Abstract
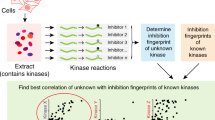
Protein kinases have proved to be largely resistant to the design of highly specific inhibitors, even with the aid of combinatorial chemistry1. The lack of these reagents has complicated efforts to assign specific signalling roles to individual kinases. Here we describe a chemical genetic strategy for sensitizing protein kinases to cell-permeable molecules that do not inhibit wild-type kinases2. From two inhibitor scaffolds, we have identified potent and selective inhibitors for sensitized kinases from five distinct subfamilies. Tyrosine and serine/threonine kinases are equally amenable to this approach. We have analysed a budding yeast strain carrying an inhibitor-sensitive form of the cyclin-dependent kinase Cdc28 (CDK1) in place of the wild-type protein. Specific inhibition of Cdc28 in vivo caused a pre-mitotic cell-cycle arrest that is distinct from the G1 arrest typically observed in temperature-sensitive cdc28 mutants3. The mutation that confers inhibitor-sensitivity is easily identifiable from primary sequence alignments. Thus, this approach can be used to systematically generate conditional alleles of protein kinases, allowing for rapid functional characterization of members of this important gene family.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Gray, N. S. et al. Exploiting chemical libraries, structure, and genomics in the search for kinase inhibitors. Science 281, 533–538 (1998).
Bishop, A. C. et al. Generation of monospecific nanomolar tyrosine kinase inhibitors via a chemical genetic approach. J. Am. Chem. Soc. 121, 627–631 (1999).
Reed, S. I. The selection of S. cerevisiae mutants defective in the start event of cell division. Genetics 95, 561– 577 (1980).
Liu, Y., Shah, K., Yang, F., Witucki, L. & Shokat, K. M. Engineering Src family protein kinases with unnatural nucleotide specificity. Chem. Biol. 5, 91 –101 (1998).
Hanke, J. H. et al. Discovery of a novel, potent, and Src family-selective tyrosine kinase inhibitor. J. Biol. Chem. 271, 695 –701 (1996).
Brown, M. T. & Cooper, J. A. Regulation, substrates and functions of src. Biochim. Biophys. Acta 1287, 121 –149 (1996).
Resh, M. D. Fyn, a Src family tyrosine kinase. Int. J. Biochem. Cell Biol. 30, 1159–1162 ( 1998).
Laneuville, P. Abl tyrosine protein kinase. Semin. Immunol. 7, 255–266 (1995).
Kelly, P. T. Calmodulin-dependent protein kinase II. Multifunctional roles in neuronal differentiation and synaptic plasticity. Mol. Neurobiol. 5, 153–177 (1991).
Morgan, D. O. Principles of CDK regulation. Nature 374, 131–134 (1995).
Lawrie, A. M. et al. Protein kinase inhibition by staurosporine revealed in details of the molecular interaction with CDK2. Nature Struct. Biol. 4, 796–801 (1997).
Wood, J. L., Stoltz, B. M., Dietrich, H. J., Pflum, D. A. & Petsch, D. T. Design and implementation of an efficient synthetic approach to furanosylated indolocarbazoles: total synthesis of (+)- and (-)-K252a. J. Am. Chem. Soc. 119, 9641–9651 (1997).
Liu, Y. et al. Structural basis for selective inhibition of Src family kinases by PP1. Chem. Biol. 6, 671– 678 (1999).
Hunter, T. & Plowman, G. D. The protein kinases of budding yeast: six score and more. Trends Biochem. Sci. 22, 18–22 (1997).
Fujimura, H. A. Yeast homolog of mammalian mitogen-activated protein kinase, Fus3/Dac2 kinase, is required both for cell fusion and for G1 arrest of the cell cycle and morphological changes by the cdc37 mutation. J. Cell. Sci. 107, 2617–2622 (1994).
Doi, S. & Yoshimura, M. Temperature-sensitive loss of sexual agglutinability in Saccharomyces cerevisiae. Arch. Microbiol. 114, 287–288 (1977).
Mendenhall, M. D. & Hodge, A. E. Regulation of Cdc28 cyclin-dependent protein kinase activity during the cell cycle of the yeast Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 62, 1191–1243 ( 1998).
King, R. W., Jackson, P. K. & Kirschner, M. W. Mitosis in transition. Cell 79, 563–571 (1994).
Surana, U. et al. The role of CDC28 and cyclins during mitosis in the budding yeast S. cerevisiae. Cell 65, 145 –161 (1991).
Bourne, Y. et al. Crystal structure and mutational analysis of the human CDK2 kinase complex with cell cycle-regulatory protein CksHs1. Cell 84, 863–874 ( 1996).
Wodicka, L., Dong, H., Mittmann, M., Ho, M. H. & Lockhart, D. J. Genome-wide expression monitoring in Saccharomyces cerevisiae. Nature Biotechnol. 15, 1359 –1367 (1997).
Espinoza, F. H., Ogas, J., Herskowitz, I. & Morgan, D. O. Cell cycle control by a complex of the cyclin HCS26 (PCL1) and the kinase PHO85. Science 266, 1388–1391 ( 1994).
Spellman, P. T. et al. Comprehensive identification of cell cycle-regulated genes of the yeast Saccharomyces cerevisiae by microarray hybridization. Mol. Biol. Cell 9, 3273– 3297 (1998).
Fitch, I. et al. Characterization of four B-type cyclin genes of the budding yeast Saccharomyces cerevisiae. Mol. Biol. Cell 3, 805–818 (1992).
Stern, B. & Nurse, P. A quantitative model for the cdc2 control of S phase and mitosis in fission yeast. Trends Genet. 12, 345–350 ( 1996).
Farrell, A. & Morgan, D. O. Cdc37 promotes the stability of protein kinases Cdc28 and Cak1. Mol. Cell. Biol. 20 , 749–754 (2000).
Sherman, F., Fink, G. & Lawrence, C. Methods in Yeast Genetics (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, 1974).
Sikorski, R. S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Schindler, T. et al. Crystal structure of Hck in complex with a Src family-selective tyrosine kinase inhibitor. Mol. Cell 3, 639–648 (1999).
Biggins, S. et al. The conserved protein kinase Ipl1 regulates microtubule binding to kinetochores in budding yeast. Genes Dev. 13, 532–544 (1999).
Acknowledgements
We thank B. Schulman and E. Harlow for early characterization of mutant cyclin-dependent kinases; C. Co, V. Nguyen and J. Li for help with flow cytometry; D. Toczyski for help with the Coulter counter; A. Su for help with transcript array analysis; S. Jaspersen, C. Carroll, C. Takizawa, E. Weiss, S. Biggins, A. Szidon, A. Rudner, D. Kahne and A. Murray for helpful advice and reagents; E. O'Shea and R. Deshaies for strains and reagents. This work was supported by funding from the National Institute of Allergy and Immunology (K.M.S.), GlaxoWellcome (K.M.S. and J.L.W.), National Institute of General Medical Sciences (D.O.M.), National Institutes of Health (J.L.W.), Bristol-Myers Squibb (J.L.W.), Yamanouchi (J.L.W.) and a predoctoral fellowship from the National Science Foundation (J.A.U.). K.M.S. is a Pew, Searle, and a Sloan Foundation Scholar.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bishop, A., Ubersax, J., Petsch, D. et al. A chemical switch for inhibitor-sensitive alleles of any protein kinase . Nature 407, 395–401 (2000). https://doi.org/10.1038/35030148
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/35030148
This article is cited by
-
Qualitative rather than quantitative phosphoregulation shapes the end of meiosis I in budding yeast
The EMBO Journal (2024)
-
The NELF pausing checkpoint mediates the functional divergence of Cdk9
Nature Communications (2023)
-
LTP induction by structural rather than enzymatic functions of CaMKII
Nature (2023)
-
Chemogenetics for cell-type-specific modulation of signalling and neuronal activity
Nature Reviews Methods Primers (2023)
-
Analog-sensitive Cdk1 as a tool to study mitotic exit: protein phosphatase 1 is required downstream from Cdk1 inactivation in budding yeast
Chromosome Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.