Abstract
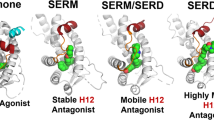
Ligand-binding dynamics control allosteric signaling through the estrogen receptor-α (ERα), but the biological consequences of such dynamic binding orientations are unknown. Here, we compare a set of ER ligands having dynamic binding orientation (dynamic ligands) with a control set of isomers that are constrained to bind in a single orientation (constrained ligands). Proliferation of breast cancer cells directed by constrained ligands is associated with DNA binding, coactivator recruitment and activation of the estrogen-induced gene GREB1, reflecting a highly interconnected signaling network. In contrast, proliferation driven by dynamic ligands is associated with induction of ERα-mediated transcription in a DNA-binding domain (DBD)-dependent manner. Further, dynamic ligands showed enhanced anti-inflammatory activity. The DBD-dependent profile was predictive of these signaling patterns in a larger diverse set of natural and synthetic ligands. Thus, ligand dynamics directs unique signaling pathways and reveals a new role of the DBD in allosteric control of ERα-mediated signaling.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Hilser, V.J., Wrabl, J.O. & Motlagh, H.N. Structural and energetic basis of allostery. Annu. Rev. Biophys. 41, 585–609 (2012).
Carroll, M.J. et al. Evidence for dynamics in proteins as a mechanism for ligand dissociation. Nat. Chem. Biol. 8, 246–252 (2012).
Shiau, A.K. et al. The structural basis of estrogen receptor/coactivator recognition and the antagonism of this interaction by tamoxifen. Cell 95, 927–937 (1998).
Nettles, K.W. et al. CBP Is a dosage-dependent regulator of nuclear factor-κB suppression by the estrogen receptor. Mol. Endocrinol. 22, 263–272 (2008).
Bruning, J.B. et al. Coupling of receptor conformation and ligand orientation determine graded activity. Nat. Chem. Biol. 6, 837–843 (2010).
Hughes, T.S. et al. Ligand and receptor dynamics contribute to the mechanism of graded PPARγ agonism. Structure 20, 139–150 (2012).
Carroll, M.J. et al. Direct detection of structurally resolved dynamics in a multiconformation receptor-ligand complex. J. Am. Chem. Soc. 133, 6422–6428 (2011).
Shang, Y., Hu, X., DiRenzo, J., Lazar, M.A. & Brown, M. Cofactor dynamics and sufficiency in estrogen receptor–regulated transcription. Cell 103, 843–852 (2000).
Meijsing, S.H. et al. DNA binding site sequence directs glucocorticoid receptor structure and activity. Science 324, 407–410 (2009).
McInerney, E.M. & Katzenellenbogen, B.S. Different regions in activation function-1 of the human estrogen receptor required for antiestrogen- and estradiol-dependent transcription activation. J. Biol. Chem. 271, 24172–24178 (1996).
Nettles, K.W. & Greene, G.L. Ligand control of coregulator recruitment to nuclear receptors. Annu. Rev. Physiol. 67, 309–333 (2005).
Darimont, B.D. et al. Structure and specificity of nuclear receptor-coactivator interactions. Genes Dev. 12, 3343–3356 (1998).
Bourguet, W., Ruff, M., Chambon, P., Gronemeyer, H. & Moras, D. Crystal structure of the ligand-binding domain of the human nuclear receptor RXR-α. Nature 375, 377–382 (1995).
Xu, H.E. et al. Structural basis for antagonist-mediated recruitment of nuclear co-repressors by PPARα. Nature 415, 813–817 (2002).
Nettles, K.W. et al. Structural plasticity in the oestrogen receptor ligand-binding domain. EMBO Rep. 8, 563–568 (2007); erratum 8, 610 (2007).
Chadwick, C.C. et al. Identification of pathway-selective estrogen receptor ligands that inhibit NF-κB transcriptional activity. Proc. Natl. Acad. Sci. USA 102, 2543–2548 (2005).
Nettles, K.W. et al. Allosteric control of ligand selectivity between estrogen receptors α and β: implications for other nuclear receptors. Mol. Cell 13, 317–327 (2004).
Laughlin, J.D. et al. Structural mechanisms of allostery and autoinhibition in JNK family kinases. Structure 20, 2174–2184 (2012).
Shang, Y. & Brown, M. Molecular determinants for the tissue specificity of SERMs. Science 295, 2465–2468 (2002).
Sicheri, F. & Kuriyan, J. Structures of Src-family tyrosine kinases. Curr. Opin. Struct. Biol. 7, 777–785 (1997).
Heery, D.M., Kalkhoven, E., Hoare, S. & Parker, M.G. A signature motif in transcriptional co-activators mediates binding to nuclear receptors. Nature 387, 733–736 (1997).
Berry, M., Metzger, D. & Chambon, P. Role of the two activating domains of the oestrogen receptor in the cell-type and promoter-context dependent agonistic activity of the anti-oestrogen 4-hydroxytamoxifen. EMBO J. 9, 2811–2818 (1990).
Pham, T.A. et al. Antiestrogen can establish nonproductive receptor complexes and alter chromatin structure at target enhancers. Proc. Natl. Acad. Sci. USA 88, 3125–3129 (1991).
Lupien, M. et al. Raloxifene and ICI182,780 increase estrogen receptor-α association with a nuclear compartment via overlapping sets of hydrophobic amino acids in activation function 2 helix 12. Mol. Endocrinol. 21, 797–816 (2007).
Planas-Silva, M.D., Shang, Y., Donaher, J.L., Brown, M. & Weinberg, R.A. AIB1 enhances estrogen-dependent induction of cyclin D1 expression. Cancer Res. 61, 3858–3862 (2001).
Xu, J., Wu, R.C. & O'Malley, B.W. Normal and cancer-related functions of the p160 steroid receptor co-activator (SRC) family. Nat. Rev. Cancer 9, 615–630 (2009).
Rae, J.M. et al. GREB 1 is a critical regulator of hormone dependent breast cancer growth. Breast Cancer Res. Treat. 92, 141–149 (2005).
Eeckhoute, J., Carroll, J.S., Geistlinger, T.R., Torres-Arzayus, M.I. & Brown, M. A cell-type-specific transcriptional network required for estrogen regulation of cyclin D1 and cell cycle progression in breast cancer. Genes Dev. 20, 2513–2526 (2006).
Wang, Y.H. et al. Knockdown of c-Myc expression by RNAi inhibits MCF-7 breast tumor cells growth in vitro and in vivo. Breast Cancer Res. 7, R220–R228 (2005).
Wang, C. et al. Estrogen induces c-myc gene expression via an upstream enhancer activated by the estrogen receptor and the AP-1 transcription factor. Mol. Endocrinol. 25, 1527–1538 (2011).
Powell, E. et al. Identification of estrogen receptor dimer selective ligands reveals growth-inhibitory effects on cells that co-express ERα and ERβ. PLoS ONE 7, e30993 (2012).
Rae, J.M. et al. GREB1 is a novel androgen-regulated gene required for prostate cancer growth. Prostate 66, 886–894 (2006).
Loven, M.A., Likhite, V.S., Choi, I. & Nardulli, A.M. Estrogen response elements alter coactivator recruitment through allosteric modulation of estrogen receptor β conformation. J. Biol. Chem. 276, 45282–45288 (2001).
Hall, J.M., McDonnell, D.P. & Korach, K.S. Allosteric regulation of estrogen receptor structure, function, and coactivator recruitment by different estrogen response elements. Mol. Endocrinol. 16, 469–486 (2002).
Wang, J.C. et al. Novel arylpyrazole compounds selectively modulate glucocorticoid receptor regulatory activity. Genes Dev. 20, 689–699 (2006).
Bapat, A.R. & Frail, D.E. Full-length estrogen receptor α and its ligand-binding domain adopt different conformations upon binding ligand. J. Steroid Biochem. Mol. Biol. 86, 143–149 (2003).
Kong, E.H. et al. Delineation of a unique protein-protein interaction site on the surface of the estrogen receptor. Proc. Natl. Acad. Sci. USA 102, 3593–3598 (2005).
Chandra, V. et al. Structure of the intact PPAR-γ-RXR- nuclear receptor complex on DNA. Nature 456, 350–356 (2008).
Rochel, N. et al. Common architecture of nuclear receptor heterodimers on DNA direct repeat elements with different spacings. Nat. Struct. Mol. Biol. 18, 564–570 (2011).
Likhite, V.S., Cass, E.I., Anderson, S.D., Yates, J.R. & Nardulli, A.M. Interaction of estrogen receptor α with 3-methyladenine DNA glycosylase modulates transcription and DNA repair. J. Biol. Chem. 279, 16875–16882 (2004).
Nwachukwu, J.C. & Nettles, K.W. The nuclear receptor signalling scaffold: insights from full-length structures. EMBO J. 31, 251–253 (2012).
Schultz-Norton, J.R., Ziegler, Y.S. & Nardulli, A.M. ERα-associated protein networks. Trends Endocrinol. Metab. 22, 124–129 (2011).
Hammes, S.R. & Levin, E.R. Minireview: recent advances in extranuclear steroid receptor actions. Endocrinology 152, 4489–4495 (2011).
Zhan, Y.Y. et al. The orphan nuclear receptor Nur77 regulates LKB1 localization and activates AMPK. Nat. Chem. Biol. 8, 807–904 (2012).
Sun, Z. et al. Inhibition of β-catenin signaling by nongenomic action of orphan nuclear receptor Nur77. Oncogene 31, 2653–2667 (2012).
National Research Council (U.S.). A New Biology for the 21st Century (eds. Connelly, T. & Sharp, P.) 12 (National Academies Press, Washington, D.C., 2009).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Schüttelkopf, A.W. & van Aalten, D.M. PRODRG: a tool for high-throughput crystallography of protein-ligand complexes. Acta Crystallogr. D Biol. Crystallogr. 60, 1355–1363 (2004).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
McNicholas, S., Potterton, E., Wilson, K.S. & Noble, M.E. Presenting your structures: the CCP4mg molecular-graphics software. Acta Crystallogr. D Biol. Crystallogr. 67, 386–394 (2011).
Acknowledgements
We thank J.L. Cleveland for critically evaluating the manuscript. This research was supported by the US National Institutes of Health (PHS 5R37 DK015556 to J.A.K.; 5R33CA132022 and 5R01DK077085 to K.W.N.). S.S. is supported by the Frenchman's Creek Women for Cancer Research.
Author information
Authors and Affiliations
Contributions
S.S. and J.C.N. designed and performed experiments and wrote the manuscript; A.A.P., V.C., J.N. and T.S.H. performed experiments; D.J.K. designed experiments; and J.A.K. and K.W.N. designed experiments and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Note (PDF 7674 kb)
Rights and permissions
About this article
Cite this article
Srinivasan, S., Nwachukwu, J., Parent, A. et al. Ligand-binding dynamics rewire cellular signaling via estrogen receptor-α. Nat Chem Biol 9, 326–332 (2013). https://doi.org/10.1038/nchembio.1214
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1214
This article is cited by
-
Chemical systems biology reveals mechanisms of glucocorticoid receptor signaling
Nature Chemical Biology (2021)
-
Sirtuins transduce STACs signals through steroid hormone receptors
Scientific Reports (2020)
-
Full antagonism of the estrogen receptor without a prototypical ligand side chain
Nature Chemical Biology (2017)
-
Competitive molecular docking approach for predicting estrogen receptor subtype α agonists and antagonists
BMC Bioinformatics (2014)