Abstract
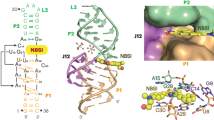
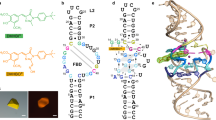
Spinach is an in vitro–selected RNA aptamer that binds a GFP-like ligand and activates its green fluorescence. Spinach is thus an RNA analog of GFP and has potentially widespread applications for in vivo labeling and imaging. We used antibody-assisted crystallography to determine the structures of Spinach both with and without bound fluorophore at 2.2-Å and 2.4-Å resolution, respectively. Spinach RNA has an elongated structure containing two helical domains separated by an internal bulge that folds into a G-quadruplex motif of unusual topology. The G-quadruplex motif and adjacent nucleotides comprise a partially preformed binding site for the fluorophore. The fluorophore binds in a planar conformation and makes extensive aromatic stacking and hydrogen bond interactions with the RNA. Our findings provide a foundation for structure-based engineering of new fluorophore-binding RNA aptamers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Yang, F., Moss, L.G. & Phillips, G.N.J. The molecular structure of green fluorescent protein. Nat. Biotechnol. 14, 1246–1251 (1996).
Tsien, R.Y. The green fluorescence protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Shaner, N.C., Steinbach, P.A. & Tsien, R.Y. A guide to choosing fluorescent proteins. Nat. Methods 2, 905–909 (2005).
Shu, X. et al. Mammalian expression of infrared fluorescent proteins engineered from a bacterial phytochrome. Science 324, 804–807 (2009).
Paige, J.S., Wu, K.Y. & Jaffrey, S.R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Strack, R.L., Disney, M.D. & Jaffrey, S.R. A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat–containing RNA. Nat. Methods 10, 1219–1224 (2013).
Strack, R.L. & Jaffrey, S.R. New approaches for sensing metabolites and proteins in live cells using RNA. Curr. Opin. Chem. Biol. 17, 651–655 (2013).
Paige, J.S., Nguyen-Duc, T., Song, W. & Jaffrey, S.R. Fluorescence imaging of cellular metabolites with RNA. Science 335, 1194 (2012).
Kellenberger, C.A., Wilson, S.C., Sales-Lee, J. & Hammond, M.C. RNA-based fluorescent biosensors for live cell imaging of second messengers cyclic di-GMP and cyclic AMP-GMP. J. Am. Chem. Soc. 135, 4906–4909 (2013).
Pothoulakis, G., Ceroni, F., Reeve, B. & Ellis, T. The Spinach RNA aptamer as a characterization tool for synthetic biology. ACS Synth. Biol. 3, 182–187 (2013).
Ye, J.D. et al. Synthetic antibodies for specific recognition and crystallization of structured RNA. Proc. Natl. Acad. Sci. USA 105, 82–87 (2008).
Koldobskaya, Y. et al. A portable RNA sequence whose recognition by a synthetic antibody facilitates structural determination. Nat. Struct. Mol. Biol. 18, 100–106 (2011).
Lim, K.W. & Phan, A.T. Structural basis of DNA quadruplex-duplex junction formation. Angew. Chem. Int. Ed. Engl. 52, 8566–8569 (2013).
Zuker, M. Mfold web server for nucleic acid folding and hybridization prediction. Nucleic Acids Res. 31, 3406–3415 (2003).
Adrian, M., Heddi, B. & Phan, A.T. NMR spectroscopy of G-quadruplexes. Methods 57, 11–24 (2012).
Burge, S., Parkinson, G.N., Hazel, P., Todd, A.K. & Neidle, S. Quadruplex DNA: sequence, topology and structure. Nucleic Acids Res. 34, 5402–5415 (2006).
Mullen, M.A., Assmann, S.M. & Bevilacqua, P.C. Toward a digital gene response: RNA G-quadruplexes with fewer quartets fold with higher cooperativity. J. Am. Chem. Soc. 134, 812–815 (2012).
Harding, M.M. Small revisions to predicted distances around metal sites in proteins. Acta Crystallogr. D Biol. Crystallogr. 62, 678–682 (2006).
Hud, N.V., Schultze, P., Sklenar, V. & Feigon, J. Binding sites and dynamics of ammonium ions in a telomere repeat DNA quadruplex. J. Mol. Biol. 285, 233–243 (1999).
Guédin, A., Gros, J., Alberti, P. & Mergny, J.L. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 38, 7858–7868 (2010).
Wang, P. et al. Photochemical properties of Spinach and its use in selective imaging. Chem. Sci. 4, 2865–2873 (2013).
Han, K.Y., Leslie, B.J., Fei, J., Zhang, J. & Ha, T. Understanding the photophysics of the Spinach−DFHBI RNA aptamer−fluorogen complex to improve live-cell RNA imaging. J. Am. Chem. Soc. 135, 19033–19038 (2013).
Song, W., Strack, R.L., Svensen, N. & Jaffrey, S.R. Plug-and-play fluorophores extend the spectral properties of Spinach. J. Am. Chem. Soc. 136, 1198–1201 (2014).
Royant, A. & Noirclerc-Savoye, M. Stabilizing role of glutamic acid 222 in the structure of enhanced green fluorescent protein. J. Struct. Biol. 174, 385–390 (2011).
Wachter, R.M., Elsliger, M.A., Kallio, K., Hanson, G.T. & Remington, S.J. Structural basis of spectral shifts in the yellow-emission variants of green fluorescent protein. Structure 6, 1267–1277 (1998).
Yang, P., De Cian, A., Teulade-Fichou, M.P., Mergny, J.L. & Monchaud, D. Engineering bisquinolinium/thiazole orange conjugates for fluorescent sensing of G-quadruplex DNA. Angew. Chem. Int. Ed. Engl. 48, 2188–2191 (2009).
Neidle, S. & Parkinson, G. Telomere maintenance as a target for anticancer drug discovery. Nat. Rev. Drug Discov. 1, 383–393 (2002).
Liberman, J.A. & Wedekind, J.E. Riboswitch structure in the ligand-free state. Wiley Interdiscip. Rev. RNA 3, 369–384 (2012).
Zaug, A.J., Podell, E.R. & Cech, T.R. Human POT1 disrupts telomeric G-quadruplexes allowing telomerase extension in vitro. Proc. Natl. Acad. Sci. USA 102, 10864–10869 (2005).
Sen, D. & Gilbert, W. Formation of parallel four-stranded complexes by guanine-rich motifs in DNA and its implications for meiosis. Nature 334, 364–366 (1988).
Macaya, R.F., Schultze, P., Smith, F.W., Roet, J.A. & Feigon, J. Thrombin-binding DNA aptamer forms a unimolecular quadruplex structure in solution. Proc. Natl. Acad. Sci. USA 90, 3745–3749 (1993).
Zhang, X.B., Kong, R.M. & Lu, Y. Metal ion sensors based on DNAzymes and related DNA molecules. Annu. Rev. Anal. Chem. (Palo Alto Calif.) 4, 105–128 (2011).
Weiss, S. et al. RNA aptamers specifically interact with the prion protein PrP. J. Virol. 71, 8790–8797 (1997).
Mashima, T., Matsugami, A., Nishikawa, F., Nishikawa, S. & Katahira, M. Unique quadruplex structure and interaction of an RNA aptamer against bovine prion protein. Nucleic Acids Res. 37, 6249–6258 (2009).
Phan, A.T. et al. Structure-function studies of FMRP RGG peptide recognition of an RNA duplex-quadruplex junction. Nat. Struct. Mol. Biol. 18, 796–804 (2011).
Bochman, M.L., Paeschke, K. & Zakian, V.A. DNA secondary structures: stability and function of G-quadruplex structures. Nat. Rev. Genet. 13, 770–780 (2012).
Bif, G., Tannahill, D., McCafferty, J. & Balasubramanian, S. Quantitative visualization of DNA G-quadruplex structures in human cells. Nat. Chem. 5, 182–186 (2013).
Kao, C., Rüdisser, S. & Zheng, M. A simple and efficient method to transcribe RNAs with reduced 3′ heterogeneity. Methods 23, 201–205 (2001).
Kim, I., McKenna, S.A., Viani Puglisi, E. & Puglisi, J.D. Rapid purification of RNAs using fast performance liquid chromatography (FPLC). RNA 13, 289–294 (2007).
Wilkinson, K.A., Merino, E.J. & Weeks, K.M. Selective 2′-hydroxyl acylation analyzed by primer extension (SHAPE): quantitative RNA structure analysis at single nucleotide resolution. Nat. Protoc. 1, 1610–1616 (2006).
Chayen, R. Rigorous filtration for protein crystallography. J. Appl. Crystallogr. 42, 743–744 (2009).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Keating, K.S. & Pyle, A.M. Semiautomated model building for RNA crystallography using a directed rotameric approach. Proc. Natl. Acad. Sci. USA 107, 8177–8182 (2010).
Chou, F.C., Sripakdeevong, P., Dibrov, S.M., Hermann, T. & Das, R. Correcting pervasive errors in RNA crystallography through enumerative structure prediction. Nat. Methods 10, 74–76 (2013).
Afonine, P.V., Grosse-Kunstleve, R.W. & Adams, P.D. The Phenix refinement framework. CCP4 Newslett. Number 42, contribution 8 (2005).
Terwilliger, T.C., Adams, P.D., Moriarty, N.W. & Cohn, J.D. Ligand identification using electron-density map correlations. Acta Crystallogr. D Biol. Crystallogr. 63, 101–107 (2007).
Haddadian, E.J. et al. Automated real-space refinement of protein structures using a realistic backbone move set. Biophys. J. 101, 899–909 (2011).
Stahley, M.R., Adams, P.L., Wang, J. & Strobel, S.A. Structural metals in the Group I Intron: a ribozyme with a multiple metal ion core. J. Mol. Biol. 372, 89–102 (2007).
Vaguine, A.A., Richelle, J. & Wodak, S.J. SFCHECK: a unified set of procedure for evaluating the quality of macromolecular structure-factor data and their agreement with atomic model. Acta Crystallogr. D Biol. Crystallogr. 55, 191–205 (1999).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Wadley, L.M. & Pyle, A.M. The identification of novel RNA structural motifs using COMPADRES: an automated approach to structural discovery. Nucleic Acids Res. 32, 6650–6659 (2004).
Das, R. SAFA: semi-automated footprinting analysis software for high-throughput quantification of nucleic acid footprinting experiments. RNA 11, 344–354 (2005).
Hura, G.L. et al. Robust, high-throughput solution structural analyses by small angle X-ray scattering (SAXS). Nat. Methods 6, 606–612 (2009).
Svergun, D.I., Barberato, C. & Koch, M.H.J. CRYSOL—a program to evaluate X-ray solution scattering of biological macromolecules from atomic coordinates. J. Appl. Crystallogr. 28, 768–773 (1995).
Acknowledgements
We are grateful to I.M. Steele for assistance in determining the structure of unbound DFHBI. We thank L. Zhang for the advice on structure and model building. We thank J.R. Fuller for refinement software support. We also thank F.C. Chou and R. Das for their aid with ERRASER software and K.N. Dyer, T.R. Sosnick and J.R. Hinshaw for the help with SAXS experiments. We thank members of the Piccirilli group, J.P. Staley and D.M.J. Lilley for helpful discussions and comments on the manuscript. The work is supported by US National Institutes of Health (NIH) grants R01-AI081987, NIH training grant T32GM008720 (to M.E.E.) and R01-GM102489 (to J.A.P.), NIH training grant T32GM007183 (to N.B.S.) and US National Institute of General Medical Sciences Medical Scientist National Research Service Award no. 5 T32GM07281 (to Y.K.). This work is based on research conducted at the Advanced Photon Source on the Northeastern Collaborative Access Team beamline 24-ID-C&E, GM/CA beamline 23-ID-D and Advanced Light Source beamline 12.3.1 SIBYLS, all supported by USA Department of Energy.
Author information
Authors and Affiliations
Contributions
H.H. and J.A.P. designed the project; H.H. conducted most of the biochemical and biophysical assays and crystallography; N.B.S. and P.A.R. made essential contributions to crystallography; N.-S.L. synthesized DFHBI and analogs; S.A.S. and M.E.E. constructed and characterized truncation mutants; Y.K. developed the Fab BL3-6 chaperone; and J.A.P. provided overall project supervision. The manuscript was prepared by H.H., P.A.R. and J.A.P.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Note, Supplementary Figures 1–21 and Supplementary Tables 1–4. (PDF 3827 kb)
Supplementary Data Set
checkCIF/PLATON report (PDF 456 kb)
Rights and permissions
About this article
Cite this article
Huang, H., Suslov, N., Li, NS. et al. A G-quadruplex–containing RNA activates fluorescence in a GFP-like fluorophore. Nat Chem Biol 10, 686–691 (2014). https://doi.org/10.1038/nchembio.1561
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1561
This article is cited by
-
Imaging the dynamics of messenger RNA with a bright and stable green fluorescent RNA
Nature Chemical Biology (2024)
-
Intracellular RNA and DNA tracking by uridine-rich internal loop tagging with fluorogenic bPNA
Nature Communications (2023)
-
Crystal structure of a highly conserved enteroviral 5′ cloverleaf RNA replication element
Nature Communications (2023)
-
Ultrasensitive and highly selective detection of strontium ions
Nature Sustainability (2023)
-
The fluorescent aptamer Squash extensively repurposes the adenine riboswitch fold
Nature Chemical Biology (2022)