Abstract
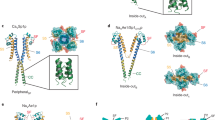
We report the crystal structure of the nonselective cation channel NaK from Bacillus cereus at a resolution of 1.6 Å. The structure reveals the intracellular gate in an open state, as opposed to the closed form reported previously, making NaK the only channel for which the three-dimensional structures of both conformations are known. Channel opening follows a conserved mechanism of inner helix bending using a flexible glycine residue, the gating hinge, seen in MthK and most other tetrameric cation channels. Additionally, distinct inter and intrasubunit rearrangements involved in channel gating are seen and characterized for the first time along with inner helix twisting motions. Furthermore, we identify a residue deeper within the cavity of the channel pore, Phe92, which is likely to form a constriction point within the open pore, restricting ion flux through the channel. Mutating this residue to alanine causes a subsequent increase in ion-conduction rates as measured by 86Rb flux assays. The structures of both the open and closed conformations of the NaK channel correlate well with those of equivalent K+ channel conformations, namely MthK and KcsA, respectively.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Hille, B. Ion Channels of Excitable Membranes. 3rd edn. (Sinauer, Sunderland, MA, 2001).
Jiang, Y. et al. Crystal structure and mechanism of a calcium-gated potassium channel. Nature 417, 515–522 (2002).
Zhou, Y. et al. Chemistry of ion coordination and hydration revealed by a K+ channel-Fab complex at 2.0 resolution. Nature 414, 43–48 (2001).
Jiang, Y. et al. The open pore conformation of potassium channels. Nature 417, 523–526 (2002).
Doyle, D.A. et al. The structure of the potassium channel: molecular basis of K+ conduction and selectivity. Science 280, 69–77 (1998).
Long, S.B. et al. Atomic structure of a voltage-dependent K+ channel in a lipid membrane-like environment. Nature 450, 376–382 (2007).
Long, S.B., Campbell, E.B. & Mackinnon, R. Crystal structure of a mammalian voltage-dependent Shaker family K+ channel. Science 309, 897–903 (2005).
Kuo, A. et al. Crystal structure of the potassium channel KirBac1.1 in the closed state. Science 300, 1922–1926 (2003).
Nishida, M. et al. Crystal structure of a Kir3.1-prokaryotic Kir channel chimera. EMBO J. 26, 4005–4015 (2007).
Alam, A., Shi, N. & Jiang, Y. Structural insight into Ca2+ specificity in tetrameric cation channels. Proc. Natl. Acad. Sci. USA 104, 15334–15339 (2007).
Shi, N. et al. Atomic structure of a Na+- and K+-conducting channel. Nature 440, 570–574 (2006).
Alam, A. & Jiang, Y. Structural analysis of ion selectivity in the NaK channel. Nat. Struct. Mol. Biol. advance online publication, doi:10.1038/nsmb.1537 (21 December 2008).
Jin, T. et al. The βγ subunits of G proteins gate a K+ channel by pivoted bending of a transmembrane segment. Mol. Cell 10, 469–481 (2002).
Magidovich, E. & Yifrach, O. Conserved gating hinge in ligand- and voltage-dependent K+ channels. Biochemistry 43, 13242–13247 (2004).
Zhao, Y. et al. A gating hinge in Na+ channels; a molecular switch for electrical signaling. Neuron 41, 859–865 (2004).
Ding, S. et al. Investigating the putative glycine hinge in Shaker potassium channel. J. Gen. Physiol. 126, 213–226 (2005).
Perozo, E., Cortes, D.M. & Cuello, L.G. Three-dimensional architecture and gating mechanism of a K+ channel studied by EPR spectroscopy. Nat. Struct. Biol. 5, 459–469 (1998).
Perozo, E., Cortes, D.M. & Cuello, L.G. Structural rearrangements underlying K+-channel activation gating. Science 285, 73–78 (1999).
Liu, Y.S., Sompornpisut, P. & Perozo, E. Structure of the KcsA channel intracellular gate in the open state. Nat. Struct. Biol. 8, 883–887 (2001).
Kelly, B.L. & Gross, A. Potassium channel gating observed with site-directed mass tagging. Nat. Struct. Biol. 10, 280–284 (2003).
Zimmer, J., Doyle, D.A. & Grossmann, J.G. Structural characterization and pH-induced conformational transition of full-length KcsA. Biophys. J. 90, 1752–1766 (2006).
Iwamoto, M. et al. Surface structure and its dynamic rearrangements of the KcsA potassium channel upon gating and tetrabutylammonium blocking. J. Biol. Chem. 281, 28379–28386 (2006).
Baker, K.A. et al. Conformational dynamics of the KcsA potassium channel governs gating properties. Nat. Struct. Mol. Biol. 14, 1089–1095 (2007).
Takeuchi, K. et al. Identification and characterization of the slowly exchanging pH-dependent conformational rearrangement in KcsA. J. Biol. Chem. 282, 15179–15186 (2007).
Shimizu, H. et al. Global twisting motion of single molecular KcsA potassium channel upon gating. Cell 132, 67–78 (2008).
Rabenstein, M.D. & Shin, Y.K. Determination of the distance between two spin labels attached to a macromolecule. Proc. Natl. Acad. Sci. USA 92, 8239–8243 (1995).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Jones, T.A. et al. Improved methods for building protein models in electron density maps and the location of errors in these models. Acta Crystallogr. A 47, 110–119 (1991).
Brunger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Smart, O.S., Goodfellow, J.M. & Wallace, B.A. The pore dimensions of gramicidin A. Biophys. J. 65, 2455–2460 (1993).
Humphrey, W., Dalke, A. & Schulten, K. VMD: visual molecular dynamics. J. Mol. Graph. 14, 33–8, 27–8 (1996).
Heginbotham, L. et al. Single Streptomyces lividans K+ channels: functional asymmetries and sidedness of proton activation. J. Gen. Physiol. 114, 551–560 (1999).
Heginbotham, L., Kolmakova-Partensky, L. & Miller, C. Functional reconstitution of a prokaryotic K+ channel. J. Gen. Physiol. 111, 741–749 (1998).
Acknowledgements
Use of the Argonne National Laboratory Structural Biology Center beamlines at the Advanced Photon Source was supported by the US Department of Energy, Office of Energy Research. We thank the beamline staff for assistance in data collection. This work was supported by the Howard Hughes Medical Institute and by grants from the the US National Institutes of Health and National Institute of General Medical Sciences (RO1 GM079179), the David and Lucile Packard Foundation and the McKnight Endowment for Neuroscience. A.A. was supported by National Institutes of Health Training Grant T32 GM008297.
Author information
Authors and Affiliations
Contributions
A.A. performed the research; A.A and Y.J designed the research, analyzed data and wrote the manuscript.
Corresponding author
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 (PDF 4190 kb)
Rights and permissions
About this article
Cite this article
Alam, A., Jiang, Y. High-resolution structure of the open NaK channel. Nat Struct Mol Biol 16, 30–34 (2009). https://doi.org/10.1038/nsmb.1531
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.1531
This article is cited by
-
Ion-dependent structure, dynamics, and allosteric coupling in a non-selective cation channel
Nature Communications (2021)
-
Inferring functional units in ion channel pores via relative entropy
European Biophysics Journal (2021)
-
Pharmacologically reversible, loss of function mutations in the TM2 and TM4 inner pore helices of TREK-1 K2P channels
Scientific Reports (2019)
-
Equilibria between the K+ binding and cation vacancy conformations of potassium channels
Protein & Cell (2019)
-
Direct knock-on of desolvated ions governs strict ion selectivity in K+ channels
Nature Chemistry (2018)