Abstract
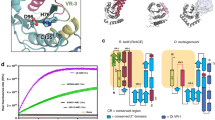
Bacterial pathogens deliver virulence proteins into host cells to facilitate entry and survival. Salmonella SopA functions as an E3 ligase to manipulate the host proinflammatory response. Here we report the crystal structure of SopA in two conformations. Although it has little sequence similarity to eukaryotic HECT-domain E3s, the C-terminal half of SopA has a bilobal architecture that is reminiscent of the N- and C-lobe arrangement of HECT domains. The SopA structure also contains a putative substrate-binding domain located near the E2-binding site. The two structures of SopA differ in the relative orientations of the C lobe, indicating that SopA possesses the conformational flexibility essential for HECT E3 function. These results suggest that SopA is a unique HECT E3 ligase evolved from the coevolutionary selective pressure at the bacterium-host interface.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Pickart, C.M. Mechanisms underlying ubiquitination. Annu. Rev. Biochem. 70, 503–533 (2001).
Huibregtse, J.M., Scheffner, M., Beaudenon, S. & Howley, P.M. A family of proteins structurally and functionally related to the E6-AP ubiquitin-protein ligase. Proc. Natl. Acad. Sci. USA 92, 2563–2567 (1995).
Scheffner, M., Nuber, U. & Huibregtse, J.M. Protein ubiquitination involving an E1–E2–E3 enzyme ubiquitin thioester cascade. Nature 373, 81–83 (1995).
Huang, L. et al. Structure of an E6AP-UbcH7 complex: insights into ubiquitination by the E2–E3 enzyme cascade. Science 286, 1321–1326 (1999).
Ogunjimi, A.A. et al. Regulation of Smurf2 ubiquitin ligase activity by anchoring the E2 to the HECT domain. Mol. Cell 19, 297–308 (2005).
Verdecia, M.A. et al. Conformational flexibility underlies ubiquitin ligation mediated by the WWP1 HECT domain E3 ligase. Mol. Cell 11, 249–259 (2003).
Patel, J.C., Rossanese, O.W. & Galán, J.E. The functional interface between Salmonella and its host cell: opportunities for therapeutic intervention. Trends Pharmacol. Sci. 26, 564–570 (2005).
Stebbins, C.E. & Galan, J.E. Structural mimicry in bacterial virulence. Nature 412, 701–705 (2001).
Zhang, Y., Higashide, W.M., McCormick, B.A., Chen, J. & Zhou, D. The inflammation-associated Salmonella SopA is a HECT-like E3 ubiquitin ligase. Mol. Microbiol. 62, 786–793 (2006).
Huibregtse, J.M., Scheffner, M. & Howley, P.M. Localization of the E6-AP regions that direct human papillomavirus E6 binding, association with p53, and ubiquitination of associated proteins. Mol. Cell. Biol. 13, 4918–4927 (1993).
Jenkins, J. & Pickersgill, R. The architecture of parallel β-helices and related folds. Prog. Biophys. Mol. Biol. 77, 111–175 (2001).
Eletr, Z.M., Huang, D.T., Duda, D.M., Schulman, B.A. & Kuhlman, B. E2 conjugating enzymes must disengage from their E1 enzymes before E3-dependent ubiquitin and ubiquitin-like transfer. Nat. Struct. Mol. Biol. 12, 933–934 (2005).
Eletr, Z.M. & Kuhlman, B. Sequence determinants of E2–E6AP binding affinity and specificity. J. Mol. Biol. 369, 419–428 (2007).
Nuber, U. & Scheffner, M. Identification of determinants in E2 ubiquitin-conjugating enzymes required for hect E3 ubiquitin-protein ligase interaction. J. Biol. Chem. 274, 7576–7582 (1999).
Zheng, N. A closer look of the HECTic ubiquitin ligases. Structure 11, 5–6 (2003).
Janjusevic, R., Abramovitch, R.B., Martin, G.B. & Stebbins, C.E. A bacterial inhibitor of host programmed cell death defenses is an E3 ubiquitin ligase. Science 311, 222–226 (2006).
Rohde, J.R., Breitkreutz, A., Chenal, A., Sansonetti, P.J. & Parsot, C. Type III secretion effectors of the IpaH family are E3 ubiquitin ligases. Cell Host & Microbe 1, 77–83 (2007).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Abrahams, J.P. & Leslie, A.G. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Collaborative Computational Project, Number 4. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Brünger, A.T. et al. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D Biol. Crystallogr. 54, 905–921 (1998).
Zhang, Y., Higashide, W., Dai, S., Sherman, D.M. & Zhou, D. Recognition and ubiquitination of Salmonella type III effector SopA by a ubiquitin E3 ligase, HsRMA1. J. Biol. Chem. 280, 38682–38688 (2005).
Acknowledgements
We thank the staff at the Advanced Photon Source beam line 23-ID for assistance with data collection. This work was supported by US National Institutes of Health grants (AI049978 to D.Z. and CA072943 to J.M.H.) and by a Pew scholarship (to J.C.).
Author information
Authors and Affiliations
Contributions
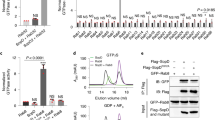
J.D. determined the structures of SopA and contributed the data for Figure 4a,b. Y.Z. subcloned most of the constructs and contributed the data for Figures 1 and 4c and Supplementary Figure 3. J.D., Y.Z., J.M.H., D.Z. and J.C. designed experiments, analyzed data and prepared the manuscript.
Corresponding authors
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 (PDF 244 kb)
Rights and permissions
About this article
Cite this article
Diao, J., Zhang, Y., Huibregtse, J. et al. Crystal structure of SopA, a Salmonella effector protein mimicking a eukaryotic ubiquitin ligase. Nat Struct Mol Biol 15, 65–70 (2008). https://doi.org/10.1038/nsmb1346
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb1346
This article is cited by
-
Structural basis for the recognition and degradation of host TRIM proteins by Salmonella effector SopA
Nature Communications (2017)
-
Bacteria-host relationship: ubiquitin ligases as weapons of invasion
Cell Research (2016)
-
Genomic characterization of Salmonella Cerro ST367, an emerging Salmonella subtype in cattle in the United States
BMC Genomics (2014)
-
Exploitation of the host ubiquitin system by human bacterial pathogens
Nature Reviews Microbiology (2014)
-
Macromolecular juggling by ubiquitylation enzymes
BMC Biology (2013)