Abstract
Asymmetric cell divisions allow stem cells to balance proliferation and differentiation. During embryogenesis, murine epidermis expands rapidly from a single layer of unspecified basal layer progenitors to a stratified, differentiated epithelium. Morphogenesis involves perpendicular (asymmetric) divisions and the spindle orientation protein LGN, but little is known about how the apical localization of LGN is regulated. Here, we combine conventional genetics and lentiviral-mediated in vivo RNAi to explore the functions of the LGN-interacting proteins Par3, mInsc and Gαi3. Whereas loss of each gene alone leads to randomized division angles, combined loss of Gnai3 and mInsc causes a phenotype of mostly planar divisions, akin to loss of LGN. These findings lend experimental support for the hitherto untested model that Par3–mInsc and Gαi3 act cooperatively to polarize LGN and promote perpendicular divisions. Finally, we uncover a developmental switch between delamination-driven early stratification and spindle-orientation-dependent differentiation that occurs around E15, revealing a two-step mechanism underlying epidermal maturation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Clayton, E. et al. A single type of progenitor cell maintains normal epidermis. Nature 446, 185–189 (2007).
Lechler, T. & Fuchs, E. Asymmetric cell divisions promote stratification and differentiation of mammalian skin. Nature 437, 275–280 (2005).
Smart, I. H. Variation in the plane of cell cleavage during the process of stratification in the mouse epidermis. Br. J. Dermatol. 82, 276–282 (1970).
Williams, S. E. & Fuchs, E. Oriented divisions, fate decisions. Curr. Opin. Cell Biol. 25, 749–758 (2013).
Knoblich, J. A. Mechanisms of asymmetric stem cell division. Cell 132, 583–597 (2008).
Siller, K. H. & Doe, C. Q. Spindle orientation during asymmetric cell division. Nat. Cell Biol. 11, 365–374 (2009).
Cowan, C. R. & Hyman, A. A. Asymmetric cell division in C. elegans: cortical polarity and spindle positioning. Annu. Rev. Cell Dev. Biol. 20, 427–453 (2004).
Martin-Belmonte, F. & Perez-Moreno, M. Epithelial cell polarity, stem cells and cancer. Nat. Rev. Cancer 12, 23–38 (2012).
Knoblich, J. A. Asymmetric cell division: recent developments and their implications for tumour biology. Nat. Rev. Mol. Cell Biol. 11, 849–860 (2010).
Gonzalez, C. Spindle orientation, asymmetric division and tumour suppression in Drosophila stem cells. Nat. Rev. Genet. 8, 462–472 (2007).
Homem, C. C. & Knoblich, J. A. Drosophila neuroblasts: a model for stem cell biology. Development 139, 4297–4310 (2012).
Prehoda, K. E. Polarization of Drosophila neuroblasts during asymmetric division. Cold Spring Harb. Perspect. Biol. 1, a001388 (2009).
Culurgioni, S., Alfieri, A., Pendolino, V., Laddomada, F. & Mapelli, M. Inscuteable and NuMA proteins bind competitively to Leu-Gly-Asn repeat-enriched protein (LGN) during asymmetric cell divisions. Proc. Natl Acad. Sci. USA 108, 20998–21003 (2011).
Mauser, J. F. & Prehoda, K. E. Inscuteable regulates the Pins-Mud spindle orientation pathway. PLoS ONE 7, e29611 (2012).
Yuzawa, S., Kamakura, S., Iwakiri, Y., Hayase, J. & Sumimoto, H. Structural basis for interaction between the conserved cell polarity proteins Inscuteable and Leu-Gly-Asn repeat-enriched protein (LGN). Proc. Natl Acad. Sci. USA 108, 19210–19215 (2011).
Zhu, J. et al. LGN/mInsc and LGN/NuMA complex structures suggest distinct functions in asymmetric cell division for the Par3/mInsc/LGN and Gαi/LGN/NuMA pathways. Mol. Cell 43, 418–431 (2011).
Williams, S. E., Beronja, S., Pasolli, H. A. & Fuchs, E. Asymmetric cell divisions promote Notch-dependent epidermal differentiation. Nature 470, 353–358 (2011).
El-Hashash, A. H. & Warburton, D. Cell polarity and spindle orientation in the distal epithelium of embryonic lung. Dev. Dyn. 240, 441–445 (2011).
Konno, D. et al. Neuroepithelial progenitors undergo LGN-dependent planar divisions to maintain self-renewability during mammalian neurogenesis. Nat. Cell Biol. 10, 93–101 (2008).
Peyre, E. et al. A lateral belt of cortical LGN and NuMA guides mitotic spindle movements and planar division in neuroepithelial cells. J. Cell Biol. 193, 141–154 (2011).
Luxenburg, C., Amalia Pasolli, H., Williams, S. E. & Fuchs, E. Developmental roles for Srf, cortical cytoskeleton and cell shape in epidermal spindle orientation. Nat. Cell Biol. 13, 203–214 (2011).
Poulson, N. D. & Lechler, T. Robust control of mitotic spindle orientation in the developing epidermis. J. Cell Biol. 191, 915–922 (2010).
Gladden, A. B., Hebert, A. M., Schneeberger, E. E. & McClatchey, A. I. The NF2 tumor suppressor, Merlin, regulates epidermal development through the establishment of a junctional polarity complex. Dev. Cell 19, 727–739 (2010).
Watt, F. M. & Green, H. Stratification and terminal differentiation of cultured epidermal cells. Nature 295, 434–436 (1982).
Dowling, J., Yu, Q. C. & Fuchs, E. Beta4 integrin is required for hemidesmosome formation, cell adhesion and cell survival. J. Cell Biol. 134, 559–572 (1996).
Beronja, S. & Fuchs, E. RNAi-mediated gene function analysis in skin. Methods Mol. Biol. 961, 351–361 (2013).
Beronja, S., Livshits, G., Williams, S. & Fuchs, E. Rapid functional dissection of genetic networks via tissue-specific transduction and RNAi in mouse embryos. Nat. Med. 16, 821–827 (2010).
Snippert, H. J. et al. Intestinal crypt homeostasis results from neutral competition between symmetrically dividing Lgr5 stem cells. Cell 143, 134–144 (2010).
Kawaguchi, D., Furutachi, S., Kawai, H., Hozumi, K. & Gotoh, Y. Dll1 maintains quiescence of adult neural stem cells and segregates asymmetrically during mitosis. Nat. Commun. 4, 1880 (2013).
El-Hashash, A. H. et al. Eya1 controls cell polarity, spindle orientation, cell fate and Notch signaling in distal embryonic lung epithelium. Development 138, 1395–1407 (2011).
Bultje, R. S. et al. Mammalian Par3 regulates progenitor cell asymmetric division via notch signaling in the developing neocortex. Neuron 63, 189–202 (2009).
Demehri, S. et al. Notch-deficient skin induces a lethal systemic B-lymphoproliferative disorder by secreting TSLP, a sentinel for epidermal integrity. PLoS Biol. 6, e123 (2008).
Schaefer, M., Shevchenko, A. & Knoblich, J. A. A protein complex containing Inscuteable and the Gα-binding protein Pins orients asymmetric cell divisions in Drosophila. Curr. Biol. 10, 353–362 (2000).
Yu, F., Morin, X., Cai, Y., Yang, X. & Chia, W. Analysis of partner of inscuteable, a novel player of Drosophila asymmetric divisions, reveals two distinct steps in inscuteable apical localization. Cell 100, 399–409 (2000).
Parmentier, M. L. et al. Rapsynoid/partner of inscuteable controls asymmetric division of larval neuroblasts in Drosophila. J. Neurosci. 20, RC84 (2000).
Postiglione, M. P. et al. Mouse inscuteable induces apical-basal spindle orientation to facilitate intermediate progenitor generation in the developing neocortex. Neuron 72, 269–284 (2011).
Schober, M., Schaefer, M. & Knoblich, J. A. Bazooka recruits Inscuteable to orient asymmetric cell divisions in Drosophila neuroblasts. Nature 402, 548–551 (1999).
Wodarz, A., Ramrath, A., Kuchinke, U. & Knust, E. Bazooka provides an apical cue for Inscuteable localization in Drosophila neuroblasts. Nature 402, 544–547 (1999).
Hao, Y. et al. Par3 controls epithelial spindle orientation by aPKC-mediated phosphorylation of apical Pins. Curr. Biol. 20, 1809–1818 (2010).
Hirose, T. et al. PAR3 is essential for cyst-mediated epicardial development by establishing apical cortical domains. Development 133, 1389–1398 (2006).
Gao, L., Macara, I. G. & Joberty, G. Multiple splice variants of Par3 and of a novel related gene, Par3L, produce proteins with different binding properties. Gene 294, 99–107 (2002).
Kohjima, M. et al. PAR3beta, a novel homologue of the cell polarity protein PAR3, localizes to tight junctions. Biochem. Biophys. Res. Commun. 299, 641–646 (2002).
Du, Q. & Macara, I. G. Mammalian Pins is a conformational switch that links NuMA to heterotrimeric G proteins. Cell 119, 503–516 (2004).
Pan, Z. et al. An autoinhibited conformation of LGN reveals a distinct interaction mode between goloco motifs and TPR motifs. Structure 21, 1007–1017 (2013).
Gotta, M. & Ahringer, J. Distinct roles for Gα and Gβγ in regulating spindle position and orientation in Caenorhabditis elegans embryos. Nat. Cell Biol. 3, 297–300 (2001).
Colombo, K. et al. Translation of polarity cues into asymmetric spindle positioning in Caenorhabditis elegans embryos. Science 300, 1957–1961 (2003).
Srinivasan, D. G., Fisk, R. M., Xu, H. & van den Heuvel, S. A complex of LIN-5 and GPR proteins regulates G protein signaling and spindle function in C elegans. Genes Dev. 17, 1225–1239 (2003).
Cai, Y., Yu, F., Lin, S., Chia, W. & Yang, X. Apical complex genes control mitotic spindle geometry and relative size of daughter cells in Drosophila neuroblast and pI asymmetric divisions. Cell 112, 51–62 (2003).
Yu, F., Cai, Y., Kaushik, R., Yang, X. & Chia, W. Distinct roles of Gαi and Gbeta13F subunits of the heterotrimeric G protein complex in the mediation of Drosophila neuroblast asymmetric divisions. J. Cell Biol. 162, 623–633 (2003).
Sanada, K. & Tsai, L. H. G protein betagamma subunits and AGS3 control spindle orientation and asymmetric cell fate of cerebral cortical progenitors. Cell 122, 119–131 (2005).
Schaefer, M., Petronczki, M., Dorner, D., Forte, M. & Knoblich, J. A. Heterotrimeric G proteins direct two modes of asymmetric cell division in the Drosophila nervous system. Cell 107, 183–194 (2001).
Kriegstein, A. & Alvarez-Buylla, A. The glial nature of embryonic and adult neural stem cells. Annu. Rev. Neurosci. 32, 149–184 (2009).
Shitamukai, A. & Matsuzaki, F. Control of asymmetric cell division of mammalian neural progenitors. Dev. Growth Differ. 54, 277–286 (2012).
Morin, X., Jaouen, F. & Durbec, P. Control of planar divisions by the G-protein regulator LGN maintains progenitors in the chick neuroepithelium. Nat. Neurosci. 10, 1440–1448 (2007).
Lancaster, M. A. & Knoblich, J. A. Spindle orientation in mammalian cerebral cortical development. Curr. Opin. Neurobiol. 22, 737–746 (2012).
Goulas, S., Conder, R. & Knoblich, J. A. The par complex and integrins direct asymmetric cell division in adult intestinal stem cells. Cell Stem Cell 11, 529–540 (2012).
Niessen, M. T. et al. aPKClambda controls epidermal homeostasis and stem cell fate through regulation of division orientation. J. Cell Biol. 202, 887–900 (2013).
Morin, X. & Bellaiche, Y. Mitotic spindle orientation in asymmetric and symmetric cell divisions during animal development. Dev. Cell 21, 102–119 (2011).
Coumailleau, F., Furthauer, M., Knoblich, J. A. & Gonzalez-Gaitan, M. Directional Delta and Notch trafficking in Sara endosomes during asymmetric cell division. Nature 458, 1051–1055 (2009).
Mummery-Widmer, J. L. et al. Genome-wide analysis of Notch signalling in Drosophila by transgenic RNAi. Nature 458, 987–992 (2009).
Rompolas, P. et al. Live imaging of stem cell and progeny behaviour in physiological hair-follicle regeneration. Nature 487, 496–499 (2012).
Iden, S. et al. Tumor type-dependent function of the par3 polarity protein in skin tumorigenesis. Cancer Cell 22, 389–403 (2012).
Tanigaki, K. et al. Notch-RBP-J signaling is involved in cell fate determination of marginal zone B cells. Nat. Immunol. 3, 443–450 (2002).
Nowak, J. A. & Fuchs, E. Isolation and culture of epithelial stem cells. Methods Mol. Biol. 482, 215–232 (2009).
Acknowledgements
We thank N. Stokes, D. Oristian and A. Aldeguer (Fuchs laboratory) and T. Anthony Curtis (Williams laboratory), for their expert technical assistance. We thank K. Byrd, K. Lough, and members of the Williams and Fuchs laboratories for critical reading of the manuscript and K. Lough for valuable input into the model presented in Fig. 8. We are grateful to S. Ohno and T. Hirose (both at Yokohama City University of Medicine, Japan) for sharing the Pard3 floxed mouse line. S.E.W. was supported by an American Cancer Society postdoctoral fellowship and E.F. is an investigator in the Howard Hughes Medical Institute. Work in the Fuchs laboratory was supported by a grant from the National Institutes of Health (E.F. R37-27883).
Author information
Authors and Affiliations
Contributions
S.E.W. designed and conducted experiments and analysed the data under the supervision of E.F. L.A.R. performed the imaging and analysis for the lineage tracing experiments. M.P.P. and J.A.K. provided mInsc mice before publication. S.E.W. and E.F. wrote the manuscript. All authors critically read and contributed to the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 4 Maturation of differentiation markers in developing epidermis.
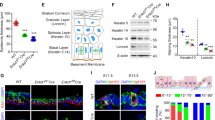
(a) Beginning around E13.5, at the stage when suprabasal cells (except the periderm) are absent, sporadic basal cells coexpress the spinous keratin, K10, along with basal keratins K5 and K14 (not shown). Note also that at this age, the basement membrane marker and hemidesmosome constituent β4-integrin is broadly and diffusely expressed throughout basal cells at E13.5, while it progressively becomes more basally restricted at later ages. (b) Sections of wild-type E14.5 back skin from anterior (less differentiated) to posterior (more differentiated). In anterior regions of single-layered epidermis, K10 and K5 are broadly coexpressed, while β4-integrin remains diffuse. In areas where β4-integrin begins to show some apical enrichment and suprabasal cells are present, K10 becomes more restricted to suprabasal cells, though the basal keratin K5 is diffusely coexpressed there. In posterior regions, the segregation of K10 and K5 becomes more apparent. (c) At E15.5, β4-integrin becomes more restricted to the epidermal-dermal boundary, while K10 and K5 are expressed in opposing domains. An exception is the appearance of sporadic cells positioned in the basal layer which coexpress K10 and K4 (arrows). We suggest that these are cells undergoing differentiation by delamination rather than asymmetric cell division.
Supplementary Figure 5 Both Notch and LGN are dispensable for early stage stratification.
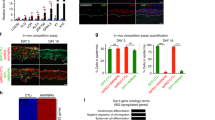
(a) (Top left) Schematic of lentiviral Notch reporter construct. Nuclear H2B-RFP is expressed under a constitutive reporter to mark cells transduced with the reporter construct, while cytosolic GFP reveals cells in which Notch signalling has been activated. At E17.5 (right), robust Notch/GFP+ cells are observed in suprabasal (K10+, blue) layers. In contrast, at E15.5 (bottom left), few cells show detectable Notch activity, and those that do are weakly positive, and appear in both basal (arrow) and suprabasal (arrowhead) layers. (b) Quantification of the percentage of cells expressing the Notch reporter construct (RFP+) which show detectable Notch activity (GFP+). At E17.5, there is a strong bias towards suprabasal Notch activity, while at E15.5 Notch activity is lower overall, and present equally in basal and suprasal cells. (c,d) Spinous differentiation as indicated by K10 (green) at E15.5 (c) and E16.5 (d) in shScramble control (top), LGN knockdown (middle) and Rbpj knockout (bottom) sections of back skin. Note that the formation of the initial spinous layer is not impacted by impairing spindle orientation or Notch. Unlike controls, however, differentiation fails to progress in these mutants. RFP (red) indicates H2B-mRFP1 in top and middle panels, and Cre-mRFP1 in bottom panels. Rbpj loss at this age was confirmed by absence of the target gene Hes1 in suprabasal layers (not shown17). Scale bars: 50 μm.
Supplementary Figure 6 Characterization of shRNA knockdown efficiency in vitro and in vivo.
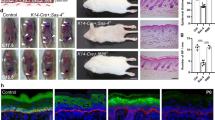
(a) Quantification of mInsc knockdown efficiency in keratinocytes stably-transduced with retroviral mInsc, necessary due to low endogenous levels of mInsc in low calcium conditions. (b) Quantification of Pard3 knockdown in keratinocytes. Red letters indicate those with the highest levels of knockdown (94–97%), subsequently used for in vivo studies. (c) Quantification of mRNA knockdown efficiency in keratinocytes for Gnai2 (left) and Gnai3 (right) shRNA clones. Those with the strongest knockdown (>97%) are shown in red letters, and were subsequently used in vivo. Bars are the mean ± s.d. (d) Back skin sections from E17.5 epidermis showing specific loss of Par3 expression in mosaic Pard3 knockout (top left) or knockdown tissue. (e) Confirmation of knockdown efficiency by immunofluorescence in E17.5 back skin. In addition to its polarized localization in mitotic basal cells, Gαi3 is also localized to cell membranes suprabasally, like Par3 (top panel). This localization is specifically lost in RFP+ regions of mosaic knockdown tissue (bottom panel). (f) Gαii3 is normally enriched apically in mitotic basal cells (left, see also Fig. 5b), but following Gnai3 knockdown, LGN cortical expression is frequently reduced (middle row, weaker hairpin) or delocalized (bottom row, stronger hairpin).
Supplementary information
Supplementary Information
Supplementary Information (PDF 889 kb)
Rights and permissions
About this article
Cite this article
Williams, S., Ratliff, L., Postiglione, M. et al. Par3–mInsc and Gαi3 cooperate to promote oriented epidermal cell divisions through LGN. Nat Cell Biol 16, 758–769 (2014). https://doi.org/10.1038/ncb3001
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3001
This article is cited by
-
Annexin A1 is a polarity cue that directs mitotic spindle orientation during mammalian epithelial morphogenesis
Nature Communications (2023)
-
Anillin governs mitotic rounding during early epidermal development
BMC Biology (2022)
-
ENKD1 promotes epidermal stratification by regulating spindle orientation in basal keratinocytes
Cell Death & Differentiation (2022)
-
ASPP2 maintains the integrity of mechanically stressed pseudostratified epithelia during morphogenesis
Nature Communications (2022)
-
High proliferation and delamination during skin epidermal stratification
Nature Communications (2021)