Featured
-
-
Article
| Open Access Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
During zygotic genome activation the embryo must re-wire the regulatory network that sustains totipotency earlier during development. Here they identify ZFP352 as an essential factor that targets retrotransposon families to facilitate dissolution of the totipotency network and enable ZGA.
- Zhengyi Li
- , Haiyan Xu
- & Hongqing Liang
-
Article
| Open Access Amniotes co-opt intrinsic genetic instability to protect germ-line genome integrity
Amniotes co-opt intrinsic genetic instability to protect germ-line genome integrity
Pachytene Piwi-interacting RNAs (piRNAs) expressed in mammalian germ lines are abundant, but their evolution and function are not fully understood. Here, the authors find that pachytene piRNA loci are hotspots of structural variation, which underlies rapid piRNA birth, divergence, and loss.
- Yu H. Sun
- , Hongxiao Cui
- & Xin Zhiguo Li
-
Article
| Open Access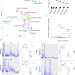 Primate-specific transposable elements shape transcriptional networks during human development
Primate-specific transposable elements shape transcriptional networks during human development
The human genome harbors more than 4.5 million transposable element (TE)-derived insertions, the result of recurrent waves of invasion and internal propagation. Here they show that TEs belonging to evolutionarily recent subfamilies go on to regulate later stages of human embryonic development, notably conditioning the expression of genes involved in gastrulation and early organogenesis.
- Julien Pontis
- , Cyril Pulver
- & Didier Trono
-
Article
| Open Access The impact of transposable elements on tomato diversity
The impact of transposable elements on tomato diversity
Transposable element insertion polymorphisms (TIPs) are a potential source of large effect alleles. Here, the authors use genome resequencing data for 602 tomato accessions together with transcriptomic and extensive phenotypic information to investigate the contribution of TIPs to tomato diversity.
- Marisol Domínguez
- , Elise Dugas
- & Leandro Quadrana
-
Article
| Open Access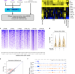 Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia
Endogenous retroviruses are a source of enhancers with oncogenic potential in acute myeloid leukaemia
Transposable elements are a potential source of transcriptional regulators, but how these sequences contribute to oncogenesis remains poorly understood. Here, the authors identify endogenous retroviruses (ERVs) with acute myeloid leukemia (AML)-associated enhancer chromatin signatures, and provide evidence that ERV activation provides an additional layer of gene regulation in AML.
- Özgen Deniz
- , Mamataz Ahmed
- & Miguel R. Branco
-
Article
| Open Access Functional disease architectures reveal unique biological role of transposable elements
Functional disease architectures reveal unique biological role of transposable elements
Transposable elements (TE) make up a large component of the human genome and have been shown to contribute to human diseases. Here, Hormozdiari
et al . estimate the contribution of TEs to the heritability of 41 complex traits and diseases and find enrichment of SINEs in blood traits.- Farhad Hormozdiari
- , Bryce van de Geijn
- & Alkes L. Price
-
Article
| Open Access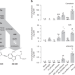 Expansion of a core regulon by transposable elements promotes Arabidopsis chemical diversity and pathogen defense
Expansion of a core regulon by transposable elements promotes Arabidopsis chemical diversity and pathogen defense
Arabidopsis plants can produce 4-hydroxyindole-3-carbonitrile (4OH-ICN) upon pathogen infection. Here, the authors show that EPCOT3, a retrotransposonderived enhancer, mediates WRKY33-binding, pathogen-responsive transcription of CYP82C2, and synthesis of 4OH-ICN.
- Brenden Barco
- , Yoseph Kim
- & Nicole K. Clay
-
Article
| Open Access Retrotranspositional landscape of Asian rice revealed by 3000 genomes
Retrotranspositional landscape of Asian rice revealed by 3000 genomes
Transposable elements (TE) are the dominant constituent of plant genomes. Here the authors develop a tool to analyze TE insertion sites in 3000 rice genomes and provide evidence for recent TE activity during cultivation and that external, rather than genetic, stimuli trigger most activations.
- Marie-Christine Carpentier
- , Ernandes Manfroi
- & Olivier Panaud
-
Article
| Open Access Genome sequence of the basal haplorrhine primate Tarsius syrichta reveals unusual insertions
Genome sequence of the basal haplorrhine primate Tarsius syrichta reveals unusual insertions
Tarsiers occupy a key node between strepsirrhines and anthropoids in the primate phylogeny. Here, Warren and colleagues present the genome of Tarsius syrichta, including a survey of transposable elements, an unusual mitochondrial insertion, and evidence for positive gene selection.
- Jürgen Schmitz
- , Angela Noll
- & Wesley C. Warren
-
Article |
 A family of transposable elements co-opted into developmental enhancers in the mouse neocortex
A family of transposable elements co-opted into developmental enhancers in the mouse neocortex
The neocortex is a mammalian-specific structure that is responsible for higher functions but details of how it evolved are lacking. Here the authors show that the transposable element family MER130 is highly enriched among the enhancers in the developing mouse neocortex, suggesting a role in the evolution of this structure.
- James H. Notwell
- , Tisha Chung
- & Gill Bejerano
-
Article
| Open Access SINE transcription by RNA polymerase III is suppressed by histone methylation but not by DNA methylation
SINE transcription by RNA polymerase III is suppressed by histone methylation but not by DNA methylation
Transcription of short interspersed nuclear elements (SINEs) allows retrotransposition that contributes to chromosomal instability. Here, the authors show that methylation of histones, rather than DNA, plays a dominant role in suppressing SINE expression.
- Dhaval Varshney
- , Jana Vavrova-Anderson
- & Robert J. White
-
Article |
 Gene silencing by CRISPR interference in mycobacteria
Gene silencing by CRISPR interference in mycobacteria
Recombination-based tools for generating targeted mutations in Mycobacterium tuberculosislack efficiency. Here the authors present a CRISPR interference approach that is able to efficiently repress the expression of target genes in mycobacteria, in a rapid and cost-effective manner.
- Eira Choudhary
- , Preeti Thakur
- & Nisheeth Agarwal
-
Article
| Open Access Transposable element islands facilitate adaptation to novel environments in an invasive species
Transposable element islands facilitate adaptation to novel environments in an invasive species
Genetic variation is key to species evolution. Here the authors sequence two phenotypically distinct populations of the ant Cardiocondyla obscurior, and find accumulations of transposable elements correlating with genetic variation that may have a role in differentiation, adaptation and speciation.
- Lukas Schrader
- , Jay W. Kim
- & Jan Oettler
-
Article |
 Autophagy supports genomic stability by degrading retrotransposon RNA
Autophagy supports genomic stability by degrading retrotransposon RNA
Retrotransposons replicate via a copy-paste mechanism involving a cytoplasmic intermediate. Guo et al.report that autophagy can suppress genetic variation by degrading cytoplasmic retrotransposon RNA, suggesting additional means by which autophagy may influence tumorigenesis.
- Huishan Guo
- , Maneka Chitiprolu
- & Derrick Gibbings
-
Article |
 SIRT6 represses LINE1 retrotransposons by ribosylating KAP1 but this repression fails with stress and age
SIRT6 represses LINE1 retrotransposons by ribosylating KAP1 but this repression fails with stress and age
Retrotransposons are repetitive sequences in the genome that can amplify themselves and whose activity has been linked to age-related pathologies. Here, Van Meter et al.report that the histone deacetylase SIRT6 represses activity of the L1 retrotransposon by ribosylating the nuclear corepressor protein, KAP1.
- Michael Van Meter
- , Mehr Kashyap
- & Vera Gorbunova
-
Article |
 RNA editing regulates transposon-mediated heterochromatic gene silencing
RNA editing regulates transposon-mediated heterochromatic gene silencing
The Hoppel transposable element mediates heterochromatin formation in Drosophila. Here Savva et al. report that the RNA-editing enzyme, ADAR, edits a long double-stranded RNA generated by the Hoppeltransposon, thereby regulating heterochromatin formation and gene expression.
- Yiannis A. Savva
- , James E. C. Jepson
- & Robert A. Reenan
-
Article |
 Recombinant SINEs are formed at high frequency during induced retrotransposition in vivo
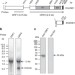
Recombinant SINEs are formed at high frequency during induced retrotransposition in vivo
SINEs are retrotransposons that insert exact copies of themselves into genomes. Using a marked copy of a SINE, Yadavet al. show that the sequences of newly transposed SINEs are a combination of marked and existing SINEs, suggesting a mechanism for the formation of mosaic SINEs.
- Vijay Pal Yadav
- , Prabhat Kumar Mandal
- & Sudha Bhattacharya
-
Article |
 Large-scale DNA editing of retrotransposons accelerates mammalian genome evolution
Large-scale DNA editing of retrotransposons accelerates mammalian genome evolution
APOBEC3 is a DNA editing enzyme that is important for antiviral responses. In this study, Carmi and colleagues show that APOBEC3 editing of retrotransposon sequences in mammalian genomes is widespread, with implications for the evolution of retrotransposons.
- Shai Carmi
- , George M. Church
- & Erez Y. Levanon
-
Article
| Open Access Mesozoic retroposons reveal parrots as the closest living relatives of passerine birds
Mesozoic retroposons reveal parrots as the closest living relatives of passerine birds
Zebra finches are passerine birds, but their phylogenetic relationship with non-passerine birds remains controversial. By examining retroposon insertion loci in avian genomes, the authors reveal that parrots are the closest relatives of passerines, which may have implications for understanding the evolution of birdsong.
- Alexander Suh
- , Martin Paus
- & Jürgen Schmitz
-
Article |
 Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences
Tumour microvesicles contain retrotransposon elements and amplified oncogene sequences
Microvesicles containing RNA are released from tumour cells. Here, the authors show that microvesicles released from tumour cells in culture have amplified levels of thec-Myconcogene, which is also found in the cell of origin, suggesting that microvesicles could be used as biomarkers.
- Leonora Balaj
- , Ryan Lessard
- & Johan Skog
-
Article |
 Insertion sequence-excision enhancer removes transposable elements from bacterial genomes and induces various genomic deletions
Insertion sequence-excision enhancer removes transposable elements from bacterial genomes and induces various genomic deletions
Insertion sequences are transposable elements that are found in the genomes of many bacteria. Here, the authors identify an enhancer element that results in a high frequency of excision of insertion elements, and suggest that the excision enhancer element coevolved with the insertion sequences.
- Masahiro Kusumoto
- , Tadasuke Ooka
- & Tetsuya Hayashi

 Transposable elements mediate genetic effects altering the expression of nearby genes in colorectal cancer
Transposable elements mediate genetic effects altering the expression of nearby genes in colorectal cancer