Article
|
Open Access
Featured
-
-
Article
| Open Access SHIELD: a platform for high-throughput screening of barrier-type DNA elements in human cells
SHIELD: a platform for high-throughput screening of barrier-type DNA elements in human cells
Chromatin boundary elements are hard to define and characterize. Here the authors report Site-specific Heterochromatin Insertion of Elements at Lamina-associated Domains (SHIELD) for high-throughput screening of barrier-type DNA elements in human cells.
- Meng Zhang
- , Mary Elisabeth Ehmann
- & Huimin Zhao
-
Article
| Open Access HMGN1 enhances CRISPR-directed dual-function A-to-G and C-to-G base editing
HMGN1 enhances CRISPR-directed dual-function A-to-G and C-to-G base editing
Limited work has been done on concurrent C-to-G and A-to-G base editing. Here the authors test how a number of chromatin-associated factors affect base editing and show that HMGN1 enhanced the efficiency; by fusing HMGN1 to GBE and ABE they develop a CRISPR-based dual-function A-to-G and C-to-G base editor (GGBE).
- Chao Yang
- , Zhenzhen Ma
- & Xueli Zhang
-
Article
| Open Access Low input capture Hi-C (liCHi-C) identifies promoter-enhancer interactions at high-resolution
Low input capture Hi-C (liCHi-C) identifies promoter-enhancer interactions at high-resolution
Here the authors present the low input capture Hi-C (liCHi-C) method, a cost-effective, flexible method to map and robustly compare promoter interactomes at high resolution. liCHi-C identifies new disease-associated genes and structural variants to ultimately illuminate their pathogenic effects.
- Laureano Tomás-Daza
- , Llorenç Rovirosa
- & Biola M. Javierre
-
Article
| Open Access Hi-TrAC reveals division of labor of transcription factors in organizing chromatin loops
Hi-TrAC reveals division of labor of transcription factors in organizing chromatin loops
It is currently not clear how architectural proteins orchestrate chromatin looping at different scales of genome organisation. Here the authors report Hi-TrAC as a proximity ligation-free method to profile genome-wide chromatin interactions at single nucleosome resolution among regulatory elements.
- Shuai Liu
- , Yaqiang Cao
- & Keji Zhao
-
Article
| Open Access Multidimensional chromatin profiling of zebrafish pancreas to uncover and investigate disease-relevant enhancers
Multidimensional chromatin profiling of zebrafish pancreas to uncover and investigate disease-relevant enhancers
Alterations in cis-regulatory elements (CREs) can contribute to pancreatic diseases. Here the authors combine chromatin profiling and interaction points with in vivo reporter assays in zebrafish to uncover functionally equivalent human CREs, helping to predict disease-relevant enhancers.
- Renata Bordeira-Carriço
- , Joana Teixeira
- & José Bessa
-
Article
| Open Access A fast Myosin super enhancer dictates muscle fiber phenotype through competitive interactions with Myosin genes
A fast Myosin super enhancer dictates muscle fiber phenotype through competitive interactions with Myosin genes
The contractile properties of adult myofibers are shaped by their Myosin heavy chain isoform content. Here the authors show that a super enhancer controls the spatiotemporal expression of the genes at the fast myosin heavy chain locus by DNA looping and that this expression profile is recapitulated in a rainbow transgenic mouse model of the locus.
- Matthieu Dos Santos
- , Stéphanie Backer
- & Pascal Maire
-
Article
| Open Access Dynamic transcriptome and chromatin architecture in granulosa cells during chicken folliculogenesis
Dynamic transcriptome and chromatin architecture in granulosa cells during chicken folliculogenesis
The domestic chicken Gallus gallus domesticus is a classic model for the study of folliculogenesis. Here the authors integrate multi-omics analyses characterizing the dynamic transcriptome and chromatin architecture in granulosa cells during chicken folliculogenesis.
- Diyan Li
- , Chunyou Ning
- & Mingzhou Li
-
Article
| Open Access A compendium of chromatin contact maps reflecting regulation by chromatin remodelers in budding yeast
A compendium of chromatin contact maps reflecting regulation by chromatin remodelers in budding yeast
The effect of ATP-dependent chromatin remodelers on 3D genome organization has not been well studied. Here the authors employ in situ Hi-C with an auxin-inducible degron system to degrade chromatin remodelers in yeast to find that the 3D structure of chromatin collapses in their absence. The chromatin remodeling can modulate 3D architecture depending on chromosomal context and cell cycle stage.
- Hyelim Jo
- , Taemook Kim
- & Daeyoup Lee
-
Article
| Open Access EpiScanpy: integrated single-cell epigenomic analysis
EpiScanpy: integrated single-cell epigenomic analysis
The authors present epiScanpy: a computational framework for the analysis of single-cell epigenomic data, both ATAC-seq and DNA methylation data, with examples for clustering, cell type identification, trajectory learning and atlas integration - and show its performance in distinguishing cell types.
- Anna Danese
- , Maria L. Richter
- & Maria Colomé-Tatché
-
Article
| Open Access Mesomelic dysplasias associated with the HOXD locus are caused by regulatory reallocations
Mesomelic dysplasias associated with the HOXD locus are caused by regulatory reallocations
Mesomelic dysplasia, a severe shortening and bending of the limb, has been linked to rearrangements in the HoxD cluster in humans and mice. Here the authors engineer a 1 Mb inversion including the HoxD gene cluster and use this model to provide a mechanistic framework to understand and unify the molecular origins of human mesomelic dysplasia associated with 2q31.
- Christopher Chase Bolt
- , Lucille Lopez-Delisle
- & Denis Duboule
-
Article
| Open Access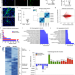 HIRA stabilizes skeletal muscle lineage identity
HIRA stabilizes skeletal muscle lineage identity
The epigenetic mechanisms coordinating the maintenance of adult cellular lineages remain poorly understood. Here the authors demonstrate that HIRA, a H3.3 histone chaperone, establishes the chromatin landscape required for skeletal muscle cell identity.
- Joana Esteves de Lima
- , Reem Bou Akar
- & Frédéric Relaix
-
Article
| Open Access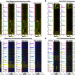 Multiple roles of H2A.Z in regulating promoter chromatin architecture in human cells
Multiple roles of H2A.Z in regulating promoter chromatin architecture in human cells
Histone variant H2A.Z has been suggested to contribute to the regulation of promoter accessibility. Here, the authors present high-depth maps of the position and accessibility of H2A.Z-containing nucleosomes for human Pol II promoters and provide evidence that H2A.Z has multiple and distinct roles in regulating gene expression dependent upon its location in a promoter.
- Lauren Cole
- , Sebastian Kurscheid
- & David J. Tremethick
-
Article
| Open Access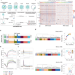 Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Single-cell multiomics sequencing reveals the functional regulatory landscape of early embryos
Extensive epigenetic reprogramming occurs during preimplantation embryo development. Here the authors develop a single cell multiomics sequencing technology that enables profiling of genome-wide chromatin accessibility, DNA methylation and RNA expression in the same individual cell and apply this method to study mouse preimplantation embryos.
- Yang Wang
- , Peng Yuan
- & Liying Yan
-
Article
| Open Access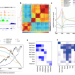 Non-CG methylation and multiple histone profiles associate child abuse with immune and small GTPase dysregulation
Non-CG methylation and multiple histone profiles associate child abuse with immune and small GTPase dysregulation
Early-life adversity is thought to increase the risk of psychopathology through epigenetic mechanisms. Here, the authors profile 6 histone marks, chromatin states and DNA methylation in the lateral amygdala in subjects with a history of early-life adversity.
- Pierre-Eric Lutz
- , Marc-Aurèle Chay
- & Gustavo Turecki
-
Article
| Open Access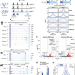 Conserved regulatory logic at accessible and inaccessible chromatin during the acute inflammatory response in mammals
Conserved regulatory logic at accessible and inaccessible chromatin during the acute inflammatory response in mammals
Genetic elements that control inflammatory gene expression are not fully elucidated. Here the authors conduct a multi-species analysis of chromatin landscape and NF-κB binding in response to the proinflammatory cytokine TNFα, finding that conserved NF-κB bound regions are linked to enhancer activity and disease.
- Azad Alizada
- , Nadiya Khyzha
- & Michael D. Wilson
-
Article
| Open Access GATA2 regulates mast cell identity and responsiveness to antigenic stimulation by promoting chromatin remodeling at super-enhancers
GATA2 regulates mast cell identity and responsiveness to antigenic stimulation by promoting chromatin remodeling at super-enhancers
Mast cells are critical effectors of allergic inflammation and protection against parasitic infections. Here the authors demonstrate that GATA2 promotes chromatin accessibility at the super-enhancers of mast cell identity genes and primes both typical and super-enhancers at genes that respond to antigenic stimulation.
- Yapeng Li
- , Junfeng Gao
- & Hua Huang
-
Article
| Open Access BET inhibition disrupts transcription but retains enhancer-promoter contact
BET inhibition disrupts transcription but retains enhancer-promoter contact
The role of BRD4 and Mediator in regulating enhancer-promoter interactions is poorly understood. Here the authors find that treatment with BET inhibitors or pharmacological degradation of BRD4 disrupts transcription while having very little effect on enhancer-promoter interactions.
- Nicholas T. Crump
- , Erica Ballabio
- & Thomas A. Milne
-
Article
| Open Access Order and stochasticity in the folding of individual Drosophila genomes
Order and stochasticity in the folding of individual Drosophila genomes
Genomes are partitioned into topologically associating domains (TADs). Here the authors present single-nucleus Hi-C maps in Drosophila at 10 kb resolution, demonstrating the presence of chromatin compartments in individual nuclei, and partitioning of the genome into non-hierarchical TADs at the scale of 100 kb, which resembles population TAD profiles.
- Sergey V. Ulianov
- , Vlada V. Zakharova
- & Sergey V. Razin
-
Article
| Open Access Primary effusion lymphoma enhancer connectome links super-enhancers to dependency factors
Primary effusion lymphoma enhancer connectome links super-enhancers to dependency factors
Primary effusion lymphoma (PEL) has a very poor prognosis. Here, the authors perform H3K27ac HiChIP in PEL cells and generate the PEL enhancer connectome, linking enhancers and promoters in PEL, as well as super-enhancers to dependency factors.
- Chong Wang
- , Luyao Zhang
- & Bo Zhao
-
Article
| Open Access SAMMY-seq reveals early alteration of heterochromatin and deregulation of bivalent genes in Hutchinson-Gilford Progeria Syndrome
SAMMY-seq reveals early alteration of heterochromatin and deregulation of bivalent genes in Hutchinson-Gilford Progeria Syndrome
Hutchinson-Gilford progeria syndrome is a genetic disease where an aberrant form of Lamin A disrupts chromatin by interfering with lamina associated domains. Here, the authors present the SAMMY-seq, a method for genome-wide characterization of heterochromatin dynamics and detect early stage alterations of heterochromatin structure in progeria primary fibroblasts, accompained by Polycomb dysfunctions.
- Endre Sebestyén
- , Fabrizia Marullo
- & Chiara Lanzuolo
-
Article
| Open Access Transcription-dependent cohesin repositioning rewires chromatin loops in cellular senescence
Transcription-dependent cohesin repositioning rewires chromatin loops in cellular senescence
Senescence is a state of stable proliferative arrest. Here, the authors perform Hi-C analysis on oncogenic RAS-induced senescence in human fibroblasts and characterize the changes in the 3D genome folding associated with the senescence-specific gene expression profile, which are mediated in part through cohesin redistribution on chromatin.
- Ioana Olan
- , Aled J. Parry
- & Masashi Narita
-
Article
| Open Access An embryonic stem cell-specific heterochromatin state promotes core histone exchange in the absence of DNA accessibility
An embryonic stem cell-specific heterochromatin state promotes core histone exchange in the absence of DNA accessibility
Nucleosome turnover concomitant with incorporation of the histone variant H3.3 is a hallmark of regulatory regions in the animal genome. Here, the authors demonstrate that fast histone turnover and H3.3 incorporation defines a dynamic heterochromatin state in pluripotent stem cells.
- Carmen Navarro
- , Jing Lyu
- & Simon J. Elsässer
-
Article
| Open Access Mapping effector genes at lupus GWAS loci using promoter Capture-C in follicular helper T cells
Mapping effector genes at lupus GWAS loci using promoter Capture-C in follicular helper T cells
T cells are a major cell type involved in systemic lupus erythematosus (SLE). Here, the authors use promoter capture-C and ATAC-seq in human follicular T helper cells to identify SLE genes distant from GWAS loci (via 3D interaction) and validate the function of key regulatory elements and genes in vitro.
- Chun Su
- , Matthew E. Johnson
- & Andrew D. Wells
-
Article
| Open Access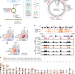 Epigenetic specifications of host chromosome docking sites for latent Epstein-Barr virus
Epigenetic specifications of host chromosome docking sites for latent Epstein-Barr virus
Epstein-Barr virus (EBV) episomes tether to the host chromosome via EBNA1. Here, using circular chromosome conformation capture (4C), Kim et al. identify attachment sites and show that EBV episomes preferentially associate with transcriptionally silenced genes in Burkitt lymphoma cells.
- Kyoung-Dong Kim
- , Hideki Tanizawa
- & Paul M. Lieberman
-
Article
| Open Access Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing
Interrogation of enhancer function by enhancer-targeting CRISPR epigenetic editing
Tissues-specific gene expression requires coordinated cis-regulatory elements. Here the authors use dCas9-based enhancer targeting to remodel local epigenetic landscapes and activate or inactive transcription.
- Kailong Li
- , Yuxuan Liu
- & Jian Xu
-
Article
| Open Access CUT&Tag for efficient epigenomic profiling of small samples and single cells
CUT&Tag for efficient epigenomic profiling of small samples and single cells
Understanding gene regulation will require mapping specific chromain features in a small number of cells at high resolution. Here the authors describe CUT&Tag, which uses antibody-mediated tethering of Tn5 transposase to a chromatin protein to generate high resolution libraries.
- Hatice S. Kaya-Okur
- , Steven J. Wu
- & Steven Henikoff
-
Article
| Open Access Deconvolution of single-cell multi-omics layers reveals regulatory heterogeneity
Deconvolution of single-cell multi-omics layers reveals regulatory heterogeneity
Heterogeneity in gene expression and epigenetic states exists across individual cells. Here, the authors develop scCAT-seq, a technique for simultaneously performing ATAC-seq and RNA-seq within the same single cell.
- Longqi Liu
- , Chuanyu Liu
- & Xun Xu
-
Article
| Open Access A rapid and robust method for single cell chromatin accessibility profiling
A rapid and robust method for single cell chromatin accessibility profiling
ATAC-seq is widely used to identify regulatory regions in the genome. Here the authors develop a simple and robust plate-based single-cell ATAC-seq method that works in fresh and cryopreserved cells.
- Xi Chen
- , Ricardo J. Miragaia
- & Sarah A. Teichmann
-
Article
| Open Access Chromatin conformation analysis of primary patient tissue using a low input Hi-C method
Chromatin conformation analysis of primary patient tissue using a low input Hi-C method
Chromatin conformation studies are limited by the large amounts of starting material required to perform current protocols. Here the authors present Low-C, a Hi-C method for low amounts of input material and produce Low-C maps from primary B-cells of a diffuse large B-cell lymphoma patient, demonstrating the suitability of Low-C to analyse rare cell populations.
- Noelia Díaz
- , Kai Kruse
- & Juan M. Vaquerizas
-
Article
| Open Access Joint single-cell DNA accessibility and protein epitope profiling reveals environmental regulation of epigenomic heterogeneity
Joint single-cell DNA accessibility and protein epitope profiling reveals environmental regulation of epigenomic heterogeneity
Cellular heterogeneity in cancer is complex and difficult to study. Here, the authors introduce Protein-indexed Assay of Transposase Accessible Chromatin (Pi-ATAC), which combines single cell chromatin and proteomic profiling to provide deep insight into the tumor microenvironment, and reveal the role of hypoxia in shaping the regulome of a subset of breast cancer cells in vivo.
- Xingqi Chen
- , Ulrike M. Litzenburger
- & Howard Y. Chang
-
Article
| Open Access Cell cycle-resolved chromatin proteomics reveals the extent of mitotic preservation of the genomic regulatory landscape
Cell cycle-resolved chromatin proteomics reveals the extent of mitotic preservation of the genomic regulatory landscape
Mitosis poses a challenge for transcriptional programs, as it is thought that several proteins lose binding on condensed chromosomes. Here, the authors analyze the chromatin-bound proteome through the cell cycle, revealing retention of most transcription factors and preservation of the regulatory landscape.
- Paul Adrian Ginno
- , Lukas Burger
- & Dirk Schübeler
-
Article
| Open Access scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
scNMT-seq enables joint profiling of chromatin accessibility DNA methylation and transcription in single cells
Relationships between DNA methylation and transcription, and methylation and DNA accessibility can be probed but interrogating all three in the same single cells has not been possible. Here, the authors report the first single-cell method for parallel chromatin accessibility, DNA methylation and transcriptome profiling.
- Stephen J. Clark
- , Ricard Argelaguet
- & Wolf Reik
-
Article
| Open Access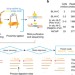 BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions
BL-Hi-C is an efficient and sensitive approach for capturing structural and regulatory chromatin interactions
Chromatin interactions and genome architecture are key regulators of gene expression. Here the authors present Bridge-Linker-Hi-C to map active chromatin loops and enhancer-promoter interactions.
- Zhengyu Liang
- , Guipeng Li
- & Yang Chen
-
Article
| Open Access Rapid and reversible epigenome editing by endogenous chromatin regulators
Rapid and reversible epigenome editing by endogenous chromatin regulators
Understanding the link between epigenetic marks and gene regulation requires the development of new tools to directly manipulate chromatin. Here the authors demonstrate a Cas9-based system to recruit chromatin remodelers to loci of interest, allowing rapid, reversible manipulation of epigenetic states.
- Simon M. G. Braun
- , Jacob G. Kirkland
- & Gerald R. Crabtree
-
Article
| Open Access Capture of associated targets on chromatin links long-distance chromatin looping to transcriptional coordination
Capture of associated targets on chromatin links long-distance chromatin looping to transcriptional coordination
Chromatin architecture is a key regulator of transcriptional processes, however current methods to investigate it have technical limitations. Here, the authors describe a novel chromatin capture technique, CATCH, which can be used to identify and characterize complex genomic interaction networks.
- Ryan J. Bourgo
- , Hari Singhal
- & Geoffrey L. Greene
-
Article
| Open Access Leukaemia cell of origin identified by chromatin landscape of bulk tumour cells
Leukaemia cell of origin identified by chromatin landscape of bulk tumour cells
A tumour’s cell of origin may influence tumour progression and response to therapy. Here, the authors demonstrate that the cell of origin determines the aggressiveness of AML in a mouse model and identify unique biomarkers of the specific leukaemia cell of origin by profiling open chromatin regions of AML samples.
- Joshy George
- , Asli Uyar
- & Jennifer J. Trowbridge
-
Article
| Open Access Nucleation and spreading of a heterochromatic domain in fission yeast
Nucleation and spreading of a heterochromatic domain in fission yeast
Chromosomes contain large heterochromatin domains. Here, the authors measure the kinetics of heterochromatin formation in fission yeast and show both global and local feedbacks by nucleosome-bound enzymes are important for formation and stability of the large heterochromatin domains.
- Michaela J. Obersriebnig
- , Emil M. H. Pallesen
- & Geneviève Thon
-
Article
| Open Access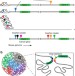 3D hotspots of recurrent retroviral insertions reveal long-range interactions with cancer genes
3D hotspots of recurrent retroviral insertions reveal long-range interactions with cancer genes
Retroviral insertional mutagenesis is used for identifying genes involved in the development of cancer. Here, the authors overlay cancer-causing insertions with genome-wide Hi-C data and find that retroviral elements tend to cluster in 3D hotspots.
- Sepideh Babaei
- , Waseem Akhtar
- & Jeroen de Ridder
-
Article
| Open Access Epigenomic footprints across 111 reference epigenomes reveal tissue-specific epigenetic regulation of lincRNAs
Epigenomic footprints across 111 reference epigenomes reveal tissue-specific epigenetic regulation of lincRNAs
Tissue-specific functions have been established for some lincRNAs. Here, by analysing 111 reference epigenomes from the NIH Roadmap Epigenomics project, the authors report tissue-specific epigenomic regulation of 3,753 lincRNAs and their strong connection with tissue-specific pathways.
- Viren Amin
- , R. Alan Harris
- & Aleksandar Milosavljevic
-
Article |
 Nanoscale chromatin profiling of gastric adenocarcinoma reveals cancer-associated cryptic promoters and somatically acquired regulatory elements
Nanoscale chromatin profiling of gastric adenocarcinoma reveals cancer-associated cryptic promoters and somatically acquired regulatory elements
Epigenetic alterations alter chromatin structure and gene expression and are known contributors to cancer development. Here, Muratani et al.profile multiple epigenetic chromatin marks in primary gastric cancers and identify hundreds of altered promoters and enhancers that drive the gene expression program in these malignancies.
- Masafumi Muratani
- , Niantao Deng
- & Patrick Tan

 Epigenomic analysis of formalin-fixed paraffin-embedded samples by CUT&Tag
Epigenomic analysis of formalin-fixed paraffin-embedded samples by CUT&Tag