Featured
-
-
Article
| Open Access Cell type specific long non-coding RNA targets identified by integrative analysis of single-cell and bulk colorectal cancer transcriptomes
Cell type specific long non-coding RNA targets identified by integrative analysis of single-cell and bulk colorectal cancer transcriptomes
- Ante Mihaljevic
- , Philip D. Rubin
- & Roberta Esposito
-
Article
| Open Access Differential expression and global analysis of miR156/SQUAMOSA promoter binding-like proteins (SPL) module in oat
Differential expression and global analysis of miR156/SQUAMOSA promoter binding-like proteins (SPL) module in oat
- Mehtab-Singh
- , Rajiv K. Tripathi
- & Jaswinder Singh
-
Article
| Open Access Identification of m6A/m5C-related lncRNA signature for prediction of prognosis and immunotherapy efficacy in esophageal squamous cell carcinoma
Identification of m6A/m5C-related lncRNA signature for prediction of prognosis and immunotherapy efficacy in esophageal squamous cell carcinoma
- Jianlin Wang
- , Huiwen Ren
- & Xinye Ni
-
Article
| Open Access Genetic determinants of plasma protein levels in the Estonian population
Genetic determinants of plasma protein levels in the Estonian population
- Anette Kalnapenkis
- , Maarja Jõeloo
- & Urmo Võsa
-
Article
| Open Access Enriched atlas of lncRNA and protein-coding genes for the GRCg7b chicken assembly and its functional annotation across 47 tissues
Enriched atlas of lncRNA and protein-coding genes for the GRCg7b chicken assembly and its functional annotation across 47 tissues
- Fabien Degalez
- , Mathieu Charles
- & Sandrine Lagarrigue
-
Article
| Open Access Genome-wide identification and expression analysis of NF-Y gene family in tobacco (Nicotiana tabacum L.)
Genome-wide identification and expression analysis of NF-Y gene family in tobacco (Nicotiana tabacum L.)
- Yue Tian
- , Kangkang Song
- & Long Yang
-
Article
| Open Access Cacao pod transcriptome profiling of seven genotypes identifies features associated with post-penetration resistance to Phytophthora palmivora
Cacao pod transcriptome profiling of seven genotypes identifies features associated with post-penetration resistance to Phytophthora palmivora
- Indrani K. Baruah
- , Jonathan Shao
- & Stephen P. Cohen
-
Article
| Open Access Multi-omics resources for the Australian southern stuttering frog (Mixophyes australis) reveal assorted antimicrobial peptides
Multi-omics resources for the Australian southern stuttering frog (Mixophyes australis) reveal assorted antimicrobial peptides
- Simon Tang
- , Emma Peel
- & Katherine A. Farquharson
-
Article
| Open Access Characterization of enhancer activity in early human neurodevelopment using Massively Parallel Reporter Assay (MPRA) and forebrain organoids
Characterization of enhancer activity in early human neurodevelopment using Massively Parallel Reporter Assay (MPRA) and forebrain organoids
- Davide Capauto
- , Yifan Wang
- & Flora M. Vaccarino
-
Article
| Open Access Whole transcriptome screening for novel genes involved in meiosis and fertility in Drosophila melanogaster
Whole transcriptome screening for novel genes involved in meiosis and fertility in Drosophila melanogaster
- Siqi Sun
- , Tyler Defosse
- & Kim S. McKim
-
Article
| Open Access GWAS-identified hyperuricemia-associated IGF1R variant rs6598541 has a limited role in urate mediated inflammation in human mononuclear cells
GWAS-identified hyperuricemia-associated IGF1R variant rs6598541 has a limited role in urate mediated inflammation in human mononuclear cells
- Orsolya I. Gaal
- , Ruiqi Liu
- & Leo A. B. Joosten
-
Article
| Open Access Cross-regulation of Aps-promoters in Lacticaseibacillus paracasei by the PsdR response regulator in response to lantibiotics
Cross-regulation of Aps-promoters in Lacticaseibacillus paracasei by the PsdR response regulator in response to lantibiotics
- Qian Zhang
- , Manuel Zúñiga
- & Ainhoa Revilla-Guarinos
-
Article
| Open Access First neurotranscriptome of adults Tambaquis (Colossoma macropomum) with characterization and differential expression between males and females
First neurotranscriptome of adults Tambaquis (Colossoma macropomum) with characterization and differential expression between males and females
- Josy Miranda
- , Ivana Veneza
- & Grazielle Evangelista-Gomes
-
Article
| Open Access An RNA-Seq analysis of coronavirus in the skin of the Pangolin
An RNA-Seq analysis of coronavirus in the skin of the Pangolin
- Siwei Deng
- , Xuechen Tian
- & Siew Woh Choo
-
Article
| Open Access A SINE-VNTR-Alu at the LRIG2 locus is associated with proximal and distal gene expression in CRISPR and population models
A SINE-VNTR-Alu at the LRIG2 locus is associated with proximal and distal gene expression in CRISPR and population models
- Ashley Hall
- , Ben Middlehurst
- & John P. Quinn
-
Article
| Open Access A disulfidptosis-related lncRNAs signature in hepatocellular carcinoma: prognostic prediction, tumor immune microenvironment and drug susceptibility
A disulfidptosis-related lncRNAs signature in hepatocellular carcinoma: prognostic prediction, tumor immune microenvironment and drug susceptibility
- Yanqiong Liu
- , Jiyu Meng
- & Xue Qin
-
Article
| Open Access Transcriptome analysis in a humanized mouse model of familial dysautonomia reveals tissue-specific gene expression disruption in the peripheral nervous system
Transcriptome analysis in a humanized mouse model of familial dysautonomia reveals tissue-specific gene expression disruption in the peripheral nervous system
- Ricardo Harripaul
- , Elisabetta Morini
- & Susan Slaugenhaupt
-
Article
| Open Access TWAS revealed significant causal loci for milk production and its composition in Murrah buffaloes
TWAS revealed significant causal loci for milk production and its composition in Murrah buffaloes
- Supriya Chhotaray
- , Vikas Vohra
- & Gopal Gowane
-
Article
| Open Access Global gene regulatory network underlying miR165a in Arabidopsis shoot apical meristem
Global gene regulatory network underlying miR165a in Arabidopsis shoot apical meristem
- Sonali Sinha
- , Sudeep Sahadevan
- & Marcus G. Heisler
-
Article
| Open Access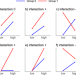 Benefit of using interaction effects for the analysis of high-dimensional time-response or dose-response data for two-group comparisons
Benefit of using interaction effects for the analysis of high-dimensional time-response or dose-response data for two-group comparisons
- Julia C. Duda
- , Carolin Drenda
- & Franziska Kappenberg
-
Article
| Open Access De novo transcriptome assembly of Dalbergia sissoo Roxb. (Fabaceae) under Botryodiplodia theobromae-induced dieback disease
De novo transcriptome assembly of Dalbergia sissoo Roxb. (Fabaceae) under Botryodiplodia theobromae-induced dieback disease
- Ummul Buneen Zafar
- , Muhammad Shahzaib
- & Iqrar Ahmad Rana
-
Article
| Open Access Transcriptomic analysis reveals that RasGEF1b deletion alters basal and LPS-induced expression of genes involved in chemotaxis and cytokine responses in macrophages
Transcriptomic analysis reveals that RasGEF1b deletion alters basal and LPS-induced expression of genes involved in chemotaxis and cytokine responses in macrophages
- Heliana B. Fernandes
- , Isadora Mafra de Oliveira
- & Aristóbolo M. Silva
-
Article
| Open Access Phylogenomics, phenotypic, and functional traits of five novel (Earth-derived) bacterial species isolated from the International Space Station and their prevalence in metagenomes
Phylogenomics, phenotypic, and functional traits of five novel (Earth-derived) bacterial species isolated from the International Space Station and their prevalence in metagenomes
- Anna C. Simpson
- , Pratyay Sengupta
- & Kasthuri Venkateswaran
-
Article
| Open Access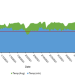 Comparative transcriptome analysis reveals candidate genes for cold stress response and early flowering in pineapple
Comparative transcriptome analysis reveals candidate genes for cold stress response and early flowering in pineapple
- Ashley G. Yow
- , Kanjana Laosuntisuk
- & Massimo Iorizzo
-
Article
| Open Access Genome-wide epigenetic modifications in sports horses during training as an adaptation phenomenon
Genome-wide epigenetic modifications in sports horses during training as an adaptation phenomenon
- Katia Cappelli
- , Samanta Mecocci
- & Stefano Capomaccio
-
Article
| Open Access Multi-breed host rumen epithelium transcriptome and microbiome associations and their relationship with beef cattle feed efficiency
Multi-breed host rumen epithelium transcriptome and microbiome associations and their relationship with beef cattle feed efficiency
- P. A. S. Fonseca
- , S. Lam
- & A. Cánovas
-
Article
| Open Access Proteomic and phosphoproteomic analyses of myectomy tissue reveals difference between sarcomeric and genotype-negative hypertrophic cardiomyopathy
Proteomic and phosphoproteomic analyses of myectomy tissue reveals difference between sarcomeric and genotype-negative hypertrophic cardiomyopathy
- Ramin Garmany
- , J. Martijn Bos
- & Michael J. Ackerman
-
Article
| Open Access Genome-wide analysis and characterization of the LRR-RLK gene family provides insights into anthracnose resistance in common bean
Genome-wide analysis and characterization of the LRR-RLK gene family provides insights into anthracnose resistance in common bean
- Caroline Marcela da Silva Dambroz
- , Alexandre Hild Aono
- & Welison Andrade Pereira
-
Article
| Open Access A novel quantitative targeted analysis of X-chromosome inactivation (XCI) using nanopore sequencing
A novel quantitative targeted analysis of X-chromosome inactivation (XCI) using nanopore sequencing
- Josefin Johansson
- , Sarah Lidéus
- & Maria Wilbe
-
Article
| Open Access Differential exon usage of developmental genes is associated with deregulated epigenetic marks
Differential exon usage of developmental genes is associated with deregulated epigenetic marks
- Hoang Thu Trang Do
- , Siba Shanak
- & Volkhard Helms
-
Article
| Open Access Identification of bromelain subfamily proteases encoded in the pineapple genome
Identification of bromelain subfamily proteases encoded in the pineapple genome
- Ashley G. Yow
- , Hamed Bostan
- & Massimo Iorizzo
-
Article
| Open Access Identification of candidate lethal haplotypes and genomic association with post-natal mortality and reproductive traits in Nellore cattle
Identification of candidate lethal haplotypes and genomic association with post-natal mortality and reproductive traits in Nellore cattle
- Patrícia Iana Schmidt
- , Lucio Flavio Macedo Mota
- & Lucia Galvão de Albuquerque
-
Article
| Open Access Rejection of Lepeophtheirus salmonis driven in part by chitin sensing is not impacted by seawater acclimitization in Coho salmon (Oncorhynchus kisutch)
Rejection of Lepeophtheirus salmonis driven in part by chitin sensing is not impacted by seawater acclimitization in Coho salmon (Oncorhynchus kisutch)
- Laura M. Braden
- , Dylan Michaud
- & Mark D. Fast
-
Article
| Open Access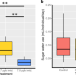 Gene expression in bumble bee larvae differs qualitatively between high and low concentration imidacloprid exposure levels
Gene expression in bumble bee larvae differs qualitatively between high and low concentration imidacloprid exposure levels
- Rubén Martín-Blázquez
- , Austin C. Calhoun
- & Sydney A. Cameron
-
Article
| Open Access Analysis of the impact of DGAT1 p.M435L and p.K232A variants on pre-mRNA splicing in a full-length gene assay
Analysis of the impact of DGAT1 p.M435L and p.K232A variants on pre-mRNA splicing in a full-length gene assay
- Nicolas Gaiani
- , Lorraine Bourgeois-Brunel
- & Arnaud Boulling
-
Article
| Open Access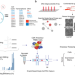 Multimodal perturbation analyses of cyclin-dependent kinases reveal a network of synthetic lethalities associated with cell-cycle regulation and transcriptional regulation
Multimodal perturbation analyses of cyclin-dependent kinases reveal a network of synthetic lethalities associated with cell-cycle regulation and transcriptional regulation
- Kyle Ford
- , Brenton P. Munson
- & Trey Ideker
-
Article
| Open Access Plasma 1H-NMR metabolic and amino acid profiles of newborn piglets from two lines divergently selected for residual feed intake
Plasma 1H-NMR metabolic and amino acid profiles of newborn piglets from two lines divergently selected for residual feed intake
- Laurence Liaubet
- , Camille Guilmineau
- & Hélène Quesnel
-
Article
| Open Access Neuropathological signatures revealed by transcriptomic and proteomic analysis in Pten-deficient mouse models
Neuropathological signatures revealed by transcriptomic and proteomic analysis in Pten-deficient mouse models
- Stanley K. K. Cheung
- , Jacinda Kwok
- & Andrew M. Chan
-
Article
| Open Access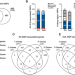 Molecular pathways identified from single nucleotide polymorphisms demonstrate mechanistic differences in systemic lupus erythematosus patients of Asian and European ancestry
Molecular pathways identified from single nucleotide polymorphisms demonstrate mechanistic differences in systemic lupus erythematosus patients of Asian and European ancestry
- Katherine A. Owen
- , Kristy A. Bell
- & Peter E. Lipsky
-
Article
| Open Access CRISPR-Cas9 genetic screen leads to the discovery of L-Moses, a KAT2B inhibitor that attenuates Tunicamycin-mediated neuronal cell death
CRISPR-Cas9 genetic screen leads to the discovery of L-Moses, a KAT2B inhibitor that attenuates Tunicamycin-mediated neuronal cell death
- Sofia Pavlou
- , Stefanie Foskolou
- & Emmanouil Metzakopian
-
Article
| Open Access Nine patients with KCNQ2-related neonatal seizures and functional studies of two missense variants
Nine patients with KCNQ2-related neonatal seizures and functional studies of two missense variants
- Suphalak Chokvithaya
- , Natarin Caengprasath
- & Vorasuk Shotelersuk
-
Article
| Open Access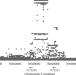 Distilling functional variations for human UGT2B4 upstream region based on selection signals and implications for phenotypes of Neanderthal and Denisovan
Distilling functional variations for human UGT2B4 upstream region based on selection signals and implications for phenotypes of Neanderthal and Denisovan
- Pin-Yi Wang
- , Yuan Yang
- & Chang Sun
-
Article
| Open Access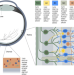 Post-GWAS screening of candidate genes for refractive error in mutant zebrafish models
Post-GWAS screening of candidate genes for refractive error in mutant zebrafish models
- Wim H. Quint
- , Kirke C. D. Tadema
- & Adriana I. Iglesias
-
Article
| Open Access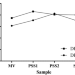 Multi-omics network model reveals key genes associated with p-coumaric acid stress response in an industrial yeast strain
Multi-omics network model reveals key genes associated with p-coumaric acid stress response in an industrial yeast strain
- F. E. Ciamponi
- , D. P. Procópio
- & M. M. Brandão
-
Article
| Open AccessUnravelling the genetics of non-random fertilization associated with gametic incompatibility
- Audrey A. A. Martin
- , Samir Id-Lahoucine
- & Flavio S. Schenkel
-
Article
| Open Access Blood-based gene expression as non-lethal tool for inferring salinity-habitat history of European eel (Anguilla anguilla)
Blood-based gene expression as non-lethal tool for inferring salinity-habitat history of European eel (Anguilla anguilla)
- Francesca Bertolini
- , Mehis Rohtla
- & Caroline M. F. Durif
-
Article
| Open Access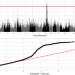 Analysis of merged transcriptomic and genomic datasets to identify genes and pathways underlying residual feed intake in growing pigs
Analysis of merged transcriptomic and genomic datasets to identify genes and pathways underlying residual feed intake in growing pigs
- Emil Ibragimov
- , Anni Øyan Pedersen
- & Peter Karlskov-Mortensen
-
Editorial
| Open AccessGenome editing
Recent advances in genome editing technologies have redefined our ability to probe and precisely edit the human genome and epigenome in vitro and in vivo. More specifically, RNA-guided CRISPR/Cas systems have revolutionized the field due to their simplicity in design and adaptability across biological systems. This Collection highlights results in CRISPR/Cas technology that increase the efficiency of precision genome editing, and allow genetic manipulation in model systems traditionally intractable to site-directed gene modification.
- Maura McGrail
- , Tetsushi Sakuma
- & Leonidas Bleris
-
Article
| Open Access C1QA, C1QB, and GZMB are novel prognostic biomarkers of skin cutaneous melanoma relating tumor microenvironment
C1QA, C1QB, and GZMB are novel prognostic biomarkers of skin cutaneous melanoma relating tumor microenvironment
- Zhuoshuai Liang
- , Lingfeng Pan
- & Lianbo Zhang

 Global transcriptome profiles provide insights into muscle cell development and differentiation on microstructured marine biopolymer scaffolds for cultured meat production
Global transcriptome profiles provide insights into muscle cell development and differentiation on microstructured marine biopolymer scaffolds for cultured meat production