Article
|
Open Access
Featured
-
-
Article |
 Structural insights into BCDX2 complex function in homologous recombination
Structural insights into BCDX2 complex function in homologous recombination
Analyses of the structure and biochemical properties of the tetrameric complex of RAD51B, RAD51C, RAD51D and XRCC2 reveal details of its role in the repair of DNA double-strand breaks.
- Yashpal Rawal
- , Lijia Jia
- & Shaun K. Olsen
-
Article
| Open Access Mechanism of AAA+ ATPase-mediated RuvAB–Holliday junction branch migration
Mechanism of AAA+ ATPase-mediated RuvAB–Holliday junction branch migration
Structures of the ATP-hydrolysing RuvAB complex captured in multiple conformations provide mechanistic insights into coordinated ATPase and motor activity during DNA recombination.
- Jiri Wald
- , Dirk Fahrenkamp
- & Thomas C. Marlovits
-
Article
| Open Access RecA finds homologous DNA by reduced dimensionality search
RecA finds homologous DNA by reduced dimensionality search
Observations of rapid repair of double-stranded DNA breaks in sister choromosomes in Escherichia coli are consistent with a reduced-dimensionality-search model of RecA-mediated repair.
- Jakub Wiktor
- , Arvid H. Gynnå
- & Johan Elf
-
Article |
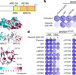 BARD1 reads H2A lysine 15 ubiquitination to direct homologous recombination
BARD1 reads H2A lysine 15 ubiquitination to direct homologous recombination
A tandem BRCT-domain-associated ubiquitin-dependent recruitment motif in BARD1 recruits BRCA1 to DNA double-strand breaks (DSBs) to promote homologous recombination and antagonize the 53BP1 DSB repair pathway that mediates non-homologous end joining.
- Jordan R. Becker
- , Gillian Clifford
- & J. Ross Chapman
-
Article |
 RNA transcripts stimulate homologous recombination by forming DR-loops
RNA transcripts stimulate homologous recombination by forming DR-loops
RNA transcripts stimulate homologous recombination through the formation of DR-loops, intermediate structures that contain both DNA–DNA and DNA–RNA hybrids.
- Jian Ouyang
- , Tribhuwan Yadav
- & Lee Zou
-
Letter |
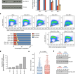 CDK12 regulates DNA repair genes by suppressing intronic polyadenylation
CDK12 regulates DNA repair genes by suppressing intronic polyadenylation
The kinase CDK12 suppresses usage of intronic polyadenylation sites and thereby promotes the expression of genes involved in homologous recombination DNA repair.
- Sara J. Dubbury
- , Paul L. Boutz
- & Phillip A. Sharp
-
Article |
 DYNLL1 binds to MRE11 to limit DNA end resection in BRCA1-deficient cells
DYNLL1 binds to MRE11 to limit DNA end resection in BRCA1-deficient cells
DYNLL1 antagonizes end resection of DNA double-strand breaks, thereby inhibiting homologous repair, and the loss of DYNLL1 correlates with poor progression-free survival of patients with BRCA1-mutant ovarian cancer.
- Yizhou Joseph He
- , Khyati Meghani
- & Dipanjan Chowdhury
-
Letter |
 EWS–FLI1 increases transcription to cause R-loops and block BRCA1 repair in Ewing sarcoma
EWS–FLI1 increases transcription to cause R-loops and block BRCA1 repair in Ewing sarcoma
The EWS–FLI1 fusion protein, expressed in Ewing sarcoma, increases global transcription, causes accumulation of R loops and replication stress, and impairs BRCA1-mediated repair.
- Aparna Gorthi
- , July Carolina Romero
- & Alexander J. R. Bishop
-
Article |
 Mechanism of tandem duplication formation in BRCA1-mutant cells
Mechanism of tandem duplication formation in BRCA1-mutant cells
BRCA1, but not BRCA2, suppresses the formation of tandem duplications at stalled replication forks in primary mammalian cells.
- Nicholas A. Willis
- , Richard L. Frock
- & Ralph Scully
-
Article |
 BRCA1–BARD1 promotes RAD51-mediated homologous DNA pairing
BRCA1–BARD1 promotes RAD51-mediated homologous DNA pairing
The tumour suppressor complex BRCA1–BARD1, which facilitates the generation of a single-stranded DNA template during homologous recombination, also binds to the recombinase RAD51 and enhances its function.
- Weixing Zhao
- , Justin B. Steinfeld
- & Patrick Sung
-
Letter |
 A mechanism for the suppression of homologous recombination in G1 cells
A mechanism for the suppression of homologous recombination in G1 cells
A mechanism for the repression of homologous recombination in G1, the stage of the cell cycle preceding replication, is determined; the critical aspects are the interaction between BRCA1 and PALB2–BRCA2, and suppression of DNA-end resection.
- Alexandre Orthwein
- , Sylvie M. Noordermeer
- & Daniel Durocher
-
Letter |
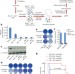 REV7 counteracts DNA double-strand break resection and affects PARP inhibition
REV7 counteracts DNA double-strand break resection and affects PARP inhibition
Loss of REV7 is shown to regulate end resection of double-stranded DNA breaks in BRCA1-deficient cells, leading to PARP inhibitor resistance and restoration of homologous recombination; REV7 dictates pathway choice in BRCA1-deficient cells and during immunoglobulin class switching.
- Guotai Xu
- , J. Ross Chapman
- & Sven Rottenberg
-
Letter |
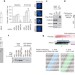 Homologous-recombination-deficient tumours are dependent on Polθ-mediated repair
Homologous-recombination-deficient tumours are dependent on Polθ-mediated repair
In studies in mammalian cells, polymerase theta (Polθ, also known as POLQ) is identified as the polymerase responsible for non-homologous end joining DNA repair; this DNA repair pathway acts in many tumours when homologous recombination is inactivated and the identification of the polymerase responsible may aid the development of new therapeutic approaches.
- Raphael Ceccaldi
- , Jessica C. Liu
- & Alan D. D’Andrea
-
Letter |
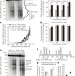 Tel1ATM-mediated interference suppresses clustered meiotic double-strand-break formation
Tel1ATM-mediated interference suppresses clustered meiotic double-strand-break formation
Meiotic recombination is initiated by a fairly uniform distribution of hundreds of DNA double-strand breaks catalysed by the Spo11 protein; here, Tel1 (orthologue of human ATM) is shown to be required for the localized inhibition that prevents double-strand breaks from forming close to one another.
- Valerie Garcia
- , Stephen Gray
- & Matthew J. Neale
-
Letter |
 Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks
Sae2 promotes dsDNA endonuclease activity within Mre11–Rad50–Xrs2 to resect DNA breaks
The MRX complex, required for double-strand break (DSB) repair by homologous recombination, has 3′ to 5′ exonuclease activity, but homologous recombination at a DSB uses a 3′-tailed molecule, which requires resection of the 5′ strand; here it is shown that in yeast, Sae2 nuclease promotes MRX to make an initial endonucleolytic cut on the 5′ strand that may allow MRX to digest the 5′ strand back to the end in a 3′ to 5′ fashion.
- Elda Cannavo
- & Petr Cejka
-
Letter |
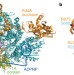 Structural basis for translocation by AddAB helicase–nuclease and its arrest at χ sites
Structural basis for translocation by AddAB helicase–nuclease and its arrest at χ sites
A dual-function helicase–nuclease, typified by RecBCD in Escherichia coli, acts on free DNA ends during bacterial double-stranded break repair until it reaches a χ sequence at which it pauses before continuing with modified enzymatic properties; here several crystal structures of the related AddAB enzyme from Bacillus subtilis bound to χ-containing DNA are presented, offering insight into χ recognition and its effect on DNA translocation.
- Wojciech W. Krajewski
- , Xin Fu
- & Dale B. Wigley
-
Letter |
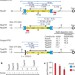 Recombination-restarted replication makes inverted chromosome fusions at inverted repeats
Recombination-restarted replication makes inverted chromosome fusions at inverted repeats
A new mechanism of chromosomal rearrangement is identified through the observation that broken or collapsed DNA replication forks restarted by homologous recombination have a high propensity for U-turns at short inverted repeats; the error-prone nature of this mechanism is suggested to contribute to gross chromosomal rearrangements and copy-number variations present in cancer and other genomic disorders.
- Ken’Ichi Mizuno
- , Izumi Miyabe
- & Johanne M. Murray
-
Letter |
 Single-molecule imaging of DNA pairing by RecA reveals a three-dimensional homology search
Single-molecule imaging of DNA pairing by RecA reveals a three-dimensional homology search
The search for DNA homology is vital to recombinational DNA repair and occurs by intersegment contact sampling wherein the three-dimensional conformational state of the double-stranded DNA target and the length of the homologous RecA–single-stranded DNA filament have important roles.
- Anthony L. Forget
- & Stephen C. Kowalczykowski
-
Letter |
 Rad51 paralogues Rad55–Rad57 balance the antirecombinase Srs2 in Rad51 filament formation
Rad51 paralogues Rad55–Rad57 balance the antirecombinase Srs2 in Rad51 filament formation
- Jie Liu
- , Ludovic Renault
- & Wolf-Dietrich Heyer
-
Letter |
 RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II
RNAi promotes heterochromatic silencing through replication-coupled release of RNA Pol II
- Mikel Zaratiegui
- , Stephane E. Castel
- & Robert A. Martienssen
-
Letter |
 Bidirectional resection of DNA double-strand breaks by Mre11 and Exo1
Bidirectional resection of DNA double-strand breaks by Mre11 and Exo1
- Valerie Garcia
- , Sarah E. L. Phelps
- & Matthew J. Neale
-
Letter |
 DNA end resection by Dna2–Sgs1–RPA and its stimulation by Top3–Rmi1 and Mre11–Rad50–Xrs2
DNA end resection by Dna2–Sgs1–RPA and its stimulation by Top3–Rmi1 and Mre11–Rad50–Xrs2
When double-strand breaks occur in DNA, the broken ends must undergo processing to prepare them for repair. Here, and in an accompanying study, this processing reaction has now been replicated in vitro using yeast proteins. Processing minimally requires the activities of a helicase, a nuclease and a single-strand-binding protein, although the reaction is enhanced by the addition of three factors that help to target the core complex and stimulate the unwinding activity.
- Petr Cejka
- , Elda Cannavo
- & Stephen C. Kowalczykowski
-
Article |
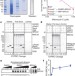 Purified human BRCA2 stimulates RAD51-mediated recombination
Purified human BRCA2 stimulates RAD51-mediated recombination
The two hereditary breast cancer susceptibility genes, BRCA1 and BRCA2, have roles in responding to DNA damage. When they are mutated or absent, genomic instability, a contributory factor to cancer development, results. Studies of BRCA2 have been hampered by its large size, which makes purification of the full-length protein challenging. These authors report the first in vitro characterization of full-length BRCA2 and delineate the different ways by which BRCA2 facilitates RAD51-mediated homologous recombination.
- Ryan B. Jensen
- , Aura Carreira
- & Stephen C. Kowalczykowski
-
Letter |
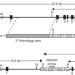 Production of p53 gene knockout rats by homologous recombination in embryonic stem cells
Production of p53 gene knockout rats by homologous recombination in embryonic stem cells
The rat is a animal model widely used for studying human physiology and disease, but functional genomics and genetic research have been stifled by the limited availability of gene targeting tools. These authors have established gene targeting by homologous recombination in rat embryonic stem cells, and have generated p53 gene knockout rats for the first time.
- Chang Tong
- , Ping Li
- & Qi-Long Ying
-
Letter |
 Double Holliday junctions are intermediates of DNA break repair
Double Holliday junctions are intermediates of DNA break repair
In meiotic cells paired homologues are joined by a set of crossovers known as a double Holliday junction (DHJ). Whether DHJs form during mitotic recombination has been unclear, as mitotic cells possess alternative repair pathways that would not require DHJ formation. Here it is demonstrated that mitotic and meiotic cells form similar DHJs, but that the levels in mitotic cells are approximately 10–fold lower, and show a preference for joints between sister chromatids rather than homologues. Consequently, in mitotic cells non–crossover outcomes are favoured.
- Malgorzata Bzymek
- , Nathaniel H. Thayer
- & Neil Hunter

 Long-molecule scars of backup DNA repair in BRCA1- and BRCA2-deficient cancers
Long-molecule scars of backup DNA repair in BRCA1- and BRCA2-deficient cancers