Abstract
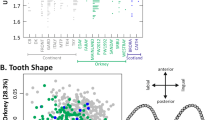
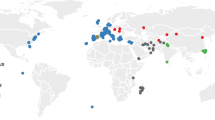
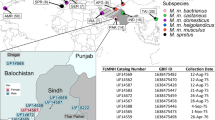
Here we provide a genome-wide, high-resolution map of the phylogenetic origin of the genome of most extant laboratory mouse inbred strains. Our analysis is based on the genotypes of wild-caught mice from three subspecies of Mus musculus. We show that classical laboratory strains are derived from a few fancy mice with limited haplotype diversity. Their genomes are overwhelmingly Mus musculus domesticus in origin, and the remainder is mostly of Japanese origin. We generated genome-wide haplotype maps based on identity by descent from fancy mice and show that classical inbred strains have limited and non-randomly distributed genetic diversity. In contrast, wild-derived laboratory strains represent a broad sampling of diversity within M. musculus. Intersubspecific introgression is pervasive in these strains, and contamination by laboratory stocks has played a role in this process. The subspecific origin, haplotype diversity and identity by descent maps can be visualized using the Mouse Phylogeny Viewer (see URLs).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Boursot, P., Auffray, J.C., Britton-Davidian, J. & Bonhomme, F. The evolution of the house mice. Annu. Rev. Ecol. Syst. 24, 119–152 (1993).
Geraldes, A. et al. Inferring the history of speciation in house mice from autosomal, X-linked, Y-linked and mitochondrial genes. Mol. Ecol. 17, 5349–5363 (2008).
Teeter, K.C. et al. Genome-wide patterns of gene flow across a house mouse hybrid zone. Genome Res. 18, 67–76 (2008).
Yonekawa, H., Takahama, S., Gotoh, O., Miyashita, N. & Moriwaki, K. Genetic diversity and geographic distribution of Mus musculus subspecies based on the polymorphism of mitochondrial DNA. in Genetics in Wild Mice. Its application to Biomedical Research (eds Moriwaki, K., Shiroishi, T. and Yonekawa, H.) 25–40 (Japan Scientific Societies Press, Tokyo, Japan, 1994).
Beck, J.A. et al. Genealogies of mouse inbred strains. Nat. Genet. 24, 23–25 (2000).
Frazer, K.A. et al. A sequence-based variation map of 8.27 million SNPs in inbred mouse strains. Nature 448, 1050–1053 (2007).
Yang, H., Bell, T.A., Churchill, G.A. & Pardo-Manuel de Villena, F . On the subspecific origin of the laboratory mouse. Nat. Genet. 39, 1100–1107 (2007).
Guénet, J.L. & Bonhomme, F. Wild mice: an ever-increasing contribution to a popular mammalian model. Trends Genet. 19, 24–31 (2003).
Mouse Genome Sequencing Consortium et al. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Sudbery, I. et al. Deep short-read sequencing of chromosome 17 from the mouse strains A/J and CAST/Ei identifies significant germline variation and candidate genes that regulate liver triglyceride levels. Genome Biol. 10, R112 (2009).
Chesler, E.J. et al. The Collaborative Cross at Oak Ridge National Laboratory: developing a powerful resource for systems genetics. Mamm. Genome 19, 382–389 (2008).
Guan, C., Ye, C., Yang, X. & Gao, J. A review of current large-scale mouse knockout efforts. Genesis 48, 73–85 (2010).
Szatkiewicz, J.P. et al. An imputed genotype resource for the laboratory mouse. Mamm. Genome 19, 199–208 (2008).
Harr, B. Genomic islands of differentiation between house mouse subspecies. Genome Res. 16, 730–737 (2006).
Boursot, P. & Belkhir, K. Mouse SNPs for evolutionary biology: beware of ascertainment biases. Genome Res. 16, 1191–1192 (2006).
White, M.A., Ané, C., Dewey, C.N., Larget, B.R. & Payseur, B.A. Fine-scale phylogenetic discordance across the house mouse genome. PLoS Genet. 5, e1000729 (2009).
Yang, H. et al. A customized and versatile high-density genotyping array for the mouse. Nat. Methods 6, 663–666 (2009).
Nagamine, C.M. et al. The musculus-type Y chromosome of the laboratory mouse is of Asian origin. Mamm. Genome 3, 84–91 (1992).
Tucker, P.K., Lee, B.K., Lundrigan, B.L. & Eicher, E.M. Geographic origin of the Y chromosomes in “old” inbred strains of mice. Mamm. Genome 3, 254–261 (1992).
Mihola, O., Trachtulec, Z., Vlcek, C., Schimenti, J.C. & Forejt, J. A mouse speciation gene encodes a meiotic histone H3 methyltransferase. Science 323, 373–375 (2009).
Ideraabdullah, F.Y. et al. Genetic and haplotype diversity among wild-derived mouse inbred strains. Genome Res. 14, 1880–1887 (2004).
Wang, J., Moore, K.J., Zhang, Q., Pardo-Manuel de Villena, F., Wang, W. & McMillan, L. Genome-wide compatible SNP intervals and their properties. Proceedings of ACM International Conference on Bioinformatics and Computational Biology (Niagara Falls, New York, USA, 2010).
Acknowledgements
This work was supported by the National Institute of General Medical Sciences (NIGMS) Centers of Excellence in Systems Biology program, grant GM-076468, by a US National Institutes of Health (NIH) grant to M.W.N. (R01 GM74245), by a grant to F.B. (ISEM 2010-141) and by a Czech Science Foundation grant to J.P. (206-08-0640). J.P.D. was partially supported by NIH Training Grant Number GM067553-04, University of North Carolina (UNC) Bioinformatics and Computational Biology Training Grant. J.P.D., R.J.B. and T.A.B. are partially supported by an NIH grant to F.P.-M.d.V. (P50 MH090338). We also thank F. Oyola for help annotating the samples genotyped in this study.
Author information
Authors and Affiliations
Contributions
F.P.-M.d.V., G.A.C. and H.Y. conceived the study design and wrote the paper. H.Y., J.R.W., J.P.D., L.M. and C.E.W. carried out the bioinformatics analyses. J.P.D., T.A.B. and R.J.B. prepared the samples and conducted the targeted PCR amplification and sequencing. F.B., P.B., A.H.-T.Y., M.W.N., J.P. and P.T. provided biological samples. All authors contributed to the interpretation of the results and the writing of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 and Supplementary Tables 2–6. (PDF 7417 kb)
Supplementary Table 1
Sample summary (XLSX 90 kb)
Rights and permissions
About this article
Cite this article
Yang, H., Wang, J., Didion, J. et al. Subspecific origin and haplotype diversity in the laboratory mouse. Nat Genet 43, 648–655 (2011). https://doi.org/10.1038/ng.847
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.847
This article is cited by
-
Insertion of short L1 sequences generates inter-strain histone acetylation differences in the mouse
Mobile DNA (2024)
-
Multi-omics analysis identifies drivers of protein phosphorylation
Genome Biology (2023)
-
A genetic locus complements resistance to Bordetella pertussis-induced histamine sensitization
Communications Biology (2023)
-
Taxonomic assessment of two wild house mouse subspecies using whole-genome sequencing
Scientific Reports (2022)
-
Population structure and inbreeding in wild house mice (Mus musculus) at different geographic scales
Heredity (2022)