Abstract
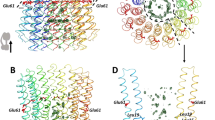
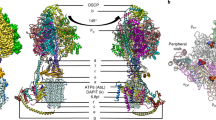
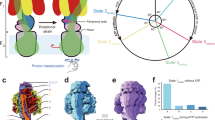
The proton pore of the F1Fo ATP synthase consists of a ring of c subunits, which rotates, driven by downhill proton diffusion across the membrane. An essential carboxylate side chain in each subunit provides a proton-binding site. In all the structures of c-rings reported to date, these sites are in a closed, ion-locked state. Structures are here presented of the c10 ring from Saccharomyces cerevisiae determined at pH 8.3, 6.1 and 5.5, at resolutions of 2.0 Å, 2.5 Å and 2.0 Å, respectively. The overall structure of this mitochondrial c-ring is similar to known homologs, except that the essential carboxylate, Glu59, adopts an open extended conformation. Molecular dynamics simulations reveal that opening of the essential carboxylate is a consequence of the amphiphilic nature of the crystallization buffer. We propose that this new structure represents the functionally open form of the c subunit, which facilitates proton loading and release.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Kabaleeswaran, V., Puri, N., Walker, J.E., Leslie, A.G. & Mueller, D.M. Novel features of the rotary catalytic mechanism revealed in the structure of yeast F1 ATPase. EMBO J. 25, 5433–5442 (2006).
Collinson, I.R., Skehel, J.M., Fearnley, I.M., Runswick, M.J. & Walker, J.E. The F1F0-ATPase complex from bovine heart mitochondria: The molar ratio of the subunits in the stalk region linking the F1 and F0 domains. Biochemistry 35, 12640–12646 (1996).
Collinson, I.R. et al. Fo membrane domain of ATP synthase from bovine heart mitochondria: purification, subunit composition, and reconstitution with F1-ATPase. Biochemistry 33, 7971–7978 (1994).
Stock, D., Leslie, A.G.W. & Walker, J.E. Molecular architecture of the rotary motor in ATP synthase. Science 286, 1700–1705 (1999).
Watt, I.N., Montgomery, M.G., Runswick, M.J., Leslie, A.G. & Walker, J.E. Bioenergetic cost of making an adenosine triphosphate molecule in animal mitochondria. Proc. Natl. Acad. Sci. USA 107, 16823–16827 (2010).
Dautant, A., Velours, J. & Giraud, M.F. Crystal structure of the Mg.ADP-inhibited state of the yeast F1c10-ATP synthase. J. Biol. Chem. 285, 29502–29510 (2010).
Kayalar, C., Rosing, J. & Boyer, P.D. An alternating site sequence for oxidative phosphorylation suggested by measurement of substrate binding patterns and exchange reaction inhibitions. J. Biol. Chem. 252, 2486–2491 (1977).
Abrahams, J.P., Leslie, A.G.W., Lutter, R. & Walker, J.E. Structure at 2.8 Å resolution of F1-ATPase from bovine heart mitochondria. Nature 370, 621–628 (1994).
Dickson, V.K., Silvester, J.A., Fearnley, I.M., Leslie, A.G. & Walker, J.E. On the structure of the stator of the mitochondrial ATP synthase. EMBO J. 25, 2911–2918 (2006).
Walker, J.E. & Dickson, V.K. The peripheral stalk of the mitochondrial ATP synthase. Biochim. Biophys. Acta 1757, 286–296 (2006).
Kabaleeswaran, V. et al. Asymmetric structure of the yeast F1 ATPase in the absence of bound nucleotides. J. Biol. Chem. 284, 10546–10551 (2009).
Rees, D.M., Leslie, A.G. & Walker, J.E. The structure of the membrane extrinsic region of bovine ATP synthase. Proc. Natl. Acad. Sci. USA 106, 21597–21601 (2009).
Meier, T. et al. Complete ion-coordination structure in the rotor ring of Na+-dependent F-ATP synthases. J. Mol. Biol. 391, 498–507 (2009).
Meier, T., Polzer, P., Diederichs, K., Welte, W. & Dimroth, P. Structure of the rotor ring of F-Type Na+-ATPase from Ilyobacter tartaricus. Science 308, 659–662 (2005).
Murata, T., Yamato, I., Kakinuma, Y., Leslie, A.G. & Walker, J.E. Structure of the rotor of the V-Type Na+-ATPase from Enterococcus hirae. Science 308, 654–659 (2005).
Pogoryelov, D., Yildiz, Ö., Faraldo-Gómez, J.D. & Meier, T. High-resolution structure of the rotor ring of a proton-dependent ATP synthase. Nat. Struct. Mol. Biol. 16, 1068–1073 (2009).
Vollmar, M., Schlieper, D., Winn, M., Buchner, C. & Groth, G. Structure of the c14 rotor ring of the proton translocating chloroplast ATP synthase. J. Biol. Chem. 284, 18228–18235 (2009).
Preiss, L., Yildiz, Ö., Hicks, D.B., Krulwich, T.A. & Meier, T. A new type of proton coordination in an F1Fo-ATP synthase rotor ring. PLoS Biol. 8, e1000443 (2010).
Mizutani, K. et al. Structure of the rotor ring modified with N,N′-dicyclohexylcarbodiimide of the Na+-transporting vacuolar ATPase. Proc. Natl. Acad. Sci. USA 108, 13474–13479 (2011).
Krah, A., Pogoryelov, D., Meier, T. & Faraldo-Gómez, J.D. On the structure of the proton-binding site in the Fo rotor of chloroplast ATP synthases. J. Mol. Biol. 395, 20–27 (2010).
Vik, S.B. & Antonio, B.J. A mechanism of proton translocation by F1F0 ATP synthases suggested by double mutants of the a subunit. J. Biol. Chem. 269, 30364–30369 (1994).
Pogoryelov, D. et al. Microscopic rotary mechanism of ion translocation in the Fo complex of ATP synthases. Nat. Chem. Biol. 6, 891–899 (2010).
Junge, W., Lill, H. & Engelbrecht, S. ATP synthase: an electrochemical transducer with rotatory mechanics. Trends Biochem. Sci. 22, 420–423 (1997).
Steed, P.R. & Fillingame, R.H. Aqueous accessibility to the transmembrane regions of subunit c of the Escherichia coli F1F0 ATP synthase. J. Biol. Chem. 284, 23243–23250 (2009).
Steed, P.R. & Fillingame, R.H. Subunit a facilitates aqueous access to a membrane-embedded region of subunit c in Escherichia coli F1F0 ATP synthase. J. Biol. Chem. 283, 12365–12372 (2008).
Rastogi, V.K. & Girvin, M.E. Structural changes linked to proton translocation by subunit c of the ATP synthase. Nature 402, 263–268 (1999).
Vincent, O.D., Schwem, B.E., Steed, P.R., Jiang, W. & Fillingame, R.H. Fluidity of structure and swiveling of helices in the subunit c ring of Escherichia coli ATP synthase as revealed by cysteine-cysteine cross-linking. J. Biol. Chem. 282, 33788–33794 (2007).
Ostermeier, C. & Michel, H. Crystallization of membrane proteins. Curr. Opin. Struct. Biol. 7, 697–701 (1997).
Pagadala, V., Vistain, L., Symersky, J. & Mueller, D.M. Characterization of the mitochondrial ATP synthase from yeast Saccharomyces cerevisae. J. Bioenerg. Biomembr. 43, 333–347 (2011).
Giraud, M.F. et al. Rotor architecture in the yeast and bovine F1-c-ring complexes of F-ATP synthase. J. Struct. Biol. 177, 490–497 (2012).
Epler, J.L., Shugart, L.R. & Barnett, W.E. N-formylmethionyl transfer ribonucleic acid in mitochondria from Neurospora. Biochemistry 9, 3575–3579 (1970).
Feldman, F. & Mahler, H.R. Mitochondrial biogenesis. Retention of terminal formylmethionine in membrane proteins and regulation of their synthesis. J. Biol. Chem. 249, 3702–3709 (1974).
Galper, J.B. & Darnell, J.E. The presence of N-formyl-methionyl-tRNA in HeLa cell mitochondria. Biochem. Biophys. Res. Commun. 34, 205–214 (1969).
Halbreich, A. & Rabinowitz, M. Isolation of Saccharomyces cerevisiae mitochondrial formyltetrahydrofolic acid:methionyl-tRNA transformylase and the hybridization of mitochondrial fMet-tRNA with mitochondrial DNA. Proc. Natl. Acad. Sci. USA 68, 294–298 (1971).
Michon, T., Galante, M. & Velours, J. NH2-terminal sequence of the isolated yeast ATP synthase subunit 6 reveals post-translational cleavage. Eur. J. Biochem. 172, 621–625 (1988).
Takeuchi, N. et al. Mammalian mitochondrial methionyl-tRNA transformylase from bovine liver. Purification, characterization, and gene structure. J. Biol. Chem. 273, 15085–15090 (1998).
Leone, V., Krah, A. & Faraldo-Gómez, J.D. On the question of hydronium binding to ATP-synthase membrane rotors. Biophys. J. 99, L53–L55 (2010).
Assadi-Porter, F.M. & Fillingame, R.H. Proton-translocating carboxyl of subunit c of F1Fo H+- ATP synthase: The unique environment suggested by the pKa determined by 1H NMR. Biochemistry 34, 16186–16193 (1995).
Williams, A. & Ibrahim, I.T. Carbodiimide chemistry: recent advances. Chem. Rev. 81, 589–636 (1981).
von Ballmoos, C. & Dimroth, P. Two distinct proton binding sites in the ATP synthase family. Biochemistry 46, 11800–11809 (2007).
Azzi, A., Casey, R.P. & Nalecz, M.J. The effect of N,N′-dicyclohexylcarbodiimide on enzymes of bioenergetic relevance. Biochim. Biophys. Acta 768, 209–226 (1984).
Krah, A. et al. Structural and energetic basis for H+ versus Na+ binding selectivity in ATP synthase Fo rotors. Biochim. Biophys. Acta 1797, 763–772 (2010).
Lightowlers, R.N., Howitt, S.M., Hatch, L., Gibson, F. & Cox, G.B. The proton pore in the Escherichia coli F0F1-ATPase: a requirement for arginine at position 210 of the a-subunit. Biochim. Biophys. Acta 894, 399–406 (1987).
Cain, B.D. & Simoni, R.D. Proton translocation by the F1F0ATPase of Escherichia coli. Mutagenic analysis of the a subunit. J. Biol. Chem. 264, 3292–3300 (1989).
Mitome, N. et al. Essential arginine residue of the Fo-a subunit in FoF1-ATP synthase has a role to prevent the proton shortcut without c-ring rotation in the Fo proton channel. Biochem. J. 430, 171–177 (2010).
Ujwal, R. & Bowie, J.U. Crystallizing membrane proteins using lipidic bicelles. Methods 55, 337–341 (2011).
Otwinowski, Z. & Minor, W. Processing of X-ray data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Laskowski, R.A., MacArthur, M.W., Moss, D.S. & Thorton, J.M. PROCHECK: a program software suite for macromolecular structure determination. J. Appl. Crystallogr. 26, 283–291 (1993).
Vagin, A. & Teplyakov, A. Molecular replacement with MOLREP. Acta Crystallogr. D Biol. Crystallogr. 66, 22–25 (2010).
Perrakis, A., Morris, R. & Lamzin, V.S. Automated protein model building combined with iterative structure refinement. Nat. Struct. Biol. 6, 458–463 (1999).
Murshudov, G.N., Vagin, A.A. & Dodson, E.J. Refinement of macromolecular structures by the maximum-likelihood method. Acta Crystallogr. D Biol. Crystallogr. 53, 240–255 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Staritzbichler, R., Anselmi, C., Forrest, L.R. & Faraldo-Gómez, J.D. GRIFFIN: a versatile methodology for optimization of protein-lipid interfaces for membrane protein simulations. J. Chem. Theory Comput. 7, 1167–1176 (2011).
Stoscheck, C.M. Quantitation of proteins. Methods Enzymol. 182, 50–68 (1990).
Baker, N.A., Sept, D., Joseph, S., Holst, M.J. & McCammon, J.A. Electrostatics of nanosystems: application to microtubules and the ribosome. Proc. Natl. Acad. Sci. USA 98, 10037–10041 (2001).
Sitkoff, D., Sharp, K.A. & Honig, B. Accurate calculation of hydration free energies using macroscopic solvent models. J. Phys. Chem. 98, 1978–1988 (1994).
Acknowledgements
The work was supported by a grant from the US National Institutes of Health R01GM66223 to D.M.M. T.M. and J.D.F.-G. are funded in part by the German Research Foundation Cluster of Excellence 'Macromolecular Complexes' (EXC115). Diffraction data were collected at the General Medicine and Cancer Institutes Collaborative Access Team (GM/CA-CAT) 23-ID and Southeast Regional Collaborative Access Team (SER-CAT) 22-ID beamlines at the Advanced Photon Source, Argonne National Laboratory. Use of the Advanced Photon Source was supported by the US Department of Energy, Office of Science, Office of Basic Energy Sciences, under contract number W-31-109-Eng-38. Computational resources were provided in part by the Jülich Supercomputing Center. We thank the reviewer for recognizing that residues 15–18 form a 310 helix.
Author information
Authors and Affiliations
Contributions
J.S. collected and analyzed diffraction data and collaborated in writing the manuscript. V.P. and D.O. designed and conducted experiments and aided in collecting diffraction data. J.D.F.-G. designed the research, analyzed data and collaborated in writing the manuscript. A.K. conducted the research and analyzed data. T.M. analyzed data and collaborated in writing the manuscript. D.M.M. designed experiments, analyzed data, aided in collecting diffraction data and collaborated in writing the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 13182 kb)
Rights and permissions
About this article
Cite this article
Symersky, J., Pagadala, V., Osowski, D. et al. Structure of the c10 ring of the yeast mitochondrial ATP synthase in the open conformation. Nat Struct Mol Biol 19, 485–491 (2012). https://doi.org/10.1038/nsmb.2284
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.2284
This article is cited by
-
Mechanisms of membrane protein crystallization in ‘bicelles’
Scientific Reports (2022)
-
Bedaquiline inhibits the yeast and human mitochondrial ATP synthases
Communications Biology (2020)
-
Molecular dynamics simulation of proton-transfer coupled rotations in ATP synthase FO motor
Scientific Reports (2020)
-
Unusual features of the c-ring of F1FO ATP synthases
Scientific Reports (2019)
-
Crucial aminoacids in the FO sector of the F1FO-ATP synthase address H+ across the inner mitochondrial membrane: molecular implications in mitochondrial dysfunctions
Amino Acids (2019)