, ubiquitylome, phosphoproteome, and acetylome analyses of sample-limited tissues
Workflow enabling deepscale immunopeptidome, proteome, ubiquitylome, phosphoproteome, and acetylome analyses of sample-limited tissues
Patient samples are often available in limited amounts, restricting the number of possible omics analyses. Here the authors present MONTE, a workflow that enables serial HLA-I and HLA-II immunopeptidome, ubiquitylome, proteome, phosphoproteome, and acetylome data collection from patient samples.
- Jennifer G. Abelin
- , Erik J. Bergstrom
- & Steven A. Carr
-
Article
| Open Access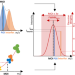 Spatial probabilistic mapping of metabolite ensembles in mass spectrometry imaging
Spatial probabilistic mapping of metabolite ensembles in mass spectrometry imaging
Spatial visualization of metabolites in tissues via mass spectrometry imaging can be prone to user perception bias. Here, the authors report the computational framework moleculaR that introduces probabilistic data-dependent molecular mapping of nonrandom spatial patterns of metabolite signals.
- Denis Abu Sammour
- , James L. Cairns
- & Carsten Hopf
-
Article
| Open Access A mass spectrum-oriented computational method for ion mobility-resolved untargeted metabolomics
A mass spectrum-oriented computational method for ion mobility-resolved untargeted metabolomics
The high dimensionality of ion mobility (IM)-resolved metabolomics data presents a great challenge to data processing. Here, authors develop a mass spectrum-oriented bottom-up assembly algorithm and the end-to-end computational framework Met4DX for IM-resolved metabolomics.
- Mingdu Luo
- , Yandong Yin
- & Zheng-Jiang Zhu
-
Article
| Open Access Neutron-encoded diubiquitins to profile linkage selectivity of deubiquitinating enzymes
Neutron-encoded diubiquitins to profile linkage selectivity of deubiquitinating enzymes
Most insights into deubiquitinase (DUB) substrate specificity originate from studies with isolated di-ubiquitins (diUb), but in cells diUbs with different linkage types coexist. Here, the authors develop a mass spectrometric DUB activity assay that can probe all diUbs simultaneously under substrate competition conditions.
- Bianca D. M. van Tol
- , Bjorn R. van Doodewaerd
- & Paul P. Geurink
-
Article
| Open Access Highly-sensitive label-free deep profiling of N-glycans released from biomedically-relevant samples
Highly-sensitive label-free deep profiling of N-glycans released from biomedically-relevant samples
Glycans can serve as disease biomarkers. Here, the authors use label-free capillary electrophoresis-mass spectrometry for N-glycan profiling of minute sample amounts, resolving and characterizing previously undetected highly sialylated glycans and linkage isomers in a single analysis.
- Anne-Lise Marie
- , Somak Ray
- & Alexander R. Ivanov
-
Article
| Open Access High-resolution separation of bioisomers using ion cloud profiling
High-resolution separation of bioisomers using ion cloud profiling
Ion mobility is used in mass spectrometers for structure analysis of biomolecules. Here, the authors show that ion mobility analysis in an ion trap under ultra-high fields enables isomer separation at resolutions over 10,000, wich they demonstrate for isomers of disaccharides, phospholipids, and peptides.
- Xiaoyu Zhou
- , Zhuofan Wang
- & Zheng Ouyang
-
Article
| Open Access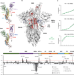 Structural dynamics in the evolution of SARS-CoV-2 spike glycoprotein
Structural dynamics in the evolution of SARS-CoV-2 spike glycoprotein
In this paper, the authors use hydrogen-deuterium exchange mass spectrometry to describe how the SARS-CoV-2 spike glycoprotein has evolved its structural dynamics features and receptor binding capability from the emergence of the original Wuhan isolate to the recent omicron variant. The findings reported shed light on the evolution of SARS-CoV-2 in the human population and the mechanisms of emergence of new variants.
- Valeria Calvaresi
- , Antoni G. Wrobel
- & Argyris Politis
-
Article
| Open Access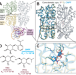 Ligand-specific changes in conformational flexibility mediate long-range allostery in the lac repressor
Ligand-specific changes in conformational flexibility mediate long-range allostery in the lac repressor
Using hydrogen-deuterium exchange, the authors propose a model explaining how a classic transcription factor undergoes changes in its conformational ensemble in response to different ligands.
- Anum Glasgow
- , Helen T. Hobbs
- & Tanja Kortemme
-
Article
| Open Access Structural mechanism for inhibition of PP2A-B56α and oncogenicity by CIP2A
Structural mechanism for inhibition of PP2A-B56α and oncogenicity by CIP2A
Tumour suppressors are inhibited in cancers and their reactivation could provide novel therapy opportunities. Here, the authors study the structural mechanism by which human tumour suppressor Protein Phosphatase 2A is inhibited in breast cancer cells by the oncoprotein CIP2A.
- Karolina Pavic
- , Nikhil Gupta
- & Jukka Westermarck
-
Article
| Open Access Four-dimensional trapped ion mobility spectrometry lipidomics for high throughput clinical profiling of human blood samples
Four-dimensional trapped ion mobility spectrometry lipidomics for high throughput clinical profiling of human blood samples
The circulatory lipidome is a valuable source for disease markers, but reliable marker discovery requires continuous development of lipidomic methods for large-scale clinical profiling. Here, the authors present a 4-dimensional lipidomics solution for confident and reproducible blood lipidome profiling.
- Raissa Lerner
- , Dhanwin Baker
- & Laura Bindila
-
Article
| Open Access Sample multiplexing-based targeted pathway proteomics with real-time analytics reveals the impact of genetic variation on protein expression
Sample multiplexing-based targeted pathway proteomics with real-time analytics reveals the impact of genetic variation on protein expression
Targeted proteomics enables robust hypothesis-driven research. Here, Yu et al. present a multiplexed approach for targeted pathway proteomics and apply it to quantify protein families across 480 fully genotyped Diversity Outbred mice, revealing impacts of genetic variation on protein expression and lipid metabolism.
- Qing Yu
- , Xinyue Liu
- & Steven P. Gygi
-
Article
| Open Access Structures of honeybee-infecting Lake Sinai virus reveal domain functions and capsid assembly with dynamic motions
Structures of honeybee-infecting Lake Sinai virus reveal domain functions and capsid assembly with dynamic motions
Viruses can cause Colony Collapse Disorder, leading to large losses of honeybee hives globally. In this study, the authors solve capsid structures of honeybee-infecting Lake Sinai viruses and identify distinct features, which advances understanding of viral dynamics, assembly and infection mechanisms.
- Nai-Chi Chen
- , Chun-Hsiung Wang
- & Chun-Jung Chen
-
Article
| Open Access Annotation of natural product compound families using molecular networking topology and structural similarity fingerprinting
Annotation of natural product compound families using molecular networking topology and structural similarity fingerprinting
Comparing experimental mass spectra to reference spectra can enable natural product identification, but these spectral libraries are often incomplete and not universally applicable. Here, the authors present SNAP-MS, a tool that allows assigning compound families without experimental or calculated reference spectra.
- Nicholas J. Morehouse
- , Trevor N. Clark
- & Roger G. Linington
-
Article
| Open Access Gas induced formation of inactive Li in rechargeable lithium metal batteries
Gas induced formation of inactive Li in rechargeable lithium metal batteries
The formation of electrochemically inactive, or “dead”, lithium limits the reversibility of lithium metal batteries. Here the authors elucidate the (electro)chemical roles of ethylene gas produced from electrolyte decomposition on the formation of inactive lithium.
- Yuxuan Xiang
- , Mingming Tao
- & Yong Yang
-
Article
| Open Access Oncogenic mutations of PIK3CA lead to increased membrane recruitment driven by reorientation of the ABD, p85 and C-terminus
Oncogenic mutations of PIK3CA lead to increased membrane recruitment driven by reorientation of the ABD, p85 and C-terminus
The PIK3CA gene is one of the most frequently mutated genes in all of human cancer. Here authors describe the molecular mechanisms underlying how different oncogenic mutations in PIK3CA mediate increased activity.
- Meredith L. Jenkins
- , Harish Ranga-Prasad
- & John E. Burke
-
Article
| Open Access Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Benchmarking commonly used software suites and analysis workflows for DIA proteomics and phosphoproteomics
Many software suites and spectral libraries have been developed for DIA proteomics data analysis. Here, the authors create benchmark data sets to evaluate four commonly used software tools combined with seven spectral libraries in both global proteomics and phosphoproteomics analysis.
- Ronghui Lou
- , Ye Cao
- & Wenqing Shui
-
Article
| Open Access Protein complex prediction using Rosetta, AlphaFold, and mass spectrometry covalent labeling
Protein complex prediction using Rosetta, AlphaFold, and mass spectrometry covalent labeling
Covalent labeling (CL) from mass spectrometry experiments provides structural information of higher-order protein structure. Here, the authors develop an algorithm which integrates experimental CL data to predict protein complexes in the Rosetta molecular modeling suite using AlphaFold models.
- Zachary C. Drake
- , Justin T. Seffernick
- & Steffen Lindert
-
Article
| Open Access Serum metabolic traits reveal therapeutic toxicities and responses of neoadjuvant chemoradiotherapy in patients with rectal cancer
Serum metabolic traits reveal therapeutic toxicities and responses of neoadjuvant chemoradiotherapy in patients with rectal cancer
Neoadjuvant chemoradiotherapy is the standard treatment for patients with locally advanced rectal cancer, however, response can be limited by development of toxicities. Here, the authors conducted metabolomics on patients with locally advanced rectal cancer enrolled in a phase III clinical study and identify serum metabolites associated with treatment response.
- Hongmiao Wang
- , Huixun Jia
- & Zheng-Jiang Zhu
-
Article
| Open Access pGlycoQuant with a deep residual network for quantitative glycoproteomics at intact glycopeptide level
pGlycoQuant with a deep residual network for quantitative glycoproteomics at intact glycopeptide level
Software tools for larger-scale intact glycopeptide quantification lag far behind, which hinders exploring the differential sitespecific glycosylation. Here, the authors report pGlycoQuant, a generic tool with a deep learning model for quantitative glycoproteomics at intact glycopeptide level.
- Siyuan Kong
- , Pengyun Gong
- & Weiqian Cao
-
Article
| Open Access Protein-Peptide Turnover Profiling reveals the order of PTM addition and removal during protein maturation
Protein-Peptide Turnover Profiling reveals the order of PTM addition and removal during protein maturation
Metabolic labeling is often used to measure protein turnover. Here the authors show that for interconvertible protein species like phosphoforms metabolic labeling does not provide information on turnover differences, but that the relative order of modification can determine the observed dynamics.
- Henrik M. Hammarén
- , Eva-Maria Geissen
- & Mikhail M. Savitski
-
Article
| Open Access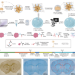 Spatially resolved proteomics via tissue expansion
Spatially resolved proteomics via tissue expansion
Spatially resolved proteomics is an emerging approach for mapping proteome heterogeneity. Here, the authors report a method based on the combination of hydrogel-based tissue transformation with mass spectrometry-based proteomics, that enables proteome profiling with a lateral resolution of 160 µm.
- Lu Li
- , Cuiji Sun
- & Kiryl D. Piatkevich
-
Article
| Open Access Structural basis for lipid and copper regulation of the ABC transporter MsbA
Structural basis for lipid and copper regulation of the ABC transporter MsbA
The bacterial ABC transporter MsbA is essential for lipopolysaccharide biogenesis. Here, the authors apply native mass spectrometry, X-ray crystallography, cryo-EM and biochemical approaches to characterize the structural basis and functional roles of MsbA binding to copper and specific lipids.
- Jixing Lyu
- , Chang Liu
- & Arthur Laganowsky
-
Article
| Open Access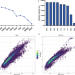 Insights into protein post-translational modification landscapes of individual human cells by trapped ion mobility time-of-flight mass spectrometry
Insights into protein post-translational modification landscapes of individual human cells by trapped ion mobility time-of-flight mass spectrometry
Single-cell proteomics is an emerging approach to study cellular heterogeneity but its coverage is still limited. Here, the authors develop a single-cell proteomics approach with improved protein sequence coverage, allowing them to quantify PTMs and characterize effects of inhibitor treatment in single human cells.
- Benjamin C. Orsburn
- , Yuting Yuan
- & Namandjé N. Bumpus
-
Article
| Open Access Reduced secretion of neuronal growth regulator 1 contributes to impaired adipose-neuronal crosstalk in obesity
Reduced secretion of neuronal growth regulator 1 contributes to impaired adipose-neuronal crosstalk in obesity
Adipose tissue is an important secretory organ, but less is known about the secretory activity of perivascular fat. Here the authors use proteomics analysis on secretomes from perivascular fat to identify neuronal growth regulator 1 as an adipocyte-derived neurotrophic factor, whose decreased secretion in obesity results in a loss of sympathetic innervation of adipose depots in mice.
- Elisa Duregotti
- , Christina M. Reumiller
- & Manuel Mayr
-
Article
| Open Access DOPAnization of tyrosine in α-synuclein by tyrosine hydroxylase leads to the formation of oligomers
DOPAnization of tyrosine in α-synuclein by tyrosine hydroxylase leads to the formation of oligomers
In this work, the authors show that α-synuclein is posttranslationally dopanized at Tyr136 by tyrosine hydroxylase, which facilitates the formation of oligomers. This modification likely impacts pathogenesis and the selective degeneration of dopaminergic neurons in Parkinson’s disease.
- Mingyue Jin
- , Sakiko Matsumoto
- & Shinji Hirotsune
-
Article
| Open Access Metabolite annotation from knowns to unknowns through knowledge-guided multi-layer metabolic networking
Metabolite annotation from knowns to unknowns through knowledge-guided multi-layer metabolic networking
Unknown metabolite annotation is a grand challenge in untargeted metabolomics. Here, the authors develop knowledge-guided multi-layer networking (KGMN) to enable global metabolite annotation from knowns to unknowns in untargeted metabolomics.
- Zhiwei Zhou
- , Mingdu Luo
- & Zheng-Jiang Zhu
-
Article
| Open Access Measurements of heterogeneity in proteomics analysis of the nanoparticle protein corona across core facilities
Measurements of heterogeneity in proteomics analysis of the nanoparticle protein corona across core facilities
Analysis of the protein corona formed around nanoparticles is important for multiple applications in nanomedicine, the methods used at core facilities used for analysis can impact the results. Here, the authors report on a study into the variability of the results obtained from 17 different core facilities and the implications of this.
- Ali Akbar Ashkarran
- , Hassan Gharibi
- & Morteza Mahmoudi
-
Article
| Open Access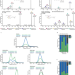 Analytical and computational workflow for in-depth analysis of oxidized complex lipids in blood plasma
Analytical and computational workflow for in-depth analysis of oxidized complex lipids in blood plasma
Oxidized lipids are prominent bioactive agents, and yet their molecular repertoire remains largely unknown. Here, the authors apply bioinformatics and LC-MS/MS to uncover the diversity and specificity of modified lipids in human blood plasma of lean and obese individuals.
- Angela Criscuolo
- , Palina Nepachalovich
- & Maria Fedorova
-
Article
| Open Access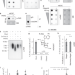 Mammalian N1-adenosine PARylation is a reversible DNA modification
Mammalian N1-adenosine PARylation is a reversible DNA modification
Poly-ADP-ribosylation (PARylation) is a well-known posttranslational modification of proteins. Here the authors show that beyond proteins also mammalian single-stranded DNA is PARylated in vitro and in vivo.
- Michael U. Musheev
- , Lars Schomacher
- & Christof Niehrs
-
Article
| Open Access Spatial snapshots of amyloid precursor protein intramembrane processing via early endosome proteomics
Spatial snapshots of amyloid precursor protein intramembrane processing via early endosome proteomics
Methods to assess organellar content are important. Here, Park et al develop a method for rapid isolation of early/sorting endosomes and demonstrate the application of the approach for analysis of endosomal proteomes and lipidomes, and for analysis of APP processing to Aβ via β and γ-Secretases.
- Hankum Park
- , Frances V. Hundley
- & J. Wade Harper
-
Article
| Open Access Inferring differential subcellular localisation in comparative spatial proteomics using BANDLE
Inferring differential subcellular localisation in comparative spatial proteomics using BANDLE
Changes in protein subcellular localization can be determined using mass spectrometry. Here, the authors present a statistical approach to determine relocalising proteins from spatial proteomics experiments.
- Oliver M. Crook
- , Colin T. R. Davies
- & Kathryn S. Lilley
-
Article
| Open Access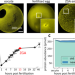 Differential nuclear import sets the timing of protein access to the embryonic genome
Differential nuclear import sets the timing of protein access to the embryonic genome
Here the authors address how embryos control the timing of specific gene activation in early frog development. They find transcription factors for early gene activation are maternally loaded and remain at constant levels, and rather that order of activation is based on their sequential entry into the nucleus based largely on their respective affinity to importins.
- Thao Nguyen
- , Eli J. Costa
- & Martin Wühr
-
Article
| Open Access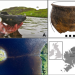 Neolithic culinary traditions revealed by cereal, milk and meat lipids in pottery from Scottish crannogs
Neolithic culinary traditions revealed by cereal, milk and meat lipids in pottery from Scottish crannogs
Despite archaeobotanical evidence for domesticated cereals, organic residue evidence is scarce. Here, the authors identify cereal-specific markers in pottery from Scottish ‘crannogs’, revealing the presence of cereals in Neolithic pottery which might have been mixed with dairy products as a milk-based gruel.
- Simon Hammann
- , Rosie R. Bishop
- & Lucy J. E. Cramp
-
Article
| Open Access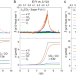 Oxidative decomposition mechanisms of lithium carbonate on carbon substrates in lithium battery chemistries
Oxidative decomposition mechanisms of lithium carbonate on carbon substrates in lithium battery chemistries
Lithium carbonate is ubiquitous in lithium battery chemistries and leads to overpotentials, however its oxidative decomposition is unclear. Here, the authors study its decomposition in ether electrolyte, clarify the role of the carbon substrate, and propose a route to limit released singlet oxygen.
- Deqing Cao
- , Chuan Tan
- & Yuhui Chen
-
Article
| Open Access Spatiotemporal-resolved protein networks profiling with photoactivation dependent proximity labeling
Spatiotemporal-resolved protein networks profiling with photoactivation dependent proximity labeling
Methods to identify protein interaction networks often suffer from poor spatiotemporal resolution. Here the authors present a light-activated proximity labeling method where the protein of interest is fused to the photosensitizer protein miniSOG, allowing temporally resolved labeling of interactors.
- Yansheng Zhai
- , Xiaoyan Huang
- & Gang Li
-
Article
| Open Access Structural basis of transcriptional regulation by a nascent RNA element, HK022 putRNA
Structural basis of transcriptional regulation by a nascent RNA element, HK022 putRNA
HK022 put is an RNA element that inhibits transcription termination without aids from protein factors. Here, authors solved cryo-EM structures of put-associated RNA polymerase and showed the structure of putRNA and its binding to the RNA polymerase.
- Seungha Hwang
- , Paul Dominic B. Olinares
- & Jin Young Kang
-
Article
| Open Access Native metabolomics identifies the rivulariapeptolide family of protease inhibitors
Native metabolomics identifies the rivulariapeptolide family of protease inhibitors
Bioactivity-guided isolation of specialized metabolites is an iterative process. Here, the authors demonstrate a native metabolomics approach that allows for fast screening of complex metabolite extracts against a protein of interest and simultaneous structure annotation.
- Raphael Reher
- , Allegra T. Aron
- & Daniel Petras
-
Article
| Open Access FLASHIda enables intelligent data acquisition for top–down proteomics to boost proteoform identification counts
FLASHIda enables intelligent data acquisition for top–down proteomics to boost proteoform identification counts
Data acquisition suitable for top-down proteomics (TDP) has the potential to significantly improve proteoform analysis. Here, the authors present FLASHIda, an intelligent online data acquisition algorithm for TDP that nearly doubles the number of proteoform-level identifications in complex samples.
- Kyowon Jeong
- , Maša Babović
- & Oliver Kohlbacher
-
Article
| Open Access TidyMass an object-oriented reproducible analysis framework for LC–MS data
TidyMass an object-oriented reproducible analysis framework for LC–MS data
Reproducibility, traceability, and transparency have been long-standing issues in metabolomics data analysis. Here, the authors present tidyMass, an R-based computational framework that allows designing traceable, shareable, and reproducible data processing and analysis workflows for untargeted metabolomics.
- Xiaotao Shen
- , Hong Yan
- & Michael P. Snyder
-
Article
| Open Access Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction
Protein shape sampled by ion mobility mass spectrometry consistently improves protein structure prediction
Collision cross sections (CCS) from ion mobility mass spectrometry provide information about protein shape and size. Here, the authors develop an algorithm to predict CCS and integrate experimental ion mobility data into Rosetta-based molecular modelling to predict protein structures from sequence.
- SM Bargeen Alam Turzo
- , Justin T. Seffernick
- & Steffen Lindert
-
Article
| Open Access The surfaceome of multiple myeloma cells suggests potential immunotherapeutic strategies and protein markers of drug resistance
The surfaceome of multiple myeloma cells suggests potential immunotherapeutic strategies and protein markers of drug resistance
The myeloma cell surface proteome regulates plasma cell biology and delineates therapy targets. Here, the authors profile the myeloma surfaceome at baseline and in drug resistance, finding the potential target CCR10, and include a streamlined approach to primary sample analysis.
- Ian D. Ferguson
- , Bonell Patiño-Escobar
- & Arun P. Wiita
-
Article
| Open Access Scalable multiplex co-fractionation/mass spectrometry platform for accelerated protein interactome discovery
Scalable multiplex co-fractionation/mass spectrometry platform for accelerated protein interactome discovery
Co-fractionation/mass spectrometry (CF/MS) allows mapping protein interactomes but efficiency and quantitative accuracy are limited. Here, the authors develop a reproducible multiplexed CF/MS method and apply it to characterize interactome rewiring in breast cancer cells.
- Pierre C. Havugimana
- , Raghuveera Kumar Goel
- & Andrew Emili
-
Article
| Open Access Mimicked synthetic ribosomal protein complex for benchmarking crosslinking mass spectrometry workflows
Mimicked synthetic ribosomal protein complex for benchmarking crosslinking mass spectrometry workflows
Cross-linking mass spectrometry is widely used to elucidate protein structures and interactions. Here, the authors generate an extensive peptide library to benchmark the most common cross-link search engines with frequently used cross-linking reagents in low and high complex sample systems.
- Manuel Matzinger
- , Adrian Vasiu
- & Karl Mechtler
-
Article
| Open Access Characterization of core fucosylation via sequential enzymatic treatments of intact glycopeptides and mass spectrometry analysis
Characterization of core fucosylation via sequential enzymatic treatments of intact glycopeptides and mass spectrometry analysis
Core fucosylation of N-linked glycoproteins has been linked to physiological and pathological processes. Here, the authors develop a mass spectrometry-based method that utilizes Endo F3 followed by PNGase F treatment to quantify site-specific glycoprotein core fucosylation in protein mixtures.
- Liwei Cao
- , T. Mamie Lih
- & Hui Zhang

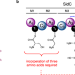 Elucidating the molecular programming of a nonlinear non-ribosomal peptide synthetase responsible for fungal siderophore biosynthesis
Elucidating the molecular programming of a nonlinear non-ribosomal peptide synthetase responsible for fungal siderophore biosynthesis