Abstract
During non-rapid eye movement (NREM) sleep, synchronous synaptic activity in the thalamocortical network generates predominantly low-frequency oscillations (<4 Hz) that are modulated by inhibitory inputs from the thalamic reticular nucleus (TRN). Whether TRN cells integrate sleep-wake signals from subcortical circuits remains unclear. We found that GABA neurons from the lateral hypothalamus (LHGABA) exert a strong inhibitory control over TRN GABA neurons (TRNGABA). We found that optogenetic activation of this circuit recapitulated state-dependent changes of TRN neuron activity in behaving mice and induced rapid arousal during NREM, but not REM, sleep. During deep anesthesia, activation of this circuit induced sustained cortical arousal. In contrast, optogenetic silencing of LHGABA-TRNGABA transmission increased the duration of NREM sleep and amplitude of delta (1–4 Hz) oscillations. Collectively, these results demonstrate that TRN cells integrate subcortical arousal inputs selectively during NREM sleep and may participate in sleep intensity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Steriade, M. The corticothalamic system in sleep. Front. Biosci. 8, d878–d899 (2003).
Lee, S.-H. & Dan, Y. Neuromodulation of brain states. Neuron 76, 209–222 (2012).
Saper, C.B., Fuller, P.M., Pedersen, N.P., Lu, J. & Scammell, T.E. Sleep state switching. Neuron 68, 1023–1042 (2010).
Steriade, M., Nuñez, A. & Amzica, F. A novel slow (< 1 Hz) oscillation of neocortical neurons in vivo: depolarizing and hyperpolarizing components. J. Neurosci. 13, 3252–3265 (1993).
Timofeev, I. & Chauvette, S. Thalamocortical oscillations: local control of EEG slow waves. Curr. Top. Med. Chem. 11, 2457–2471 (2011).
Lo˝rincz, M.L. et al. A distinct class of slow (∼0.2–2 Hz) intrinsically bursting layer 5 pyramidal neurons determines UP/DOWN state dynamics in the neocortex. J. Neurosci. 35, 5442–5458 (2015).
Huguenard, J.R. & McCormick, D.A. Thalamic synchrony and dynamic regulation of global forebrain oscillations. Trends Neurosci. 30, 350–356 (2007).
Fuentealba, P. & Steriade, M. The reticular nucleus revisited: intrinsic and network properties of a thalamic pacemaker. Prog. Neurobiol. 75, 125–141 (2005).
Cox, C.L., Huguenard, J.R. & Prince, D.A. Nucleus reticularis neurons mediate diverse inhibitory effects in thalamus. Proc. Natl. Acad. Sci. USA 94, 8854–8859 (1997).
Kim, U., Sanchez-Vives, M.V. & McCormick, D.A. Functional dynamics of GABAergic inhibition in the thalamus. Science 278, 130–134 (1997).
Golshani, P., Liu, X.B. & Jones, E.G. Differences in quantal amplitude reflect GluR4- subunit number at corticothalamic synapses on two populations of thalamic neurons. Proc. Natl. Acad. Sci. USA 98, 4172–4177 (2001).
Knyihar-Csillik, E., Chadaide, Z., Mihaly, A., Krisztin-Peva, B. & Csillik, B. Effect of electrical stimulation of the reticular nucleus of the rat thalamus upon c-fos immunoreactivity in the retrosplenial cortex. Ann. Anat. 187, 245–249 (2005).
Pita-Almenar, J.D., Yu, D., Lu, H.-C. & Beierlein, M. Mechanisms underlying desynchronization of cholinergic-evoked thalamic network activity. J. Neurosci. 34, 14463–14474 (2014).
Sun, Y.-G. et al. Biphasic cholinergic synaptic transmission controls action potential activity in thalamic reticular nucleus neurons. J. Neurosci. 33, 2048–2059 (2013).
Cox, C.L., Huguenard, J.R. & Prince, D.A. Peptidergic modulation of intrathalamic circuit activity in vitro: actions of cholecystokinin. J. Neurosci. 17, 70–82 (1997).
Brill, J., Kwakye, G. & Huguenard, J.R. NPY signaling through Y1 receptors modulates thalamic oscillations. Peptides 28, 250–256 (2007).
Le Masson, G., Renaud-Le Masson, S., Debay, D. & Bal, T. Feedback inhibition controls spike transfer in hybrid thalamic circuits. Nature 417, 854–858 (2002).
Hill, S. & Tononi, G. Modeling sleep and wakefulness in the thalamocortical system. J. Neurophysiol. 93, 1671–1698 (2005).
Destexhe, A., Contreras, D., Sejnowski, T.J. & Steriade, M. Modeling the control of reticular thalamic oscillations by neuromodulators. Neuroreport 5, 2217–2220 (1994).
Browne, S.H. et al. Kinetic and pharmacological properties of GABA(A) receptors in single thalamic neurons and GABA(A) subunit expression. J. Neurophysiol. 86, 2312–2322 (2001).
Huntsman, M.M. & Huguenard, J.R. Fast IPSCs in rat thalamic reticular nucleus require the GABAA receptor beta1 subunit. J. Physiol. (Lond.) 572, 459–475 (2006).
Schofield, C.M. & Huguenard, J.R. GABA affinity shapes IPSCs in thalamic nuclei. J. Neurosci. 27, 7954–7962 (2007).
Macdonald, K.D., Fifkova, E., Jones, M.S. & Barth, D.S. Focal stimulation of the thalamic reticular nucleus induces focal gamma waves in cortex. J. Neurophysiol. 79, 474–477 (1998).
Halassa, M.M. et al. Selective optical drive of thalamic reticular nucleus generates thalamic bursts and cortical spindles. Nat. Neurosci. 14, 1118–1120 (2011).
Kim, A. et al. Optogenetically induced sleep spindle rhythms alter sleep architectures in mice. Proc. Natl. Acad. Sci. USA 109, 20673–20678 (2012).
Adamantidis, A.R., Zhang, F., Aravanis, A.M., Deisseroth, K. & de Lecea, L. Neural substrates of awakening probed with optogenetic control of hypocretin neurons. Nature 450, 420–424 (2007).
Jego, S. et al. Optogenetic identification of a rapid eye movement sleep modulatory circuit in the hypothalamus. Nat. Neurosci. 16, 1637–1643 (2013).
Kempadoo, K.A. et al. Hypothalamic neurotensin projections promote reward by enhancing glutamate transmission in the VTA. J. Neurosci. 33, 7618–7626 (2013).
Nieh, E.H. et al. Decoding neural circuits that control compulsive sucrose seeking. Cell 160, 528–541 (2015).
Bonnavion, P., Jackson, A.C., Carter, M.E. & de Lecea, L. Antagonistic interplay between hypocretin and leptin in the lateral hypothalamus regulates stress responses. Nat. Commun. 6, 6266 (2015).
Hassani, O.K., Henny, P., Lee, M.G. & Jones, B.E. GABAergic neurons intermingled with orexin and MCH neurons in the lateral hypothalamus discharge maximally during sleep. Eur. J. Neurosci. 32, 448–457 (2010).
Jennings, J.H. et al. Visualizing hypothalamic network dynamics for appetitive and consummatory behaviors. Cell 160, 516–527 (2015).
Vong, L. et al. Leptin action on GABAergic neurons prevents obesity and reduces inhibitory tone to POMC neurons. Neuron 71, 142–154 (2011).
Leinninger, G.M. et al. Leptin acts via leptin receptor-expressing lateral hypothalamic neurons to modulate the mesolimbic dopamine system and suppress feeding. Cell Metab. 10, 89–98 (2009).
Schöne, C., Apergis-Schoute, J., Sakurai, T., Adamantidis, A. & Burdakov, D. Coreleased orexin and glutamate evoke nonredundant spike outputs and computations in histamine neurons. Cell Reports 7, 697–704 (2014).
Halassa, M.M. et al. State-dependent architecture of thalamic reticular subnetworks. Cell 158, 808–821 (2014).
Borbély, A.A., Achermann, P., Trachsel, L. & Tobler, I. Sleep initiation and initial sleep intensity: interactions of homeostatic and circadian mechanisms. J. Biol. Rhythms 4, 149–160 (1989).
Andrada, J., Livingston, P., Lee, B.J. & Antognini, J. Propofol and etomidate depress cortical, thalamic, and reticular formation neurons during anesthetic-induced unconsciousness. Anesth. Analg. 114, 661–669 (2012).
Zhang, Z. et al. Neuronal ensembles sufficient for recovery sleep and the sedative actions of α2 adrenergic agonists. Nat. Neurosci. 18, 553–561 (2015).
Steriade, M., Contreras, D., Curró Dossi, R. & Nuñez, A. The slow (< 1 Hz) oscillation in reticular thalamic and thalamocortical neurons: scenario of sleep rhythm generation in interacting thalamic and neocortical networks. J. Neurosci. 13, 3284–3299 (1993).
Contreras, D., Curró Dossi, R. & Steriade, M. Electrophysiological properties of cat reticular thalamic neurones in vivo. J. Physiol. (Lond.) 470, 273–294 (1993).
Peyron, C. et al. Neurons containing hypocretin (orexin) project to multiple neuronal systems. J. Neurosci. 18, 9996–10015 (1998).
Bittencourt, J.C. et al. The melanin-concentrating hormone system of the rat brain: an immuno- and hybridization histochemical characterization. J. Comp. Neurol. 319, 218–245 (1992).
Tortorella, S., Rodrigo-Angulo, M.L., Núñez, A. & Garzón, M. Synaptic interactions between perifornical lateral hypothalamic area, locus coeruleus nucleus and the oral pontine reticular nucleus are implicated in the stage succession during sleep-wakefulness cycle. Front. Neurosci. 7, 216 (2013).
Carter, M.E. et al. Tuning arousal with optogenetic modulation of locus coeruleus neurons. Nat. Neurosci. 13, 1526–1533 (2010).
Vyazovskiy, V.V. et al. Cortical firing and sleep homeostasis. Neuron 63, 865–878 (2009).
McCormick, D.A. & Bal, T. Sleep and arousal: thalamocortical mechanisms. Annu. Rev. Neurosci. 20, 185–215 (1997).
Contreras, D., Timofeev, I. & Steriade, M. Mechanisms of long-lasting hyperpolarizations underlying slow sleep oscillations in cat corticothalamic networks. J. Physiol. (Lond.) 494, 251–264 (1996).
Giber, K. et al. A subcortical inhibitory signal for behavioral arrest in the thalamus. Nat. Neurosci. 18, 562–568 (2015).
Barone, F.C., Cheng, J.T. & Wayner, M.J. Reticular thalamic inhibitory input to lateral hypothalamic neurons: a functional and histochemical determination. Brain Res. Bull. 33, 575–582 (1994).
Carter, M.E. et al. Mechanism for hypocretin-mediated sleep-to-wake transitions. Proc. Natl. Acad. Sci. USA 109, E2635–E2644 (2012).
Carter, M.E., Adamantidis, A., Ohtsu, H., Deisseroth, K. & de Lecea, L. Sleep homeostasis modulates hypocretin-mediated sleep-to-wake transitions. J. Neurosci. 29, 10939–10949 (2009).
Adamantidis, A. et al. Sleep architecture of the melanin-concentrating hormone receptor 1-knockout mice. Eur. J. Neurosci. 27, 1793–1800 (2008).
Sparta, D.R. Construction of implantable optical fibers for long-term optogenetic manipulation of neural circuits. Nat. Protoc. 7, 12–23 (2012).
Wulff, P. et al. Hippocampal theta rhythm and its coupling with gamma oscillations require fast inhibition onto parvalbumin-positive interneurons. Proc. Natl. Acad. Sci. USA 106, 3561–3566 (2009).
Korotkova, T., Fuchs, E.C., Ponomarenko, A., von Engelhardt, J. & Monyer, H. NMDA receptor ablation on parvalbumin-positive interneurons impairs hippocampal synchrony, spatial representations, and working memory. Neuron 68, 557–569 (2010).
Hazan, L., Zugaro, M. & Buzsáki, G. Klusters, NeuroScope, NDManager: a free software suite for neurophysiological data processing and visualization. J. Neurosci. Methods 155, 207–216 (2006).
Harris, K.D., Henze, D.A., Csicsvari, J., Hirase, H. & Buzsáki, G. Accuracy of tetrode spike separation as determined by simultaneous intracellular and extracellular measurements. J. Neurophysiol. 84, 401–414 (2000).
Franken, P., Chollet, D. & Tafti, M. The homeostatic regulation of sleep need is under genetic control. J. Neurosci. 21, 2610–2621 (2001).
Bokil, H., Andrews, P., Kulkarni, J.E., Mehta, S. & Mitra, P.P. Chronux: a platform for analyzing neural signals. J. Neurosci. Methods 192, 146–151 (2010).
Acknowledgements
We thank the Tidis laboratory members for their technical help and comments on a previous version of the manuscript. We thank the laboratory of H.-R. Widmer for the use of the fluorescence microscope, and the animal facility personnel at the Douglas Research Center. Optogenetic plasmids were kindly provided by K. Deisseroth (Stanford University). LT-HSV-GFP was kindly provided by M. Mameli (Institut du Fer a Moulin). C.G.H. was supported by a Strauss Clinical Fellowship. A.A. and T.K. were supported by the Human Frontier Science Program (RGY0076/2012). A.A. was supported by the Douglas Foundation, McGill University, Canadian Fund for Innovation (CFI), Canadian Research Chair (CRC Tier 2), Canadian Institute for Health Research (CIHR) and the Natural Science and Engineering Council of Canada (NSERC), Inselspital, Swiss National Fund and University of Bern. T.K. and A.P. were supported by the Deutsche Forschungsgemeinschaft (DFG; Exc 257 NeuroCure, T.K. and A.P.; SPP1665, A.P.).
Author information
Authors and Affiliations
Contributions
All of the authors designed the experiments. C.G.H. collected and analyzed anatomical, in vitro electrophysiological, behavioral, polysomnographic and optostimulation data. S.J. collected behavioral, polysomnographic and optostimulation preliminary data. M.C.C. and A.P. collected and M.C.C., A.P. and T.K. analyzed in vivo LH and TRN electrophysiological and optostimulation data. All of the authors discussed the results and C.G.H., M.C.C., A.P., T.K. and A.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Identification of LHGABA projections to arousal centers of the brain.
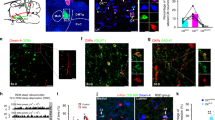
a, Photomicrographs showing antero-posterior coronal brain sections from Tg(VGAT)::IRES-Cre animal transduced with AAVdj-ChETA-EYFP in the LH area. Pictures show dense ChETA-EYFP-expressing GABA neuron terminals within sleep-related brain structures including medial septum (MS; AP +0.6 mm), basal forebrain (BF; AP +0.6 mm), medial forebrain (MBF; AP +0.1 mm), dorsal thalamus (Pvf; AP -0.58 mm), lateral habenula (LHb; AP -1.58 mm), ZI, LH and TRN (AP: -1.06 mm and -1.58 mm, respectively), peri-aqueductal gray (PAG) and ventral tegmental area (VTA, AP -3.08 mm) and the locus coeruleus (LC; AP -5.4 mm). b, Quantification of the AAVdj-DIO-ChETA-YFP genetic targeting selectively to LHGABA cells. Graph shows Mean ± s.e.m. total number of cell bodies within in a field of view at 40X where fluorescence was detected. (AP: -1.45 mm; N = 13 mice). Note the absence of YFP-expressing cells bodies outside the LH area. c, Quantification of genetic targeting to LHGABA cells using fluorescent in situ hybridization detection of GAD-67 gene transcripts and YFP immunoreactivity. Bar graphs show summary data of Mean ± s.e.m. total number of cells that are GAD-67+/YFP+, GAD67-/YFP+, and GAD67+/YFP- (N = 15 sections from N = 4 mice). d, Summary data of Mean ± s.e.m. relative fluorescence (arbitrary unit) of LHGABA terminal density in target areas shown in a-d.
Supplementary Figure 2 Optogenetic control of LHGABA neurons.
a, LHGABA neurons transfected with AAVdj-ChETA-EYFP show typical spiking in response to excitatory current injection (100 pA). b, Brief pulses of 473-nm light-evoked single action potentials in ChETA-expressing LHGABA neurons. c, Quantification of Mean ± s.e.m. voltage depolarization and inward currents of ChETA-expressing LHGABA neurons upon blue light illumination (8 cells from, 8 slices from 8 mice). d, Quantification of Mean ± s.e.m. latency to spike in LHGABA neurons upon 5-ms optical stimulation (left). Note that spontaneous firing of LHGABA cells in vitro is similar to in vivo conditions. e, Group data showing Mean ± s.e.m. ChETA-expressing LHGABA neurons fidelity response to light pulses at frequencies up to 50 Hz (N = 8 cells, 8 slices from N = 8 mice).
Supplementary Figure 3 In vivo characterization of LHGABA neuronal activity during spontaneous sleep-to-wake transitions and upon optical stimulation.
a, Photomicrograph of a representative brain hemi-section showing the track of one of the shanks of the silicon probe and the lesion in the LH (white arrow). b, Schematics for coronal sections at different anterio-posterior levels summarizing the positions of recording sites in the LH (gray dots) where 31 cells in 5 mice were recorded. c, Example of spike waveforms of a putative LHGABA cell, recorded from different octrode-probe channels. d, Representative signal trace (wide-band) showing discharge of a LHGABA cell during a photostimulation experiment (blue lines). e, Firing rates of LHGABA cells before and after wakening (N = 13 cells, P < 0.05, binomial test, 3 s. around NREM sleep-to-Wake transitions).
Supplementary Figure 4 Semi-chronic activation of LHGABA neurons.
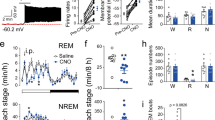
a, Schematic of the genetic targeting of LHGABA neurons. AAVdj-ChETA-YFP or AAVdj-YFP (Control) were infused into the LH of Tg(VGAT)::IRES-Cre mice (left). b, Schematic of semi-chronic optogenetic experimental timeline. Optical stimuli were delivered at 1 or 20 Hz for 10 s every minute for an hour (right). c, d, Mean ± s.e.m. of Total (c) and Episode (d) duration of wake, NREM and REM sleep during 1-h semi-chronic bilateral stimulation in control (white) and ChETA-EYFP (black) animals. Data represent > 10 events from control (N = 7) and ChETA (N = 8) mice. ***, P < 0.0001 using one-way ANOVA followed by multiple comparisons Kruskal-Wallis post hoc test.
Supplementary Figure 5 Optogenetic activation of LepRB-expressing cells in the LH area.
a, Schematic of a coronal hemi-section showing the placement of the optical fibres and the virus injection site (left). Photomicrograph showing the expression ChETA-EYFP in the LH and its projections to the TRN (right). Scale bar represents 200 µm. b, Schematic of NREM sleep-specific stimulation protocol. Tg(LepRb)::IRES-Cre animals transduced with AAV-dj-EYFP (control) or AAV-dj-ChETA-eYFP (ChETA) were stimulated after 10 s of stable NREM sleep onset for 10 sec at 1 or 20 Hz, and continuous illumination. c, Graph representing Mean latencies ± s.e.m. of wake from NREM sleep. Data represent > 10 events from 2 different animals per group. *, P < 0.05, unpaired two-tailed Student’s t-Test.
Supplementary Figure 6 Genetic mapping of monosynaptic LH-TRN circuit.
a, Schematic of a sagittal brain section showing the two injection sites; Tg(VGAT)::IRES-Cre animals were injected with AAV2-dio-mCherry in the LH at 8 weeks followed by an injection of LT-HSV-EGFP lentivirus in the anterior TRN at 10 weeks. b, Photomicrograph of a coronal hemi-section showing cells co-expressing mCherry (red) and retrogradely labeled EGFP (green) in the LH. Scale bar represents 500 μm. c, d Higher magnification photomicrographs of the red box shown in b illustrating the expression of mCherry in LH cells (red, c) and the co-expression of EGFP (merged, d). Scale bar represents 20 µm.
Supplementary Figure 7 Characterization of TRN cell firing in vivo.
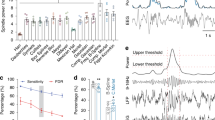
a, Histological reconstruction of a silicon probe placement. Photomicrographs showing the track of the lateral silicon probe shank in the TRN at a higher (left, scale bar: 100µm) and a lower (middle, scale bar 500 µm) magnification (Nissl staining; black arrows point to the electrode track). Right: a lesion, indicated by the white arrow, performed on the medial probe shank (right to the TRN, scale bar 500µm), overlayed with fluorescent image showing DiI (red), applied to this shank before implantation. Recordings in RTN were performed from the lateral shank. b, Schematics of coronal sections at different anterio-posterior levels summarizing the positions of recording sites in the TRN (gray dots) where 70 cells in 3 mice were recorded. c, Example of spike waveforms of a TRN cell, recorded from different octrode-probe channels. d, Average spike waveforms of a TRN cell before (black) and during LHGABA-TRN optostimulation at 20 Hz (blue) and e, the auto-correlogram of this unit. f, Firing rates of individual TRN cells during baseline (X-axis) and optogenetic stimulation of LHGABA-TRN projections (Y axis) at 20 Hz (left), 1 Hz, 5 Hz, and 2-s continuous light (right). g, Coefficient of interspike intervals variance (CV) of TRN cells upon optogenetic activation of LHGABA-TRN circuit (2-s continuous stimulation, P < 0.0001, Kruskal-Wallis test).
Supplementary Figure 8 Optical stimulation of LH GABA terminals in the posterior TRN area.
a, Schematic of the experimental timeline. b, Mean latencies ± s.e.m. of NREM sleep-to-wake transitions upon optical stimulation of LHGABA terminals in the posterior TRN field in control (white; N = 10) and ChETA-EYFP (black; N = 9) animals. c, Mean latencies ± s.e.m. of REM sleep to wake transitions upon optical stimulation of LHGABA terminals in the posterior TRN area in control (white; N = 5) and ChETA-EYFP (black; N = 7) animals. *, P <0.05; ***, P < 0.01; ***, P < 0.0001 using a unpaired two-tailed Student’s t-Test.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 905 kb)
Rights and permissions
About this article
Cite this article
Herrera, C., Cadavieco, M., Jego, S. et al. Hypothalamic feedforward inhibition of thalamocortical network controls arousal and consciousness. Nat Neurosci 19, 290–298 (2016). https://doi.org/10.1038/nn.4209
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.4209
This article is cited by
-
Proteostasis failure exacerbates neuronal circuit dysfunction and sleep impairments in Alzheimer’s disease
Molecular Neurodegeneration (2023)
-
Neuro-orchestration of sleep and wakefulness
Nature Neuroscience (2023)
-
Somatostatin neurons in prefrontal cortex initiate sleep-preparatory behavior and sleep via the preoptic and lateral hypothalamus
Nature Neuroscience (2023)
-
Activation of Ventral Pallidum CaMKIIa-Expressing Neurons Promotes Wakefulness
Neurochemical Research (2023)
-
Alterations in TRN-anterodorsal thalamocortical circuits affect sleep architecture and homeostatic processes in oxidative stress vulnerable Gclm−/− mice
Molecular Psychiatry (2022)