Abstract
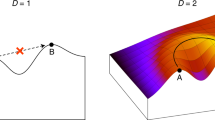
When attempting to understand evolution, we traditionally rely on analysing evolutionary outcomes, despite the fact that unseen intermediates determine its course. A handful of recent studies has begun to explore these intermediate evolutionary forms, which can be reconstructed in the laboratory. With this first view on empirical evolutionary landscapes, we can now finally start asking why particular evolutionary paths are taken.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Darwin, C. On the Origin of Species by Means of Natural Selection Ch VI (Murray, London, 1859)
Pauling, L. & Zuckerkandl, E. Chemical paleogenetics; Molecular “restoration studies” of extinct forms of life. Acta Chem. Scand. A 17, S9–S16 (1963)
Maynard Smith, J. Natural selection and the concept of a protein space. Nature 225, 563–564 (1970)
Malcolm, B. A. et al. Ancestral lysozymes reconstructed, neutrality tested, and thermostability linked to hydrocarbon packing. Nature 345, 86–89 (1990)
Stackhouse, J., Presnell, S. R., McGeehan, G. M., Nambiar, K. P. & Benner, S. A. The ribonuclease from an extinct bovid ruminant. FEBS Lett. 262, 104–106 (1990)
Ugalde, J. A., Chang, B. S. W. & Matz, M. V. Evolution of coral pigments recreated. Science 305, 1433 (2004)
Thornton, J. W. Resurrecting ancient genes: Experimental analysis of extinct molecules. Nature Rev. Genet. 5, 366–375 (2004)
Wright, S. The roles of mutation, inbreeding, crossbreeding and selection in evolution. Proc 6th Int. Cong. Genet. 1, 356–366 (1932).
Gillespie, J. H. The Causes of Molecular Evolution (Oxford Univ. Press, Oxford, 1991)
Kauffman, S. A. The Origins of Order: Self-organization and Selection in Evolution (Oxford Univ. Press, Oxford, 1993)
Gavrilets, S. Fitness Landscapes and the Origin of Species (Princeton Univ. Press, Princeton, 2004)
van Nimwegen, E. & Crutchfield, J. P. Metastable evolutionary dynamics: crossing fitness barriers or escaping via neutral paths? Bull. Math. Biol. 62, 799–848 (2000)
Weinreich, D. M., Watson, R. A. & Chao, L. Sign epistasis and genetic constraint on evolutionary trajectories. Evol. Int. J. Org. Evol. 59, 1165–1174 (2005)
Weinreich, D. M., Delaney, N. F., DePristo, M. A. & Hartl, D. L. Darwinian evolution can follow only very few mutational paths to fitter proteins. Science 312, 111–114 (2006)
Kimura, M. On the probability of fixation of mutant genes in a population. Genetics 47, 713–719 (1962)
Poelwijk, F. J., Kiviet, D. J. & Tans, S. J. Evolutionary potential of a duplicated repressor-operator pair: simulating pathways using mutation data. PLoS Comput. Biol. 2, e58 (2006)
DePristo, M. A., Weinreich, D. M. & Hartl, D. L. Missense meanderings in sequence space: a biophysical view of protein evolution. Nature Rev. Genet. 6, 678–687 (2005)
Bloom, J. D., Labthavikul, S. T., Otey, C. R. & Arnold, F. H. Protein stability promotes evolvability. Proc. Natl Acad. Sci. USA 103, 5869–5874 (2006)
Lunzer, M., Miller, S. P., Felsheim, R. & Dean, A. M. The biochemical architecture of an ancient adaptive landscape. Science 310, 499–501 (2005)
Zhu, G., Golding, G. B. & Dean, A. M. The selective cause of an ancient adaptation. Science 307, 1279–1282 (2005)
Bridgham, J. T., Carroll, S. M. & Thornton, J. W. Evolution of hormone-receptor complexity by molecular exploitation. Science 312, 97–101 (2006)
Lehming, N., Sartorius, J., Kisters-Woike, B., von Wilcken-Bergmann, B. & Müller-Hill, B. Mutant lac repressors with new specificities hint at rules for protein-DNA recognition. EMBO J. 9, 615–621 (1990)
Barkai, N. & Leibler, S. Robustness in simple biochemical networks. Nature 387, 913–917 (1997)
Kirschner, M. & Gerhart, J. Evolvability. Proc. Natl Acad. Sci. USA 95, 8420–8427 (1998)
Kitano, H. Biological robustness. Nature Rev. Genet. 5, 826–837 (2004)
Stelling, J., Sauer, U., Szallasi, Z., Doyle, F. J. & Doyle, J. Robustness of cellular functions. Cell 118, 675–685 (2004)
Thattai, M. & van Oudenaarden, A. Stochastic gene expression in fluctuating environments. Genetics 167, 523–530 (2004)
Kussell, E. & Leibler, S. Phenotypic diversity, population growth, and information in fluctuating environments. Science 309, 2075–2078 (2005)
Arnold, F. H., Wintrode, P. C., Miyazaki, K. & Gershenson, A. How enzymes adapt: lessons from directed evolution. Trends Biochem. Sci. 26, 100–106 (2001)
Elena, S. F. & Lenski, R. E. Evolution experiments with microorganisms: the dynamics and genetic bases of adaptation. Nature Rev. Genet. 4, 457–469 (2003)
Couñago, R., Chen, S. & Shamoo, Y. In vivo molecular evolution reveals biophysical origins of organismal fitness. Mol. Cell 22, 441–449 (2006)
Lenski, R. E. & Travisano, M. Dynamics of adaptation and diversification: a 10,000-generation experiment with bacterial populations. Proc. Natl Acad. Sci. USA 91, 6808–6814 (1994)
Dekel, E. & Alon, U. Optimality and evolutionary tuning of the expression level of a protein. Nature 436, 588–592 (2005)
Hurley, J. H., Dean, A. M., Koshland, D. E. & Stroud, R. M. Catalytic mechanism of NADP(+)-dependent isocitrate dehydrogenase: implications from the structures of magnesium-isocitrate and NADP+ complexes. Biochemistry 30, 8671–8678 (1991)
Hurley, J. H., Chen, R. & Dean, A. M. Determinants of cofactor specificity in isocitrate dehydrogenase: structure of an engineered NADP+ → NAD+ specificity-reversal mutant. Biochemistry 35, 5670–5678 (1996)
Kalodimos, C. G. et al. Plasticity in protein-DNA recognition: lac repressor interacts with its natural operator 01 through alternative conformations of its DNA-binding domain. EMBO J. 21, 2866–2876 (2002)
Kopke Salinas, R. et al. Altered specificity in DNA binding by the lac repressor: a mutant lac headpiece that mimics the gal repressor. ChemBioChem 6, 1628–1637 (2005)
Koradi, R., Billeter, M. & Wüthrich, K. MOLMOL: a program for display and analysis of macromolecular structures. J. Mol. Graph. 14, 51–55 (1996)
Acknowledgements
We thank A. Dean, D. Hartl, J. Thornton and W. Vos for critical reading of the manuscript, and S. Tănase-Nicola for discussions. We thank A. Bonvin and R. Salinas for supplying the data for Fig. 2b. This work is part of the research programme of the Stichting voor Fundamenteel Onderzoek der Materie (FOM), which is financially supported by the Nederlandse Organisatie voor Wetenschappelijke Onderzoek (NWO).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
Reprints and permissions information is available at www.nature.com/reprints. The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Poelwijk, F., Kiviet, D., Weinreich, D. et al. Empirical fitness landscapes reveal accessible evolutionary paths. Nature 445, 383–386 (2007). https://doi.org/10.1038/nature05451
Issue Date:
DOI: https://doi.org/10.1038/nature05451
This article is cited by
-
Early detection of emerging viral variants through analysis of community structure of coordinated substitution networks
Nature Communications (2024)
-
Antifungal alternation can be beneficial for durability but at the cost of generalist resistance
Communications Biology (2023)
-
Quantitative systems-based prediction of antimicrobial resistance evolution
npj Systems Biology and Applications (2023)
-
Statistically learning the functional landscape of microbial communities
Nature Ecology & Evolution (2023)
-
Multiple intermolecular interactions facilitate rapid evolution of essential genes
Nature Ecology & Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.