Abstract
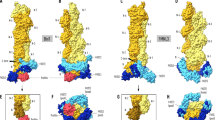
Formins are actin-assembly factors that act in a variety of actin-based processes. The conserved formin homology 2 (FH2) domain promotes filament nucleation and influences elongation through interaction with the barbed end. FMNL3 is a formin that induces assembly of filopodia but whose FH2 domain is a poor nucleator. The 3.4-Å structure of a mouse FMNL3 FH2 dimer in complex with tetramethylrhodamine-actin uncovers details of formin-regulated actin elongation. We observe distinct FH2 actin-binding regions; interactions in the knob and coiled-coil subdomains are necessary for actin binding, whereas those in the lasso-post interface are important for the stepping mechanism. Biochemical and cellular experiments test the importance of individual residues for function. This structure provides details for FH2-mediated filament elongation by processive capping and supports a model in which C-terminal non-FH2 residues of FMNL3 are required to stabilize the filament nucleus.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Pollard, T.D. & Cooper, J. Actin, a central player in cell shape and movement. Science 326, 1208–1212 (2009).
Chang, F., Drubin, D. & Nurse, P. Cdc12P, a protein required for cytokinesis in fission yeast, is a component of the cell division ring and interacts with profilin. J. Cell Biol. 137, 169–182 (1997).
Goode, B.L. & Eck, M.J. Mechanism and function of formins in the control of actin assembly. Annu. Rev. Biochem. 76, 593–627 (2007).
Xu, Y. et al. Crystal structures of a formin homology-2 domain reveal a tethered dimer architecture. Cell 116, 711–723 (2004).
Otomo, T. et al. Structural basis of actin filament nucleation and processive capping by a formin homology 2 domain. Nature 433, 488–494 (2005).
Shimada, A. et al. The core FH2 domain of diaphanous-related formins is an elongated actin binding protein that inhibits polymerization. Mol. Cell 13, 511–522 (2004).
Nezami, A., Poy, F., Toms, A., Zheng, W. & Eck, M.J. Crystal structure of a complex between amino and carboxy terminal fragments of mDia1: insights into autoinhibition of diaphanous-related formins. PLoS One 5, 1–11 (2010).
Lu, J. et al. Structure of the FH2 domain of Daam1: implications for formin regulation of actin assembly. J. Mol. Biol. 369, 1258–1269 (2007).
Yamashita, M. et al. Crystal structure of human DAAM1 formin homology 2 domain. Genes Cells 12, 1255–1265 (2007).
Paul, A.S. & Pollard, T.D. Review of the mechanism of processive actin filament elongation by formins. Cell Motil. Cytoskeleton 66, 606–617 (2009).
Shemesh, T., Otomo, T., Rosen, M.K., Bershadsky, A.D. & Kozlov, M.M. A novel mechanism of actin filament processive capping by formin: solution of the rotation paradox. J. Cell Biol. 170, 889–893 (2005).
Mizuno, H. et al. Rotational movement of the formin mDia1 along the double helical strand of an actin filament. Science 331, 80–83 (2011).
Kovar, D.R., Harris, E.S., Mahaffy, R., Higgs, H.N. & Pollard, T.D. Control of the assembly of ATP- and ADP-actin by formins and profilin. Cell 124, 423–435 (2006).
Harris, E.S., Rouiller, I., Hanein, D. & Higgs, H.N. Mechanistic differences in actin bundling activity of two mammalian formins, FRL1 and mDia2. J. Biol. Chem. 281, 14383–14392 (2006).
Moseley, J.B. & Goode, B.L. Differential activities and regulation of Saccharomyces cerevisiae formin proteins Bni1 and Bnr1 by Bud6. J. Biol. Chem. 280, 28023–28033 (2005).
Gould, C.J. et al. The formin DAD domain plays dual roles in autoinhibition and actin nucleation. Curr. Biol. 21, 384–390 (2011).
Heimsath, E.G. & Higgs, H.N. The C terminus of formin FMNL3 accelerates actin polymerization and contains a WH2 domain-like sequence that binds both monomers and filament barbed ends. J. Biol. Chem. 287, 3087–3098 (2012).
Vizcarra, C.L. et al. Structure and function of the interacting domains of Spire and Fmn-family formins. Proc. Natl. Acad. Sci. USA 108, 11884–11889 (2011).
Moseley, J., Sagot, I. & Manning, A. A conserved mechanism for Bni1-and mDia1-induced actin assembly and dual regulation of Bni1 by Bud6 and profilin. Mol. Biol. 15, 896–907 (2004).
Graceffa, P. & Dominguez, R. Crystal structure of monomeric actin in the ATP state. Structural basis of nucleotide-dependent actin dynamics. J. Biol. Chem. 278, 34172–34180 (2003).
Laskowski, R.A. PDBsum new things. Nucleic Acids Res. 37, D355–D359 (2009).
Harris, E.S., Gauvin, T.J., Heimsath, E.G. & Higgs, H.N. Assembly of filopodia by the formin FRL2 (FMNL3). Cytoskeleton 67, 755–772 (2010).
Scott, B.J., Neidt, E.M. & Kovar, D.R. The functionally distinct fission yeast formins have specific actin-assembly properties. Mol. Biol. Cell 22, 3826–3839 (2011).
Paul, A.S. & Pollard, T.D. The role of the FH1 domain and profilin in formin-mediated actin-filament elongation and nucleation. Curr. Biol. 18, 9–19 (2008).
Paul, A.S. & Pollard, T.D. Energetic requirements for processive elongation of actin filaments by FH1FH2-formins. J. Biol. Chem. 284, 12533–12540 (2009).
Galkin, V.E., Orlova, A., Kudryashov, D.S., Solodukhin, A. & Reisler, E. Remodeling of actin filaments by ADF/cofilin proteins. Proc. Natl. Acad. Sci. USA 108, 20568–20572 (2011).
Galkin, V.E., Orlova, A., Schröder, G.F. & Egelman, E.H. Structural polymorphism in F-actin. Nat. Struct. Mol. Biol. 17, 1318–1323 (2010).
Holmes, K.C., Angert, I., Kull, F.J. & Jahn, W. Electron cryo-microscopy shows how strong binding of myosin to actin releases nucleotide. Nature 425, 423–427 (2003).
Chereau, D. et al. Actin-bound structures of Wiskott-Aldrich syndrome protein (WASP)-homology domain 2 and the implications for filament assembly. Proc. Natl. Acad. Sci. USA 102, 16644–16649 (2005).
Graziano, B.R. et al. Mechanism and cellular function of Bud6 as an actin nucleation-promoting factor. Mol. Biol. Cell 22, 4016–4028 (2011).
Breitsprecher, D. et al. Rocket launcher mechanism of collaborative actin assembly defined by single-molecule imaging. Science 336, 1164–1168 (2012).
Kapust, R.B. et al. Tobacco etch virus protease: mechanism of autolysis and rational design of stable mutants with wild-type catalytic proficiency. Protein Eng. 14, 993–1000 (2001).
Harris, E.S., Li, F. & Higgs, H.N. The mouse formin, FRLalpha, slows actin filament barbed end elongation, competes with capping protein, accelerates polymerization from monomers, and severs filaments. J. Biol. Chem. 279, 20076–20087 (2004).
Spudich, J.A. & Watt, S. The regulation of rabbit skeletal muscle contraction. I. Biochemical studies of the interaction of the tropomyosin-troponin complex with actin and the proteolytic fragments of myosin. J. Biol. Chem. 246, 4866–4871 (1971).
Kabsch, W. Automatic processing of rotation diffraction data from crystals of initially unknown symmetry and cell constants. J. Appl. Crystallogr. 26, 795–800 (1993).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Kelley, L.A. & Sternberg, M.J.E. Protein structure prediction on the Web: a case study using the Phyre server. Nat. Protoc. 4, 363–371 (2009).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Afonine, P.V. et al. Joint X-ray and neutron refinement with phenix.refine. Acta Crystallogr. D Biol. Crystallogr. 66, 1153–1163 (2010).
Schuck, P. Size-distribution analysis of macromolecules by sedimentation velocity ultracentrifugation and lamm equation modeling. Biophys. J. 78, 1606–1619 (2000).
Kuhn, J.R. & Pollard, T.D. Real-time measurements of actin filament polymerization by total internal reflection fluorescence microscopy. Biophys. J. 88, 1387–1402 (2005).
Acknowledgements
We thank our undergraduates A. Kelley for mutagenesis and cloning and M. Lee for crystallization trials and screening; J. Moseley for thoughtful comments and careful reading of the manuscript; and S. Corcoran and the staff at the GM/CA CAT 23-ID-D beamline at Argonne National Laboratories Advanced Photon Source (APS) Source; and C. Bahl and D. Madden for help with the structural determination and characterization. The human α1β2 expression vector was a gift from J. Cooper (Washington University, St. Louis, Missouri, USA). This work was supported by US National Institutes of Health grants R01 GM069818 (to H.N.H.) and F31 GM089149 (to E.G.H.) as well as Howard Hughes Medical Institute predoctoral fellowship 52006921 and National Science Foundation GK-12 fellowship 0947790 (to M.E.T.). GM/CA at APS has been funded in whole or in part with US federal funds from the National Cancer Institute (Y1-CO-1020) and the National Institute of General Medical Sciences (Y1-GM-1104). Use of the APS was supported by the US Department of Energy, Basic Energy Sciences, Office of Science (contract DE-AC02-06CH11357).
Author information
Authors and Affiliations
Contributions
M.E.T. purified, crystallized and characterized the FH2–actin complex and also collected data on the crystals and determined the structure. E.G.H. performed the pyrene-actin assays. T.J.G. carried out the cellular characterization and analysis of the mutant protein constructs. The experimental design and data analysis was carried out by M.E.T., H.N.H. and F.J.K. The manuscript was prepared by M.E.T., H.N.H. and F.J.K.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9 and Supplementary Note (PDF 8519 kb)
Rights and permissions
About this article
Cite this article
Thompson, M., Heimsath, E., Gauvin, T. et al. FMNL3 FH2–actin structure gives insight into formin-mediated actin nucleation and elongation. Nat Struct Mol Biol 20, 111–118 (2013). https://doi.org/10.1038/nsmb.2462
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2462
This article is cited by
-
A structural model of the profilin–formin pacemaker system for actin filament elongation
Scientific Reports (2022)
-
Biochemical and mechanical regulation of actin dynamics
Nature Reviews Molecular Cell Biology (2022)
-
Alterations to the broad-spectrum formin inhibitor SMIFH2 modulate potency but not specificity
Scientific Reports (2022)
-
Inhibition of polar actin assembly by astral microtubules is required for cytokinesis
Nature Communications (2021)
-
Forces generated by lamellipodial actin filament elongation regulate the WAVE complex during cell migration
Nature Cell Biology (2021)