Abstract
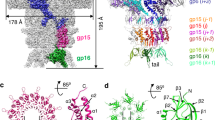
Lambda-like double-stranded (ds) DNA bacteriophage undergo massive conformational changes in their capsid shell during the packaging of their viral genomes. Capsid shells are complex organizations of hundreds of protein subunits that assemble into intricate quaternary complexes that ultimately are able to withstand over 50 atm of pressure during genome packaging1. The extensive integration between subunits in capsids requires the formation of an intermediate complex, termed a procapsid, from which individual subunits can undergo the necessary refolding and structural rearrangements needed to transition to the more stable capsid. Although various mature capsids have been characterized at atomic resolution, no such procapsid structure is available for a dsDNA virus or bacteriophage. Here we present a procapsid X-ray structure at 3.65 Å resolution, termed prohead II, of the lambda-like bacteriophage HK97, the mature capsid structure of which was previously solved to 3.44 Å (ref. 2). A comparison of the two largely different capsid forms has unveiled an unprecedented expansion mechanism that describes the transition. Crystallographic and hydrogen/deuterium exchange data presented here demonstrate that the subunit tertiary structures are significantly different between the two states, with twisting and bending motions occurring in both helical and β-sheet regions. We also identified subunit interactions at each three-fold axis of the capsid that are maintained throughout maturation. The interactions sustain capsid integrity during subunit refolding and provide a fixed hinge from which subunits undergo rotational and translational motions during maturation. Previously published calorimetric data of a closely related bacteriophage, P22, showed that capsid maturation was an exothermic process that resulted in a release of 90 kJ mol-1 of energy3. We propose that the major tertiary changes presented in this study reveal a structural basis for an exothermic maturation process probably present in many dsDNA bacteriophage and possibly viruses such as herpesvirus, which share the HK97 subunit fold4.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Smith, D. E. et al. The bacteriophage straight ϕ29 portal motor can package DNA against a large internal force. Nature 413, 748–752 (2001)
Wikoff, W. R. et al. Topologically linked protein rings in the bacteriophage HK97 capsid. Science 289, 2129–2133 (2000)
Steven, A. C., Heymann, J. B., Cheng, N., Trus, B. L. & Conway, J. F. Virus maturation: dynamics and mechanism of a stabilizing structural transition that leads to infectivity. Curr. Opin. Struct. Biol. 15, 227–236 (2005)
Baker, M. L., Jiang, W., Rixon, F. J. & Chiu, W. Common ancestry of herpesviruses and tailed DNA bacteriophages. J. Virol. 79, 14967–14970 (2005)
Gan, L. et al. Capsid conformational sampling in HK97 maturation visualized by X-ray crystallography and cryo-EM. Structure 14, 1655–1665 (2006)
Conway, J. F. et al. Virus maturation involving large subunit rotations and local refolding. Science 292, 744–748 (2001)
Lata, R. et al. Maturation dynamics of a viral capsid: visualization of transitional intermediate states. Cell 100, 253–263 (2000)
Lee, K. K. et al. Virus capsid expansion driven by the capture of mobile surface loops. Structure 16, 1491–1502 (2008)
Lee, K. K., Tsuruta, H., Hendrix, R. W., Duda, R. L. & Johnson, J. E. Cooperative reorganization of a 420 subunit virus capsid. J. Mol. Biol. 352, 723–735 (2005)
Wikoff, W. R. et al. Time-resolved molecular dynamics of bacteriophage HK97 capsid maturation interpreted by electron cryo-microscopy and X-ray crystallography. J. Struct. Biol. 153, 300–306 (2006)
Helgstrand, C. et al. The refined structure of a protein catenane: the HK97 bacteriophage capsid at 3.44 Å resolution. J. Mol. Biol. 334, 885–899 (2003)
Mandell, J. G., Baerga-Ortiz, A., Akashi, S., Takio, K. & Komives, E. A. Solvent accessibility of the thrombin-thrombomodulin interface. J. Mol. Biol. 306, 575–589 (2001)
Croy, C. H., Bergqvist, S., Huxford, T., Ghosh, G. & Komives, E. A. Biophysical characterization of the free IκBα ankyrin repeat domain in solution. Protein Sci. 13, 1767–1777 (2004)
Lee, K. K. et al. Evidence that a local refolding event triggers maturation of HK97 bacteriophage capsid. J. Mol. Biol. 340, 419–433 (2004)
Conway, J. F. et al. A thermally induced phase transition in a viral capsid transforms the hexamers, leaving the pentamers unchanged. J. Struct. Biol. 158, 224–232 (2007)
Tuma, R., Prevelige, P. E. & Thomas, G. J. Mechanism of capsid maturation in a double-stranded DNA virus. Proc. Natl Acad. Sci. USA 95, 9885–9890 (1998)
Ivanov, D. et al. Domain-swapped dimerization of the HIV-1 capsid C-terminal domain. Proc. Natl Acad. Sci. USA 104, 4353–4358 (2007)
Jiang, W. et al. Coat protein fold and maturation transition of bacteriophage P22 seen at subnanometer resolutions. Nature Struct. Biol. 10, 131–135 (2003)
Fokine, A. et al. Structural and functional similarities between the capsid proteins of bacteriophages T4 and HK97 point to a common ancestry. Proc. Natl Acad. Sci. USA 102, 7163–7168 (2005)
Agirrezabala, X. et al. Quasi-atomic model of bacteriophage t7 procapsid shell: insights into the structure and evolution of a basic fold. Structure 15, 461–472 (2007)
Morais, M. C. et al. Conservation of the capsid structure in tailed dsDNA bacteriophages: the pseudoatomic structure of ϕ29. Mol. Cell 18, 149–159 (2005)
Xie, Z. & Hendrix, R. W. Assembly in vitro of bacteriophage HK97 proheads. J. Mol. Biol. 253, 74–85 (1995)
Duda, R. L. et al. Structural transitions during bacteriophage HK97 head assembly. J. Mol. Biol. 247, 618–635 (1995)
Duda, R. L., Martincic, K. & Hendrix, R. W. Genetic basis of bacteriophage HK97 prohead assembly. J. Mol. Biol. 247, 636–647 (1995)
Conway, J. F., Duda, R. L., Cheng, N., Hendrix, R. W. & Steven, A. C. Proteolytic and conformational control of virus capsid maturation: the bacteriophage HK97 system. J. Mol. Biol. 253, 86–99 (1995)
Duda, R. L. Protein chainmail: catenated protein in viral capsids. Cell 94, 55–60 (1998)
Duda, R. L. Protein chainmail: catenated protein in viral capsids. Cell 94, 55–60 (1998)
Otwiniowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997)
Tong, L. R. & Rossmann, M. G. The locked rotation function. Acta Crystallogr. A 46, 783–792 (1990)
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D. 60, 2126–2132 (2004)
Korostelev, A., Bertram, R. & Chapman, M. S. Simulated-annealing real-space refinement as a tool in model building. Acta Crystallogr. D 58, 761–767 (2002)
Kang, S. & Prevelige, P. E. Domain study of bacteriophage p22 coat protein and characterization of the capsid lattice transformation by hydrogen/deuterium exchange. J. Mol. Biol. 347, 935–948 (2005)
Acknowledgements
We thank V. Reddy for assistance with crystallographic studies and for discussions. We thank R. Huang for providing HK97 capsomer samples and for discussions, and T. Matsui for help with X-ray data collection. We thank B. Firek and C. Moyer for mutagenesis of the HK97 constructs used in the study. We also thank B. Szymczyma for material used in the study. We also thank I. Wilson for discussions. We thank the staffs at beamlines 14-BMC and 23-ID-D of the Advanced Photon Source for assistance in data collection. This work was supported by NIH grants RO1 AI40101 (to J.E.J), RO1 GM47795 (to R.W.H) and NIH Training Grant GM08326.
Author Contributions I.G. was the lead investigator that crystallized the prohead II particles, collected the X-ray data and determined and refined the structure. He also collected and interpreted the hydrogen/deuterium exchange data and prepared the first draft of the paper. L.G. helped with the initial crystallography of the prohead II particles. M.G. helped with the initial collection and interpretation of the hydrogen/deuterium exchange data. K.L. characterized the kinetics and parameters associated with the prohead II to EI transition facilitating the hydrogen/deuterium exchange studies of the EI intermediate. J.A.S. made important contributions during the refinement of the prohead II structure. R.L.D. and R.W.H. developed the HK97 expression system that allowed the studies to be performed, prepared the prohead II mutations that facilitated the production of crystals that diffracted to high resolution, contributed valuable advice for handling the particles and helped in writing the manuscript. E.A.K. supervised the hydrogen/deuterium exchange studies that were all performed in her laboratory. J.E.J. supervised the crystallography aspect of the project, coordinated the overall project and helped in writing the manuscript.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Information
This file contains Supplementary Figure 1 with Legend, Supplementary Table 1, Supplementary Methods, Supplementary References and Supplementary Movie Legends 1-3 (PDF 204 kb)
Supplementary Movie 1
This file shows the transition of subunit F between Prohead II and Head II. (see file s1 for full legend). (MOV 4681 kb)
Supplementary Movie 2
This file shows the isolated view of hexon expansion. (see file s1 for full legend). (MOV 5698 kb)
Supplementary Movie 3
This file shows isolated helix transitions. (see file s1 for full legend). (MOV 4475 kb)
Rights and permissions
About this article
Cite this article
Gertsman, I., Gan, L., Guttman, M. et al. An unexpected twist in viral capsid maturation. Nature 458, 646–650 (2009). https://doi.org/10.1038/nature07686
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature07686
This article is cited by
-
Structural transitions during the scaffolding-driven assembly of a viral capsid
Nature Communications (2019)
-
pH-induced morphological changes of proteinaceous viral shells
Scientific Reports (2019)
-
Structural assembly of the tailed bacteriophage ϕ29
Nature Communications (2019)
-
Automated classification of tailed bacteriophages according to their neck organization
BMC Genomics (2014)
-
Nanoparticle-induced morphological transition of Bombyx mori nucleopolyhedrovirus: a novel method to treat silkworm grasserie disease
Applied Microbiology and Biotechnology (2013)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.