Featured
-
-
Article
| Open Access Inferring differential subcellular localisation in comparative spatial proteomics using BANDLE
Inferring differential subcellular localisation in comparative spatial proteomics using BANDLE
Changes in protein subcellular localization can be determined using mass spectrometry. Here, the authors present a statistical approach to determine relocalising proteins from spatial proteomics experiments.
- Oliver M. Crook
- , Colin T. R. Davies
- & Kathryn S. Lilley
-
Article
| Open Access A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
A method for multiplexed full-length single-molecule sequencing of the human mitochondrial genome
Accurate analysis of mitochondrial DNA is important for mitochondrial disease clinical research and diagnostics. Here, authors present a method using Cas9 cleavage, nanopore sequencing and a custom pipeline to identify pathogenic variants, deletions and accurately quantify heteroplasmy to below 1%.
- Ieva Keraite
- , Philipp Becker
- & Ivo Glynne Gut
-
Article
| Open Access Batch effects removal for microbiome data via conditional quantile regression
Batch effects removal for microbiome data via conditional quantile regression
Here, the authors present ConQuR, a conditional quantile regression method that removes microbiome batch effects through non-parametric modeling of complex microbial read counts, while preserving the signals of interest.
- Wodan Ling
- , Jiuyao Lu
- & Michael C. Wu
-
Article
| Open Access An analysis of 45 large-scale wastewater sites in England to estimate SARS-CoV-2 community prevalence
An analysis of 45 large-scale wastewater sites in England to estimate SARS-CoV-2 community prevalence
Wastewater surveillance could provide a means of monitoring SARS-CoV-2 prevalence that does not rely on testing individuals. Here, the authors report results from England’s national wastewater surveillance program, use it to estimate prevalence, and compare estimates with those from population-based prevalence surveys.
- Mario Morvan
- , Anna Lo Jacomo
- & Leon Danon
-
Article
| Open Access Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity
Benchmarking of analysis strategies for data-independent acquisition proteomics using a large-scale dataset comprising inter-patient heterogeneity
Data independent acquisition (DIA) has been gaining momentum in clinical proteomics. Here, the authors create a benchmark dataset comprising inter-patient heterogeneity to compare popular DIA data analysis workflows for identifying differentially abundant proteins.
- Klemens Fröhlich
- , Eva Brombacher
- & Oliver Schilling
-
Article
| Open Access EPicker is an exemplar-based continual learning approach for knowledge accumulation in cryoEM particle picking
EPicker is an exemplar-based continual learning approach for knowledge accumulation in cryoEM particle picking
Many existing deep learning algorithms for particle picking are not predictable on unseen datasets. Here the authors report an exemplar-based continual learning approach, EPicker, enabling accumulation of new knowledge of cryoEM particle picking without catastrophic forgetting of old knowledge.
- Xinyu Zhang
- , Tianfang Zhao
- & Xueming Li
-
Article
| Open Access Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry
Pytheas: a software package for the automated analysis of RNA sequences and modifications via tandem mass spectrometry
RNA modifications represent a critical aspect of RNA biology that is not well suited to sequencing methods. Here, the authors provide a software tool for automated analysis of RNA tandem mass spectra with full support of modifications, isotope labelling, and control of false discovery rate.
- Luigi D’Ascenzo
- , Anna M. Popova
- & James R. Williamson
-
Article
| Open Access Genetic analysis of over half a million people characterises C-reactive protein loci
Genetic analysis of over half a million people characterises C-reactive protein loci
Inflammation is associated with a variety of diseases. Here, the authors identify 266 genetic loci associated with C-reactive protein levels, a marker of inflammation, in >500,000 Europeans, along with associated pathways, clinical outcomes and potential causal associations with disease.
- Saredo Said
- , Raha Pazoki
- & Abbas Dehghan
-
Article
| Open Access A universal deep neural network for in-depth cleaning of single-cell RNA-Seq data
A universal deep neural network for in-depth cleaning of single-cell RNA-Seq data
Single cell RNA sequencing (scRNA-Seq) is widely used in biomedical research. Here the authors develop a novel AI model-AutoClass, which effectively cleans a wide range of noise and artifacts in scRNA-Seq data and improves downstream analyses.
- Hui Li
- , Cory R. Brouwer
- & Weijun Luo
-
Article
| Open Access Comprehensive generation, visualization, and reporting of quality control metrics for single-cell RNA sequencing data
Comprehensive generation, visualization, and reporting of quality control metrics for single-cell RNA sequencing data
Quality control (QC) is a crucial step in single-cell RNA-seq data analysis. Here, the authors present the SCTK-QC pipeline which generates and visualizes a comprehensive set of QC metrics to streamline the process of detecting and removing poor quality cells and other artifacts.
- Rui Hong
- , Yusuke Koga
- & Joshua D. Campbell
-
Article
| Open Access The spatial transcriptomic landscape of the healing mouse intestine following damage
The spatial transcriptomic landscape of the healing mouse intestine following damage
The colon is comprised of specialized cells that interact with each other to function, however, the molecular regionalization of the colon is incompletely understood. Here, the authors use spatial transcriptomics to generate a publicly available resource defining the transcriptomic regionalization of the colon during steady state and mucosal healing.
- Sara M. Parigi
- , Ludvig Larsson
- & Eduardo J. Villablanca
-
Article
| Open Access Comparative metabolomics with Metaboseek reveals functions of a conserved fat metabolism pathway in C. elegans
Comparative metabolomics with Metaboseek reveals functions of a conserved fat metabolism pathway in C. elegans
Untargeted mass spectrometry-based metabolomics can reveal new biochemistry, but data analysis is challenging. Here, the authors develop Metaboseek, an open-source software that facilitates metabolite discovery, and apply it to characterize fatty acid alpha-oxidation in C. elegans.
- Maximilian J. Helf
- , Bennett W. Fox
- & Frank C. Schroeder
-
Article
| Open Access Microbiome differential abundance methods produce different results across 38 datasets
Microbiome differential abundance methods produce different results across 38 datasets
Many microbiome differential abundance methods are available, but it lacks systematic comparison among them. Here, the authors compare the performance of 14 differential abundance testing methods on 38 16S rRNA gene datasets with two sample groups, and show ALDEx2 and ANCOM-II produce the most consistent results.
- Jacob T. Nearing
- , Gavin M. Douglas
- & Morgan G. I. Langille
-
Article
| Open Access Computational optical sectioning with an incoherent multiscale scattering model for light-field microscopy
Computational optical sectioning with an incoherent multiscale scattering model for light-field microscopy
Light-field microscopy provides volumetric imaging at high speeds, but suffers from degradation in scattering tissue. Here, the authors present an incoherent multiscale scattering model which allows for quantitative 3D reconstruction in complex environments, and demonstrate dynamic imaging in vivo.
- Yi Zhang
- , Zhi Lu
- & Qionghai Dai
-
Article
| Open Access Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020
Rapid incidence estimation from SARS-CoV-2 genomes reveals decreased case detection in Europe during summer 2020
The true number of infections from SARS-Cov-2 is unknown and believed to exceed the reported numbers by several fold. National testing policies, in particular, can strongly affect the proportion of undetected cases. Here, the authors propose a method that reconstructs incidence profiles within minutes, solely from publicly available, time-stamped viral genomes.
- Maureen Rebecca Smith
- , Maria Trofimova
- & Max von Kleist
-
Article
| Open Access ClusterMap for multi-scale clustering analysis of spatial gene expression
ClusterMap for multi-scale clustering analysis of spatial gene expression
In situ transcriptomics maps RNA expression patterns across intact tissues taking our understanding of gene expression to a new level. Here, the authors present a computational method that uncovers gene expression, cell niche, and tissue region patterns from 2D and 3D spatial transcriptomics.
- Yichun He
- , Xin Tang
- & Xiao Wang
-
Article
| Open Access Orchestrating and sharing large multimodal data for transparent and reproducible research
Orchestrating and sharing large multimodal data for transparent and reproducible research
It is no secret that a significant part of scientific research is difficult to reproduce. Here, the authors present a cloud-computing platform called ORCESTRA that facilitates reproducible processing of multimodal biomedical data using customizable pipelines and well-documented data objects.
- Anthony Mammoliti
- , Petr Smirnov
- & Benjamin Haibe-Kains
-
Article
| Open Access Copy number signatures predict chromothripsis and clinical outcomes in newly diagnosed multiple myeloma
Copy number signatures predict chromothripsis and clinical outcomes in newly diagnosed multiple myeloma
Chromothripsis is associated with unfavourable outcomes in multiple myeloma (MM), but its detection usually requires whole genome sequencing. Here the authors develop an approach to detect chromothripsis in MM based on copy-number signatures that also works with whole exome sequencing data.
- Kylee H. Maclachlan
- , Even H. Rustad
- & Francesco Maura
-
Matters Arising
| Open Access Quality control requirements for the correct annotation of lipidomics data
Quality control requirements for the correct annotation of lipidomics data
- Harald C. Köfeler
- , Thomas O. Eichmann
- & Kim Ekroos
-
Matters Arising
| Open Access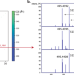 Reply to “Quality control requirements for the correct annotation of lipidomics data”
Reply to “Quality control requirements for the correct annotation of lipidomics data”
- Catherine G. Vasilopoulou
- , Karolina Sulek
- & Florian Meier
-
Article
| Open Access A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex
A genomic surveillance framework and genotyping tool for Klebsiella pneumoniae and its related species complex
Klebsiella pneumoniae is a pathogen of increasing public health concern and antimicrobial resistance is becoming more prevalent. Here, the authors describe a K. pneumoniae genotyping tool, Kleborate, that can be used to identify lineages and detect antimicrobial resistance and virulence loci.
- Margaret M. C. Lam
- , Ryan R. Wick
- & Kathryn E. Holt
-
Article
| Open Access Fractional response analysis reveals logarithmic cytokine responses in cellular populations
Fractional response analysis reveals logarithmic cytokine responses in cellular populations
Our ability to interpret single-cell multivariate signaling responses is still limited. Here the authors introduce fractional response analysis (FRA), involving fractional cell counting, capable of deconvoluting heterogeneous multivariate responses of cellular populations.
- Karol Nienałtowski
- , Rachel E. Rigby
- & Michał Komorowski
-
Article
| Open Access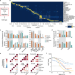 Tumor-associated hematopoietic stem and progenitor cells positively linked to glioblastoma progression
Tumor-associated hematopoietic stem and progenitor cells positively linked to glioblastoma progression
A deeper knowledge of the immune cell profile within the brain cancer tumor microenvironment (TM) could identify targets to improve immunotherapy efficacy. Here, in glioblastoma, the authors find haematopoietic stem and progenitor cells in the TM, which are associated with poor prognosis and increased immunosuppression.
- I-Na Lu
- , Celia Dobersalske
- & Igor Cima
-
Article
| Open Access Benchmarking microbiome transformations favors experimental quantitative approaches to address compositionality and sampling depth biases
Benchmarking microbiome transformations favors experimental quantitative approaches to address compositionality and sampling depth biases
Here, the authors use simulated quantitative gut microbial communities to benchmark the performance of 13 common data transformations in determining diversity as well as microbe-microbe and microbe-metadata associations, finding that quantitative approaches incorporating microbial load variation outperform computational strategies in downstream analyses, urging for a widespread adoption of quantitative approaches, or recommending specific computational transformations whenever determination of microbial load of samples is not feasible.
- Verónica Lloréns-Rico
- , Sara Vieira-Silva
- & Jeroen Raes
-
Article
| Open Access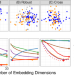 Supervised dimensionality reduction for big data
Supervised dimensionality reduction for big data
Biomedical measurements usually generate high-dimensional data where individual samples are classified in several categories. Vogelstein et al. propose a supervised dimensionality reduction method which estimates the low-dimensional data projection for classification and prediction in big datasets.
- Joshua T. Vogelstein
- , Eric W. Bridgeford
- & Mauro Maggioni
-
Article
| Open Access AutoSpill is a principled framework that simplifies the analysis of multichromatic flow cytometry data
AutoSpill is a principled framework that simplifies the analysis of multichromatic flow cytometry data
Flow cytometry allows the simultaneous quantification of many markers in and on a cell, but the analysis of such data is complicated. Here, the authors propose AutoSpill, a framework that facilitates the analysis of such data by automating parts of the analysis and requiring fewer controls.
- Carlos P. Roca
- , Oliver T. Burton
- & Adrian Liston
-
Article
| Open Access Hierarchical progressive learning of cell identities in single-cell data
Hierarchical progressive learning of cell identities in single-cell data
Classification methods for scRNA-seq data are limited in their ability to learn from multiple datasets simultaneously. Here the authors present scHPL, a hierarchical progressive learning method that automatically finds relationships between cell populations across multiple datasets and constructs a classification tree.
- Lieke Michielsen
- , Marcel J. T. Reinders
- & Ahmed Mahfouz
-
Article
| Open Access Overcoming false-positive gene-category enrichment in the analysis of spatially resolved transcriptomic brain atlas data
Overcoming false-positive gene-category enrichment in the analysis of spatially resolved transcriptomic brain atlas data
Identifying enriched gene sets in transcriptomic data is routine analysis. Here, the authors show that conventional gene category enrichment analysis (GCEA) applied to brain-wide atlas data yields biased results and develop a flexible ensemble-based null model framework to enable appropriate inference in GCEA.
- Ben D. Fulcher
- , Aurina Arnatkeviciute
- & Alex Fornito
-
Article
| Open Access Regression plane concept for analysing continuous cellular processes with machine learning
Regression plane concept for analysing continuous cellular processes with machine learning
High-content screening prompted the development of software enabling discrete phenotypic analysis of single cells. Here, the authors show that supervised continuous machine learning can drive novel discoveries in diverse imaging experiments and present the Regression Plane module of Advanced Cell Classifier.
- Abel Szkalisity
- , Filippo Piccinini
- & Peter Horvath
-
Article
| Open Access Ontology-driven weak supervision for clinical entity classification in electronic health records
Ontology-driven weak supervision for clinical entity classification in electronic health records
In the electronic health record, using clinical notes to identify entities such as disorders and their temporality can inform many important analyses. Here, the authors present a framework for weakly supervised entity classification using medical ontologies and expert-generated rules.
- Jason A. Fries
- , Ethan Steinberg
- & Nigam H. Shah
-
Article
| Open Access Automatic deep learning-driven label-free image-guided patch clamp system
Automatic deep learning-driven label-free image-guided patch clamp system
Patch clamp recording of neurons is slow and labor-intensive. Here the authors present a method for automated deep learning driven label-free image guided patch clamp physiology to perform measurements on hundreds of human and rodent neurons.
- Krisztian Koos
- , Gáspár Oláh
- & Peter Horvath
-
Article
| Open Access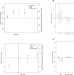 Genetic predictors of participation in optional components of UK Biobank
Genetic predictors of participation in optional components of UK Biobank
Large BioBank studies are commonly used in GWAS, but may be biased by factors affecting participation and dropout. Here the authors show that some of the factors affecting participation may have underlying genetic components.
- Jessica Tyrrell
- , Jie Zheng
- & Kate Tilling
-
Article
| Open Access Error correction enables use of Oxford Nanopore technology for reference-free transcriptome analysis
Error correction enables use of Oxford Nanopore technology for reference-free transcriptome analysis
Nanopore sequencing technologies applied to transcriptome analysis suffer from high error rates, limiting them largely to reference-based analyses. Here, the authors develop a computational error correction method for transcriptome analysis that reduces the median error rate from ~7% to ~1%.
- Kristoffer Sahlin
- & Paul Medvedev
-
Article
| Open Access Computer vision for pattern detection in chromosome contact maps
Computer vision for pattern detection in chromosome contact maps
Chromatin loops bridging distant loci within chromosomes can be detected by a variety of techniques such as Hi-C. Here the authors present Chromosight, an algorithm applied on mammalian, bacterial, viral and yeast genomes, able to detect various types of pattern in chromosome contact maps, including chromosomal loops.
- Cyril Matthey-Doret
- , Lyam Baudry
- & Axel Cournac
-
Article
| Open Access Cumulative learning enables convolutional neural network representations for small mass spectrometry data classification
Cumulative learning enables convolutional neural network representations for small mass spectrometry data classification
Convolutional Neural Networks are powerful tools for clinical diagnosis but their effectiveness decreases when the number of available samples is small. Here, the authors develop a cumulative learning method by training the same model through several classification tasks over various small Mass Spectrometry datasets.
- Khawla Seddiki
- , Philippe Saudemont
- & Arnaud Droit
-
Article
| Open Access Improved haplotype inference by exploiting long-range linking and allelic imbalance in RNA-seq datasets
Improved haplotype inference by exploiting long-range linking and allelic imbalance in RNA-seq datasets
Haplotype reconstruction of distant genetic variants is problematic in short-read sequencing. Here, the authors describe HapTree-X, a probabilistic framework that uses differential allele-specific expression to better reconstruct paternal haplotypes from diploid and polyploid genomes.
- Emily Berger
- , Deniz Yorukoglu
- & Bonnie Berger
-
Article
| Open Access Deep neural networks enable quantitative movement analysis using single-camera videos
Deep neural networks enable quantitative movement analysis using single-camera videos
In the context of diseases impairing movement, quantitative assessment of motion is critical to medical decision-making but is currently possible only with expensive motion capture systems and trained personnel. Here, the authors present a method for predicting clinically relevant motion parameters from an ordinary video of a patient.
- Łukasz Kidziński
- , Bryan Yang
- & Michael H. Schwartz
-
Article
| Open Access Strategies to enable large-scale proteomics for reproducible research
Strategies to enable large-scale proteomics for reproducible research
Clinical proteomics critically depends on the ability to acquire highly reproducible data over an extended period of time. Here, the authors assess reproducibility over four months across different mass spectrometers and develop a computational approach to mitigate variation among instruments over time.
- Rebecca C. Poulos
- , Peter G. Hains
- & Qing Zhong
-
Perspective
| Open Access Causality matters in medical imaging
Causality matters in medical imaging
Scarcity of high-quality annotated data and mismatch between the development dataset and the target environment are two of the main challenges in developing predictive tools from medical imaging. In this Perspective, the authors show how causal reasoning can shed new light on these challenges.
- Daniel C. Castro
- , Ian Walker
- & Ben Glocker
-
Article
| Open Access Deep learning for genomics using Janggu
Deep learning for genomics using Janggu
Deep learning is becoming a popular approach for understanding biological processes but can be hard to adapt to new questions. Here, the authors develop Janggu, a python library that aims to ease data acquisition and model evaluation and facilitate deep learning applications in genomics.
- Wolfgang Kopp
- , Remo Monti
- & Altuna Akalin
-
Article
| Open Access Focus on the spectra that matter by clustering of quantification data in shotgun proteomics
Focus on the spectra that matter by clustering of quantification data in shotgun proteomics
Matching mass spectra to peptide sequences is the usual first step in proteomics data analysis, often followed by peptide quantification. Here, the authors show that clustering and quantifying mass spectral features prior to peptide identification can increase the sensitivity of label-free quantitative proteomics.
- Matthew The
- & Lukas Käll
-
Article
| Open Access Discovery and quality analysis of a comprehensive set of structural variants and short tandem repeats
Discovery and quality analysis of a comprehensive set of structural variants and short tandem repeats
The complexity of structural variation (SV) and short tandem repeats (STRs) makes it necessary to apply different calling and filtering strategies to sequencing datasets. Here, Jakubosky et al. report a comprehensive SV and STR callset from whole-genome sequencing of 477 individuals from iPSCORE and HipSci using five algorithms.
- David Jakubosky
- , Erin N. Smith
- & Kelly A. Frazer
-
Article
| Open Access Deep learning enables structured illumination microscopy with low light levels and enhanced speed
Deep learning enables structured illumination microscopy with low light levels and enhanced speed
Super-resolution microscopy typically requires high laser powers which can induce photobleaching and degrade image quality. Here the authors augment structured illumination microscopy (SIM) with deep learning to reduce the number of raw images required and boost its performance under low light conditions.
- Luhong Jin
- , Bei Liu
- & Klaus M. Hahn
-
Article
| Open Access Colonic microbiota is associated with inflammation and host epigenomic alterations in inflammatory bowel disease
Colonic microbiota is associated with inflammation and host epigenomic alterations in inflammatory bowel disease
Inflammatory bowel disease (IBD) has been linked to host-microbiota interactions. Here, the authors investigate mucosa-associated microbiota using endoscopically-targeted biopsies from inflamed and non-inflamed colon in patients with Crohn’s disease and ulcerative colitis, finding associations with inflammation and host epigenomic alterations.
- F. J. Ryan
- , A. M. Ahern
- & M. J. Claesson
-
Article
| Open Access Graph embedding and unsupervised learning predict genomic sub-compartments from HiC chromatin interaction data
Graph embedding and unsupervised learning predict genomic sub-compartments from HiC chromatin interaction data
Accurate identification of sub-compartments from chromatin interaction data remains a challenge. Here, the authors introduce an algorithm combining graph embedding and unsupervised learning to predict sub-compartments using Hi-C data.
- Haitham Ashoor
- , Xiaowen Chen
- & Sheng Li
-
Article
| Open Access Abundance and diversity of resistomes differ between healthy human oral cavities and gut
Abundance and diversity of resistomes differ between healthy human oral cavities and gut
Antimicrobial resistance (AMR) represents a global health threat. Here, the authors analyse the oral and gut resistomes from metagenomes of diverse populations and find that the oral resistome harbours higher abundance but lower diversity of antimicrobial resistance genes than the gut resistome.
- Victoria R. Carr
- , Elizabeth A. Witherden
- & David L. Moyes
-
Article
| Open Access Agreement between two large pan-cancer CRISPR-Cas9 gene dependency data sets
Agreement between two large pan-cancer CRISPR-Cas9 gene dependency data sets
Integrating independent large-scale pharmacogenomic screens can enable unprecedented characterization of genetic vulnerabilities in cancers. Here, the authors show that the two largest independent CRISPR-Cas9 gene-dependency screens are concordant, paving the way for joint analysis of the data sets.
- Joshua M. Dempster
- , Clare Pacini
- & Francesco Iorio
-
Article
| Open Access The SIGMA rat brain templates and atlases for multimodal MRI data analysis and visualization
The SIGMA rat brain templates and atlases for multimodal MRI data analysis and visualization
Magnetic resonance imaging (MRI) is widely used to study the rat brain. Here, the authors provide standardized MRI brain templates and descriptive atlases for the rat, incorporating both structural and functional MRI data, along with associated resources.
- D. A. Barrière
- , R. Magalhães
- & S. Mériaux
-
Article
| Open Access The Escherichia coli transcriptome mostly consists of independently regulated modules
The Escherichia coli transcriptome mostly consists of independently regulated modules
Mechanistic insight into the regulation of transcriptional modules remains scarce. Here, the authors identify statistically independent gene sets by applying independent component analysis to a high-quality E. coli RNA-seq data compendium and find that most gene sets represent the effects of specific transcriptional regulators.
- Anand V. Sastry
- , Ye Gao
- & Bernhard O. Palsson

 Metabolite annotation from knowns to unknowns through knowledge-guided multi-layer metabolic networking
Metabolite annotation from knowns to unknowns through knowledge-guided multi-layer metabolic networking