Abstract
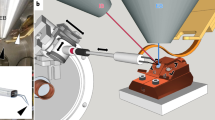
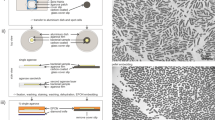
This protocol details methods for the isolation of yeast nuclei from budding yeast (Saccharomyces cerevisiae) and fission yeast (Schizosaccharomyces pombe), immuno-gold labeling of proteins and visualization by field emission scanning electron microscopy (FESEM). This involves the removal of the yeast cell wall and isolation of the nucleus from within, followed by subsequent processing for high-resolution microscopy. The nuclear isolation step can be performed in two ways: enzymatic treatment of yeast cells to rupture the cell wall and generate spheroplasts (cells that have partially lost their cell wall and their characteristic shape), followed by isolation of the nuclei by centrifugation or homogenization; and whole cell freezing followed by manual cell rupture and centrifugation. This protocol has been optimized for the visualization of the yeast nuclear envelope (NE), nuclear pore complexes (NPCs) and associated cyto-skeletal structures. Samples once processed for FESEM can be stored under vacuum for weeks, allowing considerable time for image acquisition.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
References
Nurse, P. Genetic control of cell size at cell division in yeast. Nature 256, 547–551 (1975).
Nurse, P. Coordinating histone transcription and DNA replication. Nature 302, 378 (1983).
Fraser, R.S. & Nurse, P. Novel cell cycle control of RNA synthesis in yeast. Nature 271, 726–730 (1978).
Nurse, P. & Bissett, Y. Gene required in G1 for commitment to cell cycle and in G2 for control of mitosis in fission yeast. Nature 292, 558–560 (1981).
Sillevis Smitt, W.W., Nanni, G., Rozijn, T.H. & Tonino, G.J. Sedimentation characteristics of RNA from isolated yeast nuclei. Exp. Cell Res. 59, 440–446 (1970).
Mason, J.A. & Mellor, J. Isolation of nuclei for chromatin analysis in fission yeast. Nucleic Acids Res. 25, 4700–4701 (1997).
Zhang, Z. & Reese, J.C. Isolation of yeast nuclei and micrococcal nuclease mapping of nucleosome positioning. Methods Mol. Biol. 313, 245–255 (2006).
Rout, M.P. & Blobel, G. Isolation of the yeast nuclear pore complex. J. Cell Biol. 123, 771–783 (1993).
Winey, M., Yarar, D., Giddings, T.H. Jr. & Mastronarde, D.N. Nuclear pore complex number and distribution throughout the Saccharomyces cerevisiae cell cycle by three-dimensional reconstruction from electron micrographs of nuclear envelopes. Mol. Biol. Cell 8, 2119–2132 (1997).
Yang, Q., Rout, M.P. & Akey, C.W. Three-dimensional architecture of the isolated yeast nuclear pore complex: functional and evolutionary implications. Mol. Cell 1, 223–234 (1998).
Allen, J.L. & Douglas, M.G. Organization of the nuclear pore complex in Saccharomyces cerevisiae. J. Ultrastruct. Mol. Struct. Res. 102, 95–108 (1989).
Heymann, J.A. et al. Site-specific 3D imaging of cells and tissues with a dual beam microscope. J. Struct. Biol. 155, 63–73 (2006).
Kiseleva, E. et al. Yeast nuclear pore complexes have a cytoplasmic ring and internal filaments. J. Struct. Biol. 145, 272–288 (2004).
Rozjin, T.H. & Tonino, G.J. Studies on the yeast nucleus. I. The isolation of nuclei. Biochim. Biophys. Acta. 91, 105–112 (1964).
Ho, A.K. et al. Assembly and preferential localization of Nup116p on the cytoplasmic face of the nuclear pore complex by interaction with Nup82p. Mol. Cell Biol. 20, 5736–5748 (2000).
Allen, T. et al. A protocol for isolating Xenopus oocyte nuclear envelope for visualization and characterization by scanning electron microscopy (SEM) or transmission electron microscopy (TEM). Nat. Protoc. 2, 1166–1172 (2007).
Acknowledgements
T.D.A, S.A.R., S.M. and S.P.D. would like to acknowledge the support of Cancer Research (CR)-UK. E.K. acknowledges The Wellcome Trust and Russian Federation for Basic Research, M.W.G acknowledges The Wellcome Trust, and F.G. acknowledges the MRC. The authors thank Professor M. Rout (The Rockefeller University, New York, NY) for his advice in isolation of nuclei from frozen yeast.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Kiseleva, E., Allen, T., Rutherford, S. et al. A protocol for isolation and visualization of yeast nuclei by scanning electron microscopy (SEM). Nat Protoc 2, 1943–1953 (2007). https://doi.org/10.1038/nprot.2007.251
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2007.251
This article is cited by
-
Enhancing co-translational folding of heterologous protein by deleting non-essential ribosomal proteins in Pichia pastoris
Biotechnology for Biofuels (2019)
-
The tethering of chromatin to the nuclear envelope supports nuclear mechanics
Nature Communications (2015)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.