Abstract
Type II diabetes mellitus (T2DM) is often accompanied by non-alcoholic steatohepatitis (NASH) and associated with hypercholesterolemia, that is, increased levels of plasma low-density lipoprotein (LDL) and oxidized LDL (ox-LDL). Approximately one-third of NASH develops hepatic fibrosis. The role of hypercholesterolemia in T2DM and NASH-associated hepatic fibrogenesis remains obscure. We previously reported that the phytochemical curcumin inhibited the activation of hepatic stellate cells (HSCs), the major effector cells during hepatic fibrogenesis, and protected the liver from fibrogenesis in vitro and in vivo. The aims of this study are to evaluate the role of ox-LDL in activation of HSCs, to assess curcumin effects on eliminating the role of ox-LDL, and to further explore the underlying mechanisms. In this report, we observe that ox-LDL alters the expression of genes closely relevant to HSC activation, which is eliminated by curcumin. Curcumin suppresses gene expression of lectin-like oxidized LDL receptor-1 (LOX-1), leading to the blockade of the transport of extracellular ox-LDL into cells. This suppressive effect of curcumin results from the interruption of Wnt signaling and the activation of peroxisome proliferator-activated receptor-gamma (PPARγ). In conclusion, these results support our initial hypothesis and demonstrate that ox-LDL stimulates HSC activation, which is eliminated by curcumin by suppressing lox-1 expression by interrupting Wnt signaling and stimulating PPARγ activity. These results provide novel insights into the role of ox-LDL in T2DM and NASH-associated hepatic fibrogenesis and mechanisms by which curcumin suppresses ox-LDL-induced HSC activation, as well as the implication of curcumin in the treatment of T2DM and NASH-associated hepatic fibrosis.
Similar content being viewed by others
Main
Almost 6% of the world's adult population now lives with diabetes mellitus. Type II diabetes mellitus (T2DM), that is non-insulin-dependent diabetes mellitus, represents over 80% of all diabetics and is dramatically increasing in incidence as a result of changes in human behavior and increased body mass index.1 T2DM is often associated with non-alcoholic steatohepatitis (NASH).2, 3 NASH is characterized by fat accumulation and inflammation in the liver. Approximately one-third of NASH patients develop hepatic fibrosis and even cirrhosis.4 Both T2DM and NASH are most commonly present in obese patients with hypercholesterolemia, that is increased levels of plasma low-density lipoprotein (LDL) and oxidized LDL (ox-LDL).4 However, the role of hypercholesterolemia in hepatic fibrogenesis remains obscure.
Low-density lipoprotein becomes the highly reactive form of ox-LDL after lipid peroxidative modification. High concentrations of circulating ox-LDL are associated with high incidences of metabolic syndromes, such as T2DM and coronary heart disease.5 Cellular uptake of ox-LDL is mediated by binding to its scavenger receptors, such as lectin-like oxidized LDL receptor-1 (LOX-1), leading to the elevation of intracellular levels of ox-LDL and reactive oxygen species (ROS), as well as to the activation of intracellular signaling.6, 7 LOX-1 was originally identified as a major scavenger receptor for ox-LDL in endothelial cells, and was subsequently detected in many other cell types.7, 8 LOX-1 gene expression is induced by ox-LDL9 and is upregulated in obese patients with hyperlipidemia.6
Hepatic fibrosis is a progressive disorder characterized by the accumulation of extracellular matrix (ECM) components.10, 11 Hepatic stellate cells (HSCs) are the major effector cells during hepatic fibrogenesis and are the primary source of ECM production in the liver.10, 11 During liver injury, quiescent HSCs undergo dramatic phenotypic changes from vitamin A, fat-storing cells to proliferative myofibroblast-like cells with acquisition of fibrogenic properties.10, 11 This process is coupled with the activation of signaling pathways for pro-fibrogenic transforming growth factor-beta (TGF-β),12 pro-mitogenic platelet-derived growth factor-beta (PDGF-β),13 and Wnt signaling,14, 15, 16 as well as the depletion of peroxisome proliferator activated receptor-gamma (PPARγ).17, 18, 19 It is important to note that culturing quiescent HSCs on plastic plates causes spontaneous activation, mimicking the process seen in vivo, which provides a good model for elucidating underlying mechanisms of HSC activation and for studying therapeutic intervention of the process.10, 11
Curcumin (diferuloylmethane), the yellow pigment in curry from turmeric, is one of the best-studied natural compounds. Although the underlying mechanisms remain elusive, curcumin has shown diverse and versatile beneficial effects, including anti-inflammatory, anti-oxidative stress, anti-viral, anti-hypercholesterolemic, anti-infective, and anti-carcinogenic effects.20 Curcumin has recently received attention as a promising dietary supplement for liver protection.21 We recently reported that curcumin inhibited HSC activation by inhibiting cell proliferation, inducing apoptosis, and attenuating oxidative stress in vitro and in vivo.22, 23, 24, 25, 26, 27 In addition, we demonstrated that curcumin dramatically induced the gene expression of endogenous PPARγ and stimulated its activity in activated HSCs in vitro and in vivo, which is required for curcumin to inhibit HSC activation.22, 23, 24 Furthermore, curcumin suppressed the gene expression of LDL receptor in activated HSCs in vitro by activating PPARγ and regulating gene expression of the transcription factor sterol regulatory element binding proteins (SREBPs), leading to the reduction in the level of intracellular cholesterol and to the attenuation of the stimulatory effects of LDL on HSC activation (Kang and Chen, manuscript accepted by British Journal of Pharmacology, in press).
The aims of this study were to evaluate the role of ox-LDL in activating HSCs, to assess the effects of curcumin on eliminating the stimulatory role, and to further explore the underlying mechanisms. Results in the current report supported our initial hypothesis that ox-LDL might stimulate HSC activation, which could be eliminated by curcumin by suppressing gene expression of LOX-1.
MATERIALS AND METHODS
Isolation and Culture of Rat HSCs and Chemicals
Primary HSCs were isolated from male Sprague–Dawley rats (200–250 g) as previously described.23 Passaged HSCs were grown in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% (v/v) fetal bovine serum (FBS). Cultured HSCs were used at Passage #4–9. Human ox-LDL was purchased from Intracel Company (Frederick, MD, USA) and used before expiration. The presence of endotoxin in ox-LDL was <0.1 U/ml, as measured by Limulus Amebocyte lysate assay kit (Whittaker M.A. Bioproducts, Walkersville, MD, USA). Wnt3a was purchased from R&D Systems (Minneapolis, MN, USA). PPARγ antagonist PD68235 was kindly provided by Pfizer (Ann Arbor, MI, USA). Curcumin (purity >94%) and κ-carrageenan were purchased from Sigma (St Louis, MO, USA). 15-deoxy-Δ12, 14-prostaglandin J2 (PGJ2), a natural PPARγ agonist, was a product of Cayman Chemical Company (Ann Arbor, MI, USA).
Western Blotting Analyses
Protein samples were prepared from whole cell extracts, separated by 10% SDS-PAGE, and electro-blotted onto PVDF membrane. Target proteins were visualized by using the ECL method (Amersham, Piscataway, NJ, USA). Mouse anti-LOX-1 monoclonal antibody was kindly provided by Dr Tatsuya Sawamura (National Cardiovascular Center Research Institute, Japan). Other primary and secondary antibodies were presented in Table 1. Densities of bands in Western blotting analyses were normalized with the internal invariable control. Levels of target protein bands were densitometrically determined by using Quantity One 4.4.1 (Bio-Rad, Hercules, CA, USA). Variations in the density were expressed as fold changes compared with the control in the blot.
Immunostaining
Serum-starved HSCs were treated with or without human ox-LDL (10 μg/ml) in the presence or absence of curcumin (20 μM) for 24 h before the fixation with 4% paraformaldehyde for 15 min at room temperature (RT). After blocking with 5% BSA for 1 h, fixed cells were incubated with anti-human ox-LDL antibody (1:100) (cat# AB3230, Chemicon Company, Billerica, MA, USA) overnight at 4°C. After three washes with phosphate-buffered saline (PBS), sections on slides were incubated with goat anti-rabbit Alexa Fluor 488-conjugated secondary antibodies (1:500) (Invitrogen, Carlsbad, CA, USA) at RT for 1 h. Sections incubated with secondary antibodies alone were used as negative controls (data not shown). DAPI (100 ng/ml) (Molecular Probes Company, Eugene, OR, USA) was added in the mounting solution for nuclear staining. Sections were viewed under Leica DM 400 B microscope (North Central Instruments, Maryland Heights, MO, USA).
Preparation of Nuclear Protein Extracts
Nuclear extracts were prepared as previously described.28 In brief, after washing twice with PBS, cells were evenly re-suspended in Buffer A (10-mM HEPES–KOH pH7.9, 1.5-mM MgCl2, 10-mM KCl, 1-mM DTT, and 1-mM PMSF). After incubation on ice for 10 min, cells were vortexed for 30 s and centrifuged at 8000 g for 10 s. Pellets were re-suspended in Buffer C (20-mM HEPES–KOH pH7.9, 25% glycerol, 420-mM NaCl, 1.5-mM MgCl2, 0.2-mM EDTA, 1-mM DTT, and 1-mM PMSF) and incubated on ice for 15 min before vortexing. The lysates were centrifuged at 8000 g at 4°C for 2 min, and the resulting supernatants were taken as nuclear protein extracts, and stored at −80°C until use.
Plasmids and Transient Transfection Assays
The lox-1 promoter luciferase reporter plasmid (p-2336/+36-Luc), containing a 2336 bp fragment of the 5′-flanking promoter region of the LOX-1 gene, and some of the reporter plasmids with various lengths of the lox-1 promoter were kindly provided by Dr Jawahar L. Mehta (Department of Internal Medicine, University of Arkansas for Medical Sciences, Little Rock, AR, USA).29 Additional lox-1 promoter luciferase reporter constructs with shorter promoter regions were created by PCR and enzyme ligation using p-2336/+36-Luc as a template (see the following for details). The PDGF-βR promoter luciferase reporter plasmid pPDGF-βR-Luc (pβ12) was a gift from Dr Keiko Funa (Ludwig Institute for Cancer Research, Uppsala, Sweden).30 The type I TGF-β receptor promoter luciferase reporter plasmid pTβ-RI-Luc (pES1.0) was kindly provided by Dr Michael Centrella (Yale University, New Haven, CT, USA).31 The PPARγ cDNA expression plasmid pPPARγ, containing a full size of PPARγ cDNA, was a gift from Dr Reed Graves (Department of Medicine, University of Chicago, Chicago, IL, USA). The dominant negative PPARγ expression construct pdn-PPARγ was a gift from Dr Krishna V. Chatterjee.32 The Wnt signaling luciferase reporter plasmids TOPflash and its mutant FOPflash were kindly provided by Dr Randall T. Moon (Department of Pharmacology, School of Medicine, University of Washington, Seattle, WA, USA).33 TOPflash contained eight copies of TCF–LEF binding sites. FOPflash was a counterpart control for TOPflash with site-directed mutations in the TCF–LEF binding sites.33
Transient transfection of semi-confluent HSCs in 6-well plates was carried out using the LipofectAMINE reagent (Invitrogen Carlsbad, CA, USA), as previously described.23 Each treatment had triplicates in every experiment. Each experiment was repeated, at least, three times. Luciferase activity assays were conducted as we previously described.23 Transfection efficiency was determined by co-transfection of a β-galactosidase reporter, pSV-β-gal (Promega, Madison, WI, USA). β-galactosidase assays were carried out using an assay kit from Promega, according to the manufacturer's instruction. Luciferase activities were expressed as a relative unit after normalization with β-galactosidase activities. Results were combined from, at least, three independent experiments.
Generation of a LOX-1 cDNA Expression Plasmid
Total RNA was extracted from HSCs using Trizol reagent and reversely transcribed into cDNA, which was used as a template for PCR amplification of LOX-1 cDNA. The PCR primers were designed according to GenBank accession number: NM_138648, and used the following primers: (F) 5′-CCCAAGCTTATGACTTTTGATGACAAGATGAAGC-3′; (R) 5′-GGGGTACCCCCTAAATTTGCAAATGATTTGTCTTC-3′. The primers were tailed with HindIII site (the forward primer) and KpnI site (the reverse primer). The PCR product was subcloned into pCDNA3.1/Zeo+ expression vector (Invitrogen) at HindIII/KpnI sites. The subcloned LOX-1 cDNA was verified by DNA sequencing.
Promoter Deletion and Site-Directed Mutagenesis
The lox-1 promoter luciferase reporter plasmid p-2336/+36-Luc, also termed pLOX-1-Luc, was used as a PCR template to generate constructs with various lengths of the gene promoter. Primers in Table 2 were used for generating the plasmids by PCR. The primers were tailed with KpnI site (forward primers) or BglII site (reverse primer), and the PCR products were subcloned into pGL3-Basic (Promega) at KpnI/BglII sites. The site-directed mutagenesis was carried out using the GeneTailor Site-Directed Mutagenesis kit (Invitrogen), according to the manufacture's instruction. The sequence of 5′-GGCACATTTTTTACAAATGTAGTGTGACTTACTCTCTTTGAATTTCAGTTTC-3′ (−310 to −258) containing the putative TCF–LEF-1 binding site in the plasmid p-2336/+36-Luc, was mutated to 5′-GGCACATTTTTTACAAATGTAGTGTGACTTACTCTCATAGAATTTCAGTTTC-3′ in p-2336/+36-Mut-Luc. The primers used for mutagenesis were in Table 3. The mutations were verified by DNA sequencing.
Real-Time PCR
Total RNA was extracted from cells using Trizol reagent (Invitrogen) following the manufacturer's instruction. Real-time PCR was carried out using SYBR green, as previously described.34 Total RNA was treated with DNase I before the synthesis of the first strand of cDNA. First strand cDNA was synthesized using total RNA as templates and oligodT as primers. Samples were run on a Bio-Rad MyiQ real-time PCR machine (Bio-Rad). mRNA levels were expressed as fold changes after normalization with endogenous glyceraldehyde-3-phosphate dehydrogenase (GAPDH) as suggested by Schmittgen et al.35 The primers for PCR were presented in Table 3.
Electrophoretic Mobility Shift Assays (EMSAs)
Synthesized single-stranded oligonucleotide probes were biotin-labeled using the Biotin 3′ End DNA Labeling kit (Pierce, Rockford, IL, USA), and annealed to form oligonucleotide duplex after labeling. EMSAs were conducted using LightShift Chemiluminescent EMSA kit (Pierce), following the protocol provided by the manufacturer. Briefly, the annealed, biotin-labeled double-stranded oligonucleotide probes were incubated with 3 μg of nuclear extracts at room temperature for 30 min. The mixtures were subjected to 6% non-denatured PGE, and electroblotted onto a nylon membrane. Proteins bound to the biotin end-labeled oligonucleotide probes were detected using the streptavidin–horseradish peroxidase conjugate and the chemiluminescent substrate. The following probes were used. P(TCFwt) contained a putative TCF-1 binding site: 5′-GTGTGACTTACTCTCTTTGAATTTCAGTTTCTGTC-3′ (−289 to −259). P(TCFmut) contained a mutated TCF-1 binding site: 5′-GTGTGACTTACTCTC A T A GAATTTCAGTTTCTGTC-3′.
Statistical Analyses
Differences between means were evaluated by an unpaired two-sided Student's t-test (P<0.05 considered as significant). Where appropriate, comparisons of multiple treatment conditions with controls were analyzed by ANOVA with the Dunnett's test for post hoc analysis.
RESULTS
Ox-LDL Induces Expression of Genes Relevant to the Activation of HSCs, Which is Abrogated by Curcumin in Cultured HSCs
It was previously demonstrated that ox-LDL prompted the biosynthesis of ECM components in HSCs, implying ox-LDL being a pro-fibrogenic stimulator.36 We further examined the effect of ox-LDL on the expression of genes closely relevant to HSC activation. Semi-confluent cultured HSCs were serum-starved for 24 h before the stimulation with exogenous ox-LDL at various concentrations (0–15 μg/ml) in serum-depleted media for additional 24 h. Serum starvation rendered cells more sensitive to stimuli, including exogenous ox-LDL. Subsequent culture in serum-depleted media excluded the interference from other factors in FBS.37, 38 Total RNA and whole cell protein extracts were prepared from the cells for real-time PCR and Western blotting analyses, respectively. As shown in Figure 1a and b, ox-LDL increased gene expression of α(I) collagen and alpha-smooth muscle actin (α-SMA), the markers of activated HSCs, in a dose-dependent manner. In addition, ox-LDL dose-dependently stimulated the expression of pro-fibrogenic genes, including connective tissue growth factor (CTGF) and type I and II TGF-β receptors (Tβ-RI & Tβ-RII), as well as pro-mitogenic genes, including receptors for PDGF-β (PDGF-βR) and EGF (EGFR), as well as cyclin D1, a key regulator in cell cycle progression. These data collectively indicated that exogenous ox-LDL markedly induced expression of genes closely relevant to HSC activation, suggesting the role of ox-LDL in the stimulation of HSC activation in vitro.
Oxidized low-density lipoprotein (ox-LDL) stimulates expression of genes relevant to hepatic stellate cell (HSC) activation, which is dose-dependently eliminated by curcumin. Serum-starved HSCs were stimulated with ox-LDL at indicated concentrations in serum-depleted media for 24 h in the absence (a and b) or presence (c and d) of curcumin at 0–30 μM. Total RNA and whole cell protein extracts were respectively prepared from the cells for real-time PCR (a and c) and Western blotting analyses (b and d). *P<0.05 vs cells without treatment (n=3) (the first corresponding column on the left); ‡P<0.05 vs cells treated with ox-LDL only (the second corresponding column). β-actin was used in Western blotting analyses as an internal control for equal protein loading. Representatives were from three independent experiments.
To evaluate the effect of curcumin on the elimination of the stimulatory role of ox-LDL in the activation of HSCs, serum-starved HSCs were stimulated with ox-LDL (10 μg/ml) in the presence of curcumin at various concentrations (0–30 μM) in serum-depleted media for 24 h. Total RNA and whole cell extracts were prepared to analyze expression of genes related to HSC activation by real-time PCR and Western blotting analyses. As indicated in Figure 1c and d, curcumin dose-dependently inhibited expression of the pro-fibrogenic and pro-mitogenic genes induced by ox-LDL in cultured HSCs. Taken together, these results demonstrated that ox-LDL induced expression of genes closely relevant to the activation of HSCs, which was dose-dependently abrogated by curcumin.
LOX-1 has a Mediating Role in ox-LDL-Induced HSC Activation and in the Curcumin Elimination of the Stimulatory Effect of ox-LDL
Oxidized LDL receptor-1 has been identified as a specific receptor for ox-LDL, responsible for transporting extracellular ox-LDL into cells.6, 7 LOX-1, as a scavenger receptor, facilitates the accumulation of ox-LDL and stimulates the transformation of smooth muscle cells and monocyte/macrophages into foam cells.39 LOX-1 gene expression is induced by ox-LDL.9 We assumed that LOX-1 mediated the stimulatory effects of ox-LDL on HSC activation and fibrogenesis. To elucidate the link role of LOX-1 in ox-LDL-induced HSC activation, serum-starved HSCs were pre-treated with or without κ-carrageenan (250 μg/ml), a LOX-1 antagonist,40, 41 for 1 h before the stimulation with ox-LDL (10 μg/ml) for additional 24 h. Whole cell extracts were prepared for Western blotting analyses of genes relevant to HSC activation. As shown in Figure 2a, compared with the untreated control (the corresponding first wells), ox-LDL, as expected, significantly increased the abundance of proteins closely relevant to HSC activation, including pro-mitogenic PDGF-βR and EGFR and pro-fibrogenic CTGF, Tβ-RI, and Tβ-RII (the corresponding second wells). It was of interest to observe that the pretreatment with the LOX-1 antagonist κ-carrageenan dramatically eliminated the stimulatory effects of ox-LDL on the abundance of proteins (the corresponding fourth wells). These results suggested a direct link of LOX-1 between ox-LDL and HSC activation, as well as fibrogenesis.
Oxidized low-density lipoprotein receptor-1 (LOX-1) has a mediating role in oxidized low-density lipoprotein (ox-LDL)-induced hepatic stellate cell (HSC) activation and in the curcumin elimination of the stimulatory effect of ox-LDL. (a) Serum-starved HSCs were pre-treated with or without the LOX-1 antagonist κ-carrageenan at 250 μg/ml for 1 h before the stimulation with or without ox-LDL (10 μg/ml) for additional 24 h. Whole cell extracts were prepared for Western blotting analyses of genes relevant to HSC activation. β-actin was used in Western blotting analyses as an internal control for equal protein loading. Representatives were from three independent experiments. Italic numbers beneath blots were fold changes in the densities of the bands compared with the control without treatment in the blot (n=3), after normalization with the internal invariable control β-actin. Owing to limited space, s.d. were not presented. (b) HSCs were co-transfected with the LOX-1 cDNA expression plasmid pLOX-1 at indicated doses plus pPDGF-βR-Luc, or pTβ-RI-Luc. A total of 4.5 μg of plasmid DNA per well was used for co-transfection of HSCs in 6-well culture plates. It included 2 μg of pPDGF-βR-Luc or pTβ-RI-Luc, 0.5 μg of pSV-β-gal, and 2.0 μg of pLOX-1 at indicated doses plus the empty vector pcDNA. The latter was used to ensure an equal amount of total DNA in transfection assays. After recovery, cells were stimulated with or without ox-LDL (10 μg/ml) in the presence or absence of curcumin (20 μM) in serum-depleted media for 24 h. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.; n≥6). *P<0.05 vs cells without treatment (the corresponding first column); **P<0.05 vs cells treated only with ox-LDL (the corresponding second column); ‡P<0.05 vs cells treated with ox-LDL plus curcumin (the corresponding third column). The floating schema denoted the luciferase reporter construct pPDGF-βR-Luc or pTβ-RI-Luc in use and forced expression of LOX-1 cDNA in the system.
To determine the role of LOX-1 in the curcumin elimination of the stimulatory effects of ox-LDL, HSCs were co-transfected with LOX-1 cDNA expression plasmid pLOX-1 at various doses plus the PDGF-βR gene promoter luciferase reporter plasmid pPDGF-βR-Luc, or the Tβ-RI gene promoter luciferase reporter plasmid pTβ-RI-Luc. A total of 4.5 μg of plasmid DNA per well was used for co-transfection of HSCs in 6-well culture plates. It included 2 μg of pPDGF-βR-Luc or pTβ-RI-Luc, 0.5 μg of pSV-β-gal, and 2.0 μg of pLOX-1 at indicated doses plus the empty vector pcDNA. The latter was used to ensure an equal amount of total DNA in transfection assays. After recovery, cells were stimulated with or without ox-LDL (10 μg/ml) in the presence or absence of curcumin (20 μM) in serum-depleted media for 24 h. As shown in Figure 2b by luciferase activity assays, compared with the untreated control (the corresponding first columns), ox-LDL, as expected, significantly increased luciferase activities in the cells (the corresponding second columns). Curcumin significantly diminished the ox-LDL-elevated luciferase activities (the corresponding third columns). Further experiments revealed that forced expression of LOX-1 cDNA dose-dependently eliminated the inhibitory role of curcumin and elevated luciferase activities (the last three corresponding columns). These results indicated that forced expression of LOX-1 cDNA eliminated the inhibitory effect of curcumin on the ox-LDL-induced promoter activities of pro-mitogenic PDGF-βR and pro-fibrogenic Tβ-RI genes. Taken together, our results demonstrated the mediating role of LOX-1 in ox-LDL-induced HSC activation and in the curcumin elimination of stimulatory effect of ox-LDL.
Curcumin Suppresses lox-1 Expression in Cultured HSCs
Based on the above observations, we hypothesized that curcumin might eliminate the stimulatory effects of ox-LDL on the activation of HSCs by suppressing gene expression of LOX-1, leading to a reduction in the bioavailability of the receptor to transport extracellular ox-LDL into HSCs. To test the hypothesis, cultured HSCs were transiently transfected with the LOX-1 promoter luciferase reporter plasmid pLOX-1-Luc. This plasmid contained a 5′-flanking fragment (2336 bp) of the LOX-1 gene promoter, subcloned in a luciferase reporter plasmid.29 After overnight recovery, cells were treated with curcumin at various concentrations (0–30 μM) for 24 h. As shown in Figure 3a by luciferase activity assays, curcumin caused a dose-dependent reduction in luciferase activities, suggesting that curcumin reduced the promoter activity of LOX-1 gene in HSCs. To verify the observation, semi-confluent passaged HSCs were treated with curcumin at various concentrations (0–30 μM) for 24 h. Total RNA and whole cell extracts were prepared for real-time PCR and Western blotting analyses. As shown in Figure 3b and c, curcumin dose-dependently reduced the steady state mRNA levels and the protein abundance of LOX-1 in the cells.
Curcumin suppresses lox-1 expression in activated hepatic stellate cells (HSCs) in vitro. (a) Luciferase activity assays of cells transiently transfected with the lox-1 promoter luciferase reporter plasmid pLOX-1-Luc, and treated with curcumin after transfection. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.; n≥6). *P<0.05 vs cells without treatment (the first column on the left). The floating schema denoted the luciferase reporter construct pLOX-1-Luc in use and the application of curcumin to the system. (b) Real-time PCR analyses of the steady-state levels of LOX-1 mRNA in cells treated with curcumin. mRNA fold changes were calculated as stated in MATERIALS and METHODS. Values were expressed as means±s.d. (n≥3). *P<0.05 vs the untreated control (the first column on the left); (c) Western blotting analyses of LOX-1 in cells treated with curcumin at indicated concentrations. β-actin was used as an internal control for equal protein loading. Representative was from three independent experiments. Italic numbers beneath the blot were fold changes in the densities of the bands compared with the control without treatment in the blot (n=3), after normalization with the internal invariable control β-actin. (d) Immuno-staining of cultured HSCs treated with or without oxidized low-density lipoprotein (ox-LDL) (10 μg/ml) plus or minus curcumin (10 μM) for evaluating the impact of curcumin on the abundance of intracellular ox-LDL. DAPI in mounting solution was used for staining nuclei. Representative views were presented.
To evaluate the effect of curcumin on the level of intracellular ox-LDL, serum-starved HSCs were treated with or without human ox-LDL (10 μg/ml) in the presence or absence of curcumin (20 μM) for 24 h. Results from immuno-cytochemistry in Figure 3d indicated that ox-LDL treatment significantly increased the abundance of intracellular ox-LDL, which was apparently ameliorated by curcumin. Taken together, these results collectively demonstrated that curcumin suppressed the gene expression of LOX-1 in activated HSCs in vitro, which likely contributed to the reduction in the abundance of intracellular ox-LDL.
The Activation of PPARγ Likely has a Critical Role in the Curcumin-Caused Inhibition of lox-1 Expression in Activated HSCs
We have previously reported that curcumin induces the expression of endogenous PPARγ gene and stimulates its activity in activated HSCs, which is a prerequisite for curcumin to inhibit HSC activation in vitro.23, 24 To elucidate the mechanisms by which curcumin suppressed gene expression of LOX-1 in HSCs, we postulated that PPARγ mediated the inhibitory effect of curcumin on lox-1 expression. To test this postulation, HSCs were transiently transfected with the LOX-1 promoter luciferase reporter pLOX-1-Luc. After recovery, cells were pretreated with or without the PPARγ antagonist PD68235 (20 μM) for 30 min before the addition of curcumin at 20 μM for additional 24 h. As shown in Figure 4a by luciferase activity assays, compared with the untreated control cells (the first column on the left), cells treated with curcumin showed a significant reduction in luciferase activity (the second column), confirming the inhibitory effect of curcumin on the lox-1 promoter activity in HSCs. In great contrast, the pretreatment of cells with the PPARγ antagonist PD68235 abrogated this inhibitory effect of curcumin (the fourth column), suggesting the necessity of the activation of PPARγ in the curcumin-caused inhibition of the lox-1 promoter activity. To verify the observation, HSCs were pretreated with or without PD68235 (20 μM) for 30 min before the addition of curcumin (20 μM) for additional 24 h. Whole cell extracts were prepared for Western blotting analyses. As demonstrated in Figure 4b, compared with the untreated control (the first well), curcumin, as expected, significantly reduced the abundance of LOX-1 in the cells (the second well). The inhibition of the PPARγ activity by the pretreatment with PD68235 apparently eliminated the inhibitory effect of curcumin on the abundance of LOX-1 (the fourth well). These results collectively suggested that the activation of PPARγ might has a critical role in the curcumin-caused inhibition of lox-1 expression in activated HSCs in vitro.
The activation of peroxisome proliferator-activated receptor-gamma (PPARγ) is required for curcumin to inhibit lox-1 expression in activated hepatic stellate cells (HSCs) in vitro. Semi-confluent HSCs were pretreated with or without the PPARγ antagonist PD68235 (20 μM) for 30 min before the addition of curcumin at 20 μM for additional 24 h. (a) Luciferase activity assays of HSCs transfected with the plasmid pLOX-1-Luc, followed by the above treatment. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.; n≥6). *P<0.05 vs cells with no treatment (the first column on the left). **P<0.05 vs cells treated with curcumin only (the second column). (b) Western blotting analyses of oxidized low-density lipoprotein receptor-1 (LOX-1) in cultured HSCs with the above treatment. β-actin was used as an internal control for equal protein loading. Representative was from three independent experiments. Italic numbers beneath the blot were fold changes in the densities of the bands compared to the control without treatment in the blot (n=3), after normalization with β-actin. Because of the limited space, standard deviations were not presented.
The Activation of PPARγ Results in the Suppression of lox-1 Expression in Activated HSCs In Vitro
To verify the role of PPARγ activation in the inhibition of lox-1 expression, semi-confluent HSCs were co-transfected with pLOX-1-Luc and the plasmid pPPARγ, or the plasmid pdn-PPARγ, at indicated doses. The cDNA expression plasmid pPPARγ contained a full size of wild-type PPARγ cDNA.24 pdn-PPARγ contained a full length of cDNA encoding dominant negative PPARγ (dn-PPARγ).32 A total of 4.5 μg of plasmid DNA per well was used for co-transfection of HSCs in 6-well culture plates. It included 2 μg of pLOX-1-Luc, 0.5 μg of pSV-β-gal, and 2.0 μg of pPPARγ, or pdn-PPARγ, at indicated doses plus the empty vector pcDNA. The latter was used to ensure an equal amount of total DNA in transfection assays. After recovery, cells were treated with or without curcumin (20 μM) in DMEM with FBS (10%) for 24 h. Previous experiments have suggested that 10% of FBS in the medium contains enough agonists to activate PPARγ in HSCs.19, 23, 24 Luciferase activity assays demonstrated that forced expression of wild-type PPARγ cDNA dose-dependently reduced luciferase activities (Figure 5a). In contrast, forced expression of dn-PPARγ dose-dependently abrogated the inhibitory effect of curcumin on the lox-1 promoter activity (Figure 5b). These results collectively suggested the inhibitory role of PPARγ activation and its requirement in the curcumin-caused inhibition of the lox-1 promoter activity.
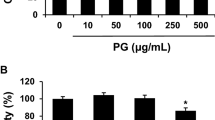
The activation of peroxisome proliferator-activated receptor-gamma (PPARγ) results in the suppression of lox-1 expression in activated hepatic stellate cells (HSCs) in vitro. (a and b) Semi-confluent HSCs were co-transfected with the plasmid pLOX-1-Luc and the plasmid pPPARγ (a), or its mutant counterpart pdn-PPARγ (b), at indicated doses. After recovery, cells were treated with or without curcumin (20 μM) for 24 h. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.; n≥6). *P<0.05 vs cells without pPPARγ, or pdn-PPARγ, (the first column on the left). ‡P<0.05 vs cells transfected with no pdn-PPARγ and treated with curcumin only (the second column). The floating schema denoted pLOX-1-Luc in use and co-transfected plasmid pPPARγ or pdn-PPARγ±curcumin in the system. (c and d) Serum-starved HSCs were treated with the natural PPARγ agonist prostaglandin J2 (PGJ2) at 0–15 μM for 24 h. (c). Real-time PCR assays of oxidized low-density lipoprotein receptor-1 (LOX-1) mRNA. mRNA fold changes were expressed as means±s.d. (n≥3). *P<0.05 vs the untreated control (the first column on the left); (d) Western blotting analyses of LOX-1. β-actin was used as an internal control for equal protein loading. Representative was from three independent experiments. Italic numbers beneath the blot were fold changes in the densities of the bands compared with the control without treatment in the blot (n=3), after normalization with β-actin. Because of the limited space, s.d. were not presented.
To further confirm the inhibitory role of PPARγ activation, serum-starved HSCs were treated with the natural PPARγ agonist 15-deoxy-Δ.12, 14-PGJ2 at 0–15 μM for 24 h. Total RNA and whole cell extracts were prepared for real-time PCR and Western blotting analyses. As showed in Figure 5c and d, the activation of PPARγ by PGJ2 dose-dependently reduced the levels of the transcript and protein of LOX-1. Taken together, our data demonstrated that the activation of PPARγ resulted in the suppression of lox-1 expression in activated HSCs in vitro.
Identification of Putative Curcumin Response Element(s) in the lox-1 Promoter
Additional experiments were conducted to elucidate the molecular mechanisms by which curcumin inhibited the lox-1 promoter activity in HSCs. To localize curcumin response element(s) in the lox-1 promoter, a group of luciferase reporter plasmids with various lengths of the lox-1 promoter region, including 2336, 996, 289, 259 and 99 bp nucleotides, were used. Passaged HSCs were transfected with the group of luciferase reporter plasmids. After recovery, cells were treated with or without curcumin at 20 μM for 24 h. As shown in Figure 6a, compared with the corresponding untreated control, curcumin significantly reduced luciferase activities by approximately 50% in cells transfected with a plasmid containing a 5′-flanking fragment longer than −289 bp nucleotides of the lox-1 promoter. However, loss of the promoter region between −289 and −259 bp nucleotides in p259-Luc resulted in a marked reduction in luciferase activities and in response to curcumin. Although cells transfected with p259 or pp99 still showed the response to the inhibitory effect of curcumin, the difference in luciferase activities between cells with or without curcumin treatment was dramatically reduced approximately from 50 to 30%. We could not exclude the presence of additional curcumin response element(s) within the promoter region of −259 bp, even −99 bp. However, our results suggested that a major curcumin response element(s) might be located within the fragment between −289 and −259 bp, which also controlled the basal transcription activity of the promoter. We, therefore, paid our major attention to this region and studied the major curcumin response element(s) in the lox-1 promoter.
Identification of putative curcumin response elements in the lox-1 promoter in hepatic stellate cells (HSCs). Semi-confluent HSCs were transiently transfected with a group of oxidized low-density lipoprotein receptor-1 (LOX-1) promoter luciferase reporter plasmids. A total of 3.5 μg of plasmid DNA per well was added to HSCs in 6-well culture plates. It included 3 μg of a LOX-1 promoter reporter plasmid and 0.5 μg of pSV-β-gal. After recovery, cells were treated with or without curcumin at 20 μM for 24 h. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.; n≥6). The percentages indicated the reduction in luciferase activities in cells treated with curcumin, compared with those in corresponding cells without curcumin treatment. (a) Luciferase activity assays of cells transfected with a group of plasmids with various lengths of the lox-1 5′-flanking promoter region. (b) Luciferase activity assays of HSCs transfected with the wild-type pLOX-1-Luc, or its mutant counterpart pLOX-1(mut)-Luc with site-directed mutations in the putative TCF–LEF-1 binding site.
Computer-aided analyses found a putative TCF–LEF binding site, that is, CTTTGA, at −275 to −269 bp within the promoter region. TCF–LEF binding site is the target of canonical Wnt signaling and bound by a complex of β-catenin with TCF–LEF, mediating the regulation of target gene transcription.42, 43 The plasmid pLOX-1(mut)-Luc with site-directed mutation in the TCF–LEF-1 binding site was generated from the parental plasmid pLOX-1-Luc. Passaged HSCs were transfected with pLOX-1-Luc or pLOX-1(mut)-Luc and subsequently treated with or without curcumin (20 μM) for 24 h. Luciferase activity assays in Figure 6b demonstrated that compared with the untreated control, curcumin significantly reduced, as expected, luciferase activities by 50.5% in cells transfected with wild-type pLOX-1-Luc. However, the site-directed mutation of the TCF–LEF-1 binding site in pLOX-1(mut)-Luc resulted in a significant reduction approximately from 50 to 25% in the difference of luciferase activities between cells treated with or without curcumin, indicating a significant loss in response to curcumin. In addition, compared with pLOX-1-Luc, pLOX-1(mut)-Luc also led to an apparent reduction in luciferase activities in cells without curcumin treatment, suggesting an important role of the TCF–LEF-1 binding site in regulating the basal promoter activity of LOX-1 in HSCs. Taken together, these results suggested that the TCF–LEF-1 binding site might be responsible for both controlling the basal promoter activity as well as responding to the curcumin treatment. It might be a major curcumin response element, which played a critical role in the curcumin-caused inhibition of the lox-1 promoter activity.
The TCF–LEF-1 Binding Site in the lox-1 Promoter Mediates Canonical Wnt Signaling and PPARγ in the Regulation of the lox-1 Promoter Activity in HSCs
Our earlier results demonstrated the roles of curcumin and PPARγ activation in the inhibition of the lox-1 promoter activity and gene expression (Figures 3, 4 and 5). Results in Figure 6 suggested that the TCF–LEF-1 binding site was the curcumin response element, which might mediate the curcumin-caused inhibition of the lox-1 promoter activity. To explore the underlying mechanisms, it is plausible to hypothesize that the curcumin-activated PPARγ interrupted canonical Wnt signaling, leading to the reduction in lox-1 promoter activity and to the inhibition of lox-1 expression in HSCs. To test the hypothesis, HSCs were co-transfected with the plasmid pLOX-1-Luc, or the mutant plasmid pLOX-1(mut)-Luc, and the PPARγ cDNA expression plasmid pPPARγ or the empty control vector pcDNA. After recovery, cells were cultured for 24 h in DMEM with FBS (10%), which contained enough agonists to activate PPARγ in HSCs.19, 23, 24 Luciferase activity assays in Figure 7a demonstrated that forced expression of PPARγ from pPPARγ significantly reduced luciferase activity by 63.4% in cells co-transfected with the wild-type plasmid pLOX-1-Luc (the upper black column), compared with the empty control vector pcDNA (the upper white column). In great contrast, forced expression of PPARγ from pPPARγ failed to reduce luciferase activity in cells co-transfected with the mutant plasmid pLOX-1(mut)-Luc (the lower black column), compared with the empty control vector pcDNA (the lower white column). Luciferase activity was much lower in cells transfected with pLOX-1(mut)-Luc (the lower white column), compared with that in cells transfected with the wild-type pLOX-1-Luc (the upper white column), confirming the role of the TCF–LEF-1 binding site in controlling the basal activity of the gene promoter observed in Figure 6.
The TCF–LEF-1 binding site in the lox-1 promoter mediates Wnt signaling and peroxisome proliferator-activated receptor-gamma (PPARγ) in the regulation of the lox-1 promoter activity in hepatic stellate cells (HSCs). (a). Luciferase activity assays of HSCs co-transfected with pLOX-1-Luc, or pLOX-1(mut)-Luc, plus pPPARγ, or the empty control vector pcDNA. A total of 4.5 μg of plasmid DNA per well was used for co-transfection of HSCs in 6-well culture plates. It included 2 μg of pLOX-1-Luc, or pLOX-1(mut)-Luc, 0.5 μg of pSV-β-gal and 2.0 μg of pPPARγ or pcDNA. After recovery, cells were cultured in DMEM with FBS (10%) for 24 h. (b) Luciferase activity assays of HSCs transfected with pLOX-1-Luc, or pLOX-1(mut)-Luc. Cells were treated with or without Wnt3a (50 ng/ml) in serum-free medium for 24 h. A total of 3.5 μg of plasmid DNA per well was added to HSCs in 6-well culture plates. It included 3 μg of pLOX-1-Luc, or pLOX-1(mut)-Luc, and 0.5 μg of pSV-β-gal. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.; n≥6). The percentages indicated the changes in luciferase activities, compared with corresponding control cells (the white column).
To further test our hypothesis, HSCs were transfected with pLOX-1-Luc, or the mutant pLOX-1(mut)-Luc. After overnight recovery, cells were serum-starved for 24 h before the stimulation with or without exogenous Wnt3a (50 ng/ml), a stimulator of canonical Wnt signaling, in serum-free medium for another 24 h. As shown in Figure 7b by luciferase activity assays, exogenous Wnt3a significantly increased luciferase activity by 38.5% in cells transfected with the wild-type pLOX-1-Luc. However, Wnt3a showed no stimulatory effect on luciferase activity in cells transfected with the mutant pLOX-1(mut)-Luc, verifying the critical role of the TCF–LEF-1 binding site in regulating lox-1 promoter activity in HSCs. These results collectively indicated that the TCF–LEF-1 binding site in lox-1 promoter mediated both canonical Wnt signaling and PPARγ in the regulation of lox-1 promoter activity in HSCs.
Activation of Canonical Wnt Signaling Induces lox-1 Expression, which is Dose-Dependently Eliminated by Curcumin in Activated HSCs
Additional experiments were conducted to clarify the effect of canonical Wnt signaling on regulating lox-1 expression in activated HSCs. HSCs were transfected with the lox-1 promoter luciferase reporter plasmid pLOX-1-Luc. After recovery, cells were serum-starved for 3 h before the stimulation with Wnt3a at 0–100 ng/ml in serum-free medium for additional 24 h. Luciferase activity assays demonstrated that the activation of canonical Wnt signaling by exogenous Wnt3a dose-dependently increased luciferase activities in the cells (Figure 8a), suggesting the stimulatory effect of Wnt signaling on the lox-1 promoter activity. This observation was confirmed by Western blotting analyses in serum-starved HSCs treated with Wnt3a at 0–100 ng/ml in serum-depleted media for 24 h (Figure 8b). Wnt3a dose-dependently increased the abundance of LOX-1 in the cells.
Activation of canonical Wnt signaling induces lox-1 expression in activated hepatic stellate cells (HSCs), which is dose-dependently eliminated by curcumin. (a) HSCs were transiently transfected with pLOX-1-Luc and subsequently treated with Wnt3a at indicated concentrations in serum-free medium for 24 h. *P<0.05 vs cells with no treatment (the first column). Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.) (n=6). The floating schema denoted pLOX-1-Luc in use and the application of Wnt3a to the system. (b and c) Western blotting analyses of oxidized low-density lipoprotein receptor-1 (LOX-1) in HSCs treated with Wnt3a at indicated concentrations in the absence (b) or presence (c) of curcumin at various concentrations in serum-depleted media for 24 h. β-actin was used as an internal control for equal protein loading. Representative was from three independent experiments. Italic numbers beneath the blot were fold changes in the densities of the bands compared with the control without treatment in the blot (n=3), after normalization with β-actin. Because of the limited space, s.d. were not presented.
Further experiments examined whether curcumin could antagonize the stimulatory effect of Wnt3a on lox-1 expression. Serum-starved HSCs were pretreated with curcumin at different concentrations for 30 min before the stimulation with exogenous Wnt3a (50 ng/ml) for 24 h. Whole cell extracts were prepared for Western blotting analyses. It was revealed that curcumin dose-dependently attenuated the stimulatory effect of Wnt3a on the abundance of LOX-1 in activated HSCs (Figure 8c). Taken together, these results demonstrated that the activation of Wnt signaling induced lox-1 expression in activated HSCs in vitro, which was dose- dependently eliminated by curcumin.
Curcumin Interrupts Canonical Wnt Signaling in Activated HSCs in vitro
To explore the mechanisms by which curcumin eliminated the stimulatory effect of Wnt signaling on the induction of lox-1 expression, we assumed that the activation of PPARγ by curcumin antagonistically interacted with the Wnt signaling pathway and led to the interruption of canonical Wnt signaling in activated HSCs. To test the assumption, passaged HSCs were transfected with the plasmid TOPflash or FOPflash. TOPflash was a canonical Wnt signaling luciferase reporter, which contained eight copies of TCF–LEF binding sites.33 FOPflash was used as a control luciferase reporter, which contained eight copies of mutant TCF–LEF binding sites.33 After recovery, cells were treated with curcumin at various concentrations (0–30 μM) in DMEM with FBS (10%) for 24 h. Luciferase activity assays in Figure 9a demonstrated that curcumin caused a dose-dependent reduction in luciferase activities in cells transfected with TOPflash. However, curcumin had no effect on luciferase activities in cells transfected with FOPflash. These results suggested that curcumin interrupted canonical Wnt signaling in HSCs.
Curcumin interrupts canonical Wnt signaling in activated hepatic stellate cells (HSCs) in vitro. (a) Luciferase activity assays of HSCs transiently transfected with the plasmid TOPflash or FOPflash, followed by the treatment with curcumin at indicated concentrations for 24 h. (n=6). *P<0.05 vs cells with no treatment (the first column). The floating schema denoted the canonical Wnt signaling luciferase reporter construct TOPflash or its mutant counterpart FOPflash in use and the application of curcumin to the system. (b) Semi-confluent HSCs were treated with curcumin at indicated concentrations for 24 h. Total nuclear extracts were prepared for Western blotting analyses of β-catenin. Histone H1 was used as an invariant control for equal nuclear protein loading. Representative was from three independent experiments. Italic numbers beneath the blot were fold changes in the densities of the bands compared with the control without treatment in the blot (n=3), after normalization with Histone H1. Because of the limited space, standard deviations were not presented. (c) Electrophoretic mobility shift assay (EMSA) of nuclear protein extracts from HSCs treated with various concentrations of curcumin using the biotin-labeled probe P(TCFwt), which contained the consensus TCF–LEF binding site found in the lox-1 promoter. (d) EMSA competition assays of nuclear protein extracts from HSCs treated with or without curcumin (Cur) at 20 μM using the biotin-labeled probe P(TCFwt) and a 10-, 50-, or 100-fold excess of the unlabeled P(TCFwt) (lanes 3–5), or an unlabeled probe P(TCFmut) (lanes 6–8). The latter probe contained the consensus TCF–LEF binding site found in the lox-1 promoter with site-directed mutations. Representatives of EMSA were shown from three independent experiments.
Canonical Wnt signaling is activated only when dephosphorylated β-catenin is translocated into the nucleus.42 To verify the effect of curcumin on interrupting the Wnt signaling pathway, HSCs were treated with curcumin at various concentrations (0–30 μM) for 24 h. Total nuclear proteins were prepared and the abundance of nuclear β-catenin was evaluated by Western blotting analyses. It was revealed in Figure 9b that curcumin dose-dependently reduced the abundance of nuclear β-catenin. The nuclear protein histone H1 was used as an invariant control.
Nuclear β-catenin forms a complex with the transcription factor TCF–LEF to bind to the promoter of target genes and to stimulate target gene transcription.42, 43 EMSAs were carried out to further verify the effect of curcumin on interrupting canonical Wnt signaling. Nuclear protein extracts from HSCs treated with curcumin at 0–30 μM for 24 h were incubated with biotin-labeled double-stranded oligonucleotide probe P(TCFwt). The probe P(TCFwt) contained the consensus TCF–LEF binding site found in the lox-1 promoter. As shown in Figure 9c, curcumin dose-dependently reduced the DNA binding activity of the transcription factor TCF–LEF to the probe. To examine the DNA binding specificity of TCF–LEF to the probe, competition assays were carried out using a 10-, 50-, or 100-fold excess of the unlabeled probe P(TCFwt) (lanes 3–5 in Figure 9d), or the unlabeled probe P(TCFmut) (lanes 6–8 in Figure 9d). The probe P(TCFmut) contained the TCF–LEF binding site found in the lox-1 promoter with site-directed mutations. It was found that the amount of TCF–LEF binding to the biotin-labeled probe P(TCFwt)) was competitively reduced by the unlabeled probe P(TCFwt), but not by the mutant probe P(TCFwt). These results indicated that curcumin dose- dependently reduced the DNA binding activity of TCF–LEF to the promoter of LOX-1 gene in activated HSCs. Taken together, these results indicated that curcumin interrupted canonical Wnt signaling in activated HSCs by reducing the abundance of nuclear β-catenin and reducing the DNA binding activity of TCF–LEF.
The Activation of PPARγ by Curcumin Results in the Interruption of Canonical Wnt Signaling in Activated HSCs in vitro
Additional experiments were conducted to further test our above assumption. To evaluate the role of PPARγ activation in the curcumin-caused interruption of canonical Wnt signaling, HSCs were transfected with the plasmid TOPflash or FOPflash. The cells were pretreated with the PPARγ antagonist PD68235 (20 μM) for 30 min before the addition of curcumin (20 μM) for additional 24 h. Luciferase activity assays in Figure 10a indicated that compared with the untreated control (the first black column on the left), curcumin reduced, as expected, luciferase activities in cells transfected with TOPflash (the second black column). The pretreatment with PD68235 significantly attenuated the inhibitory effect of curcumin (the fourth black column). Curcumin with or without PD68235 caused no significant changes in luciferase activities in cells transfected with the control plasmid FOPflash (white columns). These results suggested that the activation of PPARγ had a critical role in the curcumin-caused interruption of canonical Wnt signaling in HSCs.
The activation of peroxisome proliferator-activated receptor-gamma (PPARγ) by curcumin interrupts canonical Wnt signaling in activated hepatic stellate cells (HSCs) in vitro. (a and b) HSCs were transiently transfected with the plasmid TOPflash or FOPflash. After recovery, cells were incubated in DMEM with FBS (10%) with treatment for 24 h. *P<0.05 vs cells with no treatment (the first column). Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.) (n=6). The floating schemas denote the plasmid TOPflash or its mutant FOPflash in use and the application of treatments to the system. (a) Luciferase activity assays of cells pretreated with PD68235 (20 μM) for 30 min before the addition of curcumin (20 μM) for additional 24 h. (b) Luciferase activity assays of cells co-transfected with pPPARγ at indicated doses. A total of 4.5 μg of plasmid DNA per well was used for co-transfection of HSCs in 6-well culture plates. It included 2 μg of TOPflash, or FOPflash, 0.5 μg of pSV-β-gal and 2.0 μg of pPPARγ plus pcDNA. The latter was used to ensure an equal amount of total DNA in transfection assays. (c) HSCs were transfected with TOPflash and treated with prostaglandin J2 (PGJ2) at indicated doses in serum-depleted medium for 24 h. Luciferase activities were expressed as relative units after β-galactosidase normalization (means±s.d.) (n=6). *P<0.05 vs cells with no treatment (the first column). The floating schema denoted the plasmid TOPflash in use and the application of PGJ2 to the system.
To further confirm the inhibitory role of PPARγ activation in the interruption of canonical Wnt signaling, HSCs were co-transfected with the plasmid TOPflash or FOPflash, plus the cDNA expression plasmid pPPARγ. Transfected cells were cultured for 24 h in media with 10% FBS, which contains enough agonists to activate PPARγ in HSCs.19, 23, 24 Luciferase activity assays in Figure 10b revealed that forced expression of PPARγ dose dependent reduced luciferase activities in cells transfected with TOPflash, but not in cells transfected with FOPflash, confirming that the activation of PPARγ interrupted canonical Wnt signaling in HSCs. Additional experiments verified these observations and demonstrated that the activation of PPARγ by PGJ2 dose-dependently reduced luciferase activities in HSCs transfected with TOPflash (Figure 10c). Taken together, these results indicated that the activation of PPARγ by curcumin resulted in the interruption of canonical Wnt signaling in activated HSCs in vitro.
DISCUSSION
Non-alcoholic steatohepatitis patients are often associated with high levels of lipid peroxidation products, such as ox-LDL.44 It has been suggested that the elevated levels of lipid peroxidation might make an important contribution to the pathogenesis of NASH,45 leading to hepatic fibrosis. In the current report, we demonstrated that ox-LDL stimulated the activation of HSCs in vitro, which was attenuated by curcumin by suppressing gene expression of LOX-1 via the activation of PPARγ and the interruption of Wnt signaling.
The precise range of plasma ox-LDL concentrations in human remains unclear. Previous studies suggested that ox-LDL was only a minor fraction of LDL ranging from 0.001% in healthy controls46 to approximately 5% in patients with acute coronary events.47 A recent report indicated that the level of ox-LDL in healthy human was in 0.58±0.23 ng/5 μg LDL protein.48 However, it bears emphasis that because the in vivo system is multi-factorial, directly extrapolating in vitro conditions and results, for example effective concentrations, to the in vivo system might be misleading. ox-LDL at 10 μg/ml was chosen for most of our experiments. It showed no apparent cytotoxicity to cultured HSCs (data not presented here), though higher concentrations (>25 μg/ml) of ox-LDL might result in the reduction in the production of ECM components and even cytotoxicity to cultured HSCs.36
Uptake and transport of ox-LDL into cells is mediated by a variety of scavenger receptors, including LOX-1, SR-AI/II, CD36, and SR-BI.49 Depending on cell types, these scavenger receptors show distinct expression profiles. For example, CD36 is highly expressed in macrophages, and has a fundamental role in the uptake of ox-LDL in macrophages,50 whereas LOX-1 is the principal ox-LDL receptor in epithelial cells.7 CD36 is a multiligand scavenger receptor that recognizes and binds many ligands, including ox-LDL and long-chain free fatty acids.51 Our preliminary experiments indicated that compared with that of CD36, the expression of LOX-1 in activated HSCs is more inducible and highly expressed at both transcript and protein levels (unpublished observations), suggesting that LOX-1 might have a more important role in the uptake and transport of extracellular ox-LDL into activated HSCs. We, therefore, focused our primary attention to the effects of curcumin on LOX-1 in this study. It bears emphasis that we do not exclude the roles of CD36 and other scavenger receptors in the uptake and transport of ox-LDL into HSCs and their effects on the ox-LDL-induced HSC activation. On the other hand, we could not exclude the role of curcumin in inhibiting gene expression of CD36 and other scavenger receptors in HSCs. Our preliminary experiments indicated that curcumin indeed suppressed gene expression of CD36 in activated HSCs in vitro (data not shown), which might collectively contribute to the curcumin-caused reduction in the level of intracellular ox-LDL observed in Figure 3d.
During HSC activation, the abundance of PPARγ is significantly reduced. We have previously shown that curcumin dramatically induces expression of endogenous PPARγ gene and its activity in cultured HSCs, which is required for curcumin to inhibit HSC activation.23, 24, 27 We have also observed that although significantly reduced, PPARγ is still detectable in cultured HSCs and responds to the stimulation with PGJ2, a natural PPARγ agonist.23, 38 To evaluate the role of PPARγ activation in the curcumin-caused inhibition of LOX-1 gene expression in cultured HSCs, PPARγ activation was blocked by its specific antagonist PD68235. Our results in Figure 4 indicated that the blockade of PPARγ activation apparently eliminated the role of curcumin in inhibiting gene expression of LOX-1 in cultured HSCs. Additional experiments revealed that forced expression of PPARγ cDNA dose- dependently reduced the gene promoter activity of LOX-1 (Figure 5a). On the other hand, forced expression of dominant negative PPARγ cDNA dose-dependently eliminated the effect of curcumin on reducing the promoter activity of LOX-1 gene (Figure 5b). Furthermore, the activation of PPARγ by its natural agonist PGJ2 dose-dependently suppressed gene expression of LOX-1 at both levels of transcription and translation (Figure 5c and d). Taken together, our results demonstrated that the activation of PPARγ had a critical role in the curcumin-caused inhibition of LOX-1 gene expression in activated HSCs in vitro. Our current observations are consistent with previous other reports.6, 52 The activation of PPARγ by its ligand pioglitazone inhibited LOX-1 expression in coronary artery endothelial cells and fibroblasts.6, 52 In addition, PPARγ activation inhibited TNF-α-induced LOX-1 expression in aortic endothelial cells.53 On the other hand, the stimulatory effect of PPARγ on lox-1 expression was also reported.54 PPARγ bound to the peroxisome proliferator response elements (PPREs) in the lox-1 promoter and stimulated LOX-1 gene transcription and expression in adipocytes.54 These observations indicate that the roles of PPARγ in the regulation of lox-1 expression are divergent and might greatly depend on cell types. Additional experiments are necessary to clarify the underlying mechanisms.
In this study, we reported that the activation of PPARγ by curcumin antagonized canonical Wnt signaling (Figure 10), likely leading to the suppression of the lox-1 expression and the inhibition of ox-LDL-induced HSC activation. To determine the role of Wnt signaling in the regulation of LOX-1 gene expression and in the curcumin-caused inhibition of LOX-1 gene expression in activated HSCs in vitro, we observed that the activation of Wnt signaling by its agonist Wnt3a dose-dependently induced gene expression of LOX-1 in HSCs, including increasing the promoter activity (Figure 8a) and elevating the abundance of LOX-1 protein (Figure 8b). In addition, curcumin dose-dependently eliminated the stimulatory effect of the Wnt signaling activated by Wnt3a on expression of LOX-1 gene in cultured HSCs (Figure 8c). Additional experiments revealed that curcumin interrupted Wnt signaling (Figure 9a), reduced the abundance of nuclear β-catenin (Figure 9b), decreased the DNA binding activity of Wnt signaling target TCF (Figure 9c). Taken together, our results demonstrated that the interruption of Wnt signaling pathway had a key role in the curcumin-caused inhibition of LOX-1 gene expression in activated HSCs in vitro.
In contrast to non-canonical β-catenin-independent Wnt signaling, canonical Wnt signaling involves a series of events that is initiated by binding an extracellular Wnt protein, including Wnt3a, to a member of the family of Frizzled transmembrane receptors.42, 43 This interaction activates the Dishevelled protein by hyper-phosphorylation, which results in the prevention of the phosphorylation of β-catenin by the degradation complex (GSK-3, APC, and axin) and the release of β-catenin from the degradation complex. Free β-catenin forms a complex with TCF–LEF and translocates into the nucleus, stimulating target gene transcription.42, 43 Although its effect remains controversial,55 accumulating evidence has indicated that activation of the canonical Wnt signaling pathway stimulates HSC activation and hepatic fibrogenesis.56, 57 We demonstrated in this report that curcumin significantly reduced the level of nuclear β-catenin in HSCs. In addition, the activation of PPARγ by curcumin interrupted Wnt signaling, leading to the suppression of lox-1 expression and the inhibition of ox-LDL-induced HSC activation. Our results are supported by previous observations that the activation of PPARγ antagonistically interrupted Wnt signaling by inducing the degradation of β-catenin.58, 59, 60 In addition, the interruption of Wnt signaling inhibited HSC activation.15, 16 Other studies also showed the roles of the Wnt signaling pathway and the transcription factor TCF–LEF-1 in mediating curcumin actions. For example, curcumin treatment caused p53- and p21-independent G(2)/M phase arrest and apoptosis in colon cancer cell lines. Further experiments found that curcumin caused a caspase-3-mediated cleavage of β-catenin, decreased trans-activation of β-catenin–TCF, decreased promoter DNA binding activity of the β-catenin–TCF complex, and decreased levels of c-Myc protein in the colon cancer cells.61 Studies also indicated that curcumin and its derivatives attenuated the Wnt–β-catenin pathway through downregulation of the transcriptional coactivator p300.62 It was even suggested that curcumin and its derivative were excellent inhibitors of β-catenin–TCF signaling in cancer cell lines.63 The reduced β-catenin–TCF transcriptional activity was due to the decreased nuclear β-catenin and TCF-4.63
Based on these results, we propose a simplified action model for curcumin to inhibit ox-LDL-induced HSC activation (Figure 11). Extracellular ox-LDL is transported into HSCs mediated by LOX-1, leading to the stimulation of HSC activation. This process is blocked by the curcumin-caused suppression of LOX-1 gene expression by activating PPARγ and interrupting Wnt signaling. It bears emphasis that the underlying mechanisms are certainly more complex than what is described here. In addition, our results do not exclude the possible involvement of any other signaling pathways and mechanisms in the curcumin-caused suppression of LOX-1 gene expression and in the inhibition of ox-LDL-induced HSC activation. Our observations provide new insights into the molecular mechanisms of curcumin in inhibiting ox-LDL-induced HSC activation and offer a natural anti-fibrotic candidate for the therapeutic treatment and prevention of T2DM and NASH-associated hepatic fibrosis.
A simplified action model for curcumin to inhibit oxidized low-density lipoprotein (ox-LDL)-induced hepatic stellate cell (HSC) activation. Extracellular ox-LDL is transported into HSCs, mediated by oxidized low-density lipoprotein receptor-1 (LOX-1), leading to the stimulation of HSC activation. This process is blocked by the curcumin-caused suppression of LOX-1 gene expression by activating peroxisome proliferator-activated receptor-gamma (PPARγ) and interrupting Wnt signaling.
Accession codes
References
Maiese K, Morhan SD, Chong ZZ . Oxidative stress biology and cell injury during type 1 and type 2 diabetes mellitus. Curr Neurovasc Res 2007;4:63–71.
Tsochatzis E, Papatheodoridis GV, Manesis EK, et al. Metabolic syndrome is associated with severe fibrosis in chronic viral hepatitis and non-alcoholic steatohepatitis. Aliment Pharmacol Ther 2008;27:80–89.
Jacqueminet S, Lebray P, Morra R, et al. Screening for liver fibrosis by using a noninvasive biomarker in patients with diabetes. Clin Gastroenterol Hepatol 2008;6:828–831.
Clark JM . The epidemiology of nonalcoholic fatty liver disease in adults. J Clin Gastroenterol 2006;40 (Suppl 1):S5–S10.
Holvoet P, Lee DH, Steffes M, et al. Association between circulating oxidized low-density lipoprotein and incidence of the metabolic syndrome. JAMA 2008;299:2287–2293.
Mehta JL, Chen J, Hermonat PL, et al. Lectin-like, oxidized low-density lipoprotein receptor-1 (LOX-1): a critical player in the development of atherosclerosis and related disorders. Cardiovasc Res 2006;69:36–45.
Sawamura T, Kume N, Aoyama T, et al. An endothelial receptor for oxidized low-density lipoprotein. Nature 1997;386:73–77.
Dunn S, Vohra RS, Murphy JE, et al. The lectin-like oxidized low-density-lipoprotein receptor: a pro-inflammatory factor in vascular disease. Biochem J 2008;409:349–355.
Li D, Mehta JL . Upregulation of endothelial receptor for oxidized LDL (LOX-1) by oxidized LDL and implications in apoptosis of human coronary artery endothelial cells: evidence from use of antisense LOX-1 mRNA and chemical inhibitors. Arterioscler Thromb Vasc Biol 2000;20:1116–1122.
Bataller R, Brenner DA . Liver fibrosis. J Clin Invest 2005;115:209–218.
Friedman SL . Mechanisms of hepatic fibrogenesis. Gastroenterology 2008;134:1655–1669.
Friedman SL, Yamasaki G, Wong L . Modulation of transforming growth factor beta receptors of rat lipocytes during the hepatic wound healing response. Enhanced binding and reduced gene expression accompany cellular activation in culture and in vivo. J Biol Chem 1994;269:10551–10558.
Pinzani M, Milani S, Herbst H, et al. Expression of platelet-derived growth factor and its receptors in normal human liver and during active hepatic fibrogenesis. Am J Pathol 1996;148:785–800.
Cheng JH, She H, Han YP, et al. Wnt antagonism inhibits hepatic stellate cell activation and liver fibrosis. Am J Physiol Gastrointest Liver Physiol 2008;294:G39–G49.
Jiang F, Parsons CJ, Stefanovic B . Gene expression profile of quiescent and activated rat hepatic stellate cells implicates Wnt signaling pathway in activation. J Hepatol 2006;45:401–409.
Myung SJ, Yoon JH, Gwak GY, et al. Wnt signaling enhances the activation and survival of human hepatic stellate cells. FEBS Lett 2007;581:2954–2958.
Galli A, Crabb D, Price D, et al. Peroxisome proliferator-activated receptor gamma transcriptional regulation is involved in platelet-derived growth factor-induced proliferation of human hepatic stellate cells. Hepatology 2000;31:101–108.
Marra F, Efsen E, Romanelli RG, et al. Ligands of peroxisome proliferator-activated receptor gamma modulate profibrogenic and proinflammatory actions in hepatic stellate cells. Gastroenterology 2000;119:466–478.
Miyahara T, Schrum L, Rippe R, et al. Peroxisome proliferator-activated receptors and hepatic stellate cell activation. J Biol Chem 2000;275:35715–35722.
Joe B, Vijaykumar M, Lokesh BR . Biological properties of curcumin-cellular and molecular mechanisms of action. Crit Rev Food Sci Nutr 2004;44:97–111.
O’Connell MA, Rushworth SA . Curcumin: potential for hepatic fibrosis therapy? Br J Pharmacol 2008;153:403–405.
Fu Y, Zheng S, Lin J, et al. Curcumin protects the rat liver from CCl4-caused injury and fibrogenesis by attenuating oxidative stress and suppressing inflammation. Mol Pharmacol 2008;73:399–409.
Xu J, Fu Y, Chen A . Activation of peroxisome proliferator-activated receptor-gamma contributes to the inhibitory effects of curcumin on rat hepatic stellate cell growth. Am J Physiol Gastrointest Liver Physiol 2003;285:G20–G30.
Zheng S, Chen A . Activation of PPARgamma is required for curcumin to induce apoptosis and to inhibit the expression of extracellular matrix genes in hepatic stellate cells in vitro. Biochem J 2004;384: 149–157.
Zheng S, Chen A . Curcumin suppresses the expression of extracellular matrix genes in activated hepatic stellate cells by inhibiting gene expression of connective tissue growth factor. Am J Physiol Gastrointest Liver Physiol 2006;290:G883–G893.
Zheng S, Chen A . Disruption of transforming growth factor-beta signaling by curcumin induces gene expression of peroxisome proliferator-activated receptor-gamma in rat hepatic stellate cells. Am J Physiol Gastrointest Liver Physiol 2007;292:G113–G123.
Zheng S, Yumei F, Chen A . De novo synthesis of glutathione is a prerequisite for curcumin to inhibit hepatic stellate cell (HSC) activation. Free Radic Biol Med 2007;43:444–453.
Chen A, Davis BH . UV irradiation activates JNK and increases alphaI (I) collagen gene expression in rat hepatic stellate cells. J Biol Chem 1999;274:158–164.
Chen J, Liu Y, Liu H, et al. Lectin-like oxidized low-density lipoprotein receptor-1 (LOX-1) transcriptional regulation by Oct-1 in human endothelial cells: implications for atherosclerosis. Biochem J 2006;393:255–265.
Ballagi AE, Ishizaki A, Nehlin JO, et al. Isolation and characterization of the mouse PDGF beta-receptor promoter. Biochem Biophys Res Commun 1995;210:165–173.
Ji C, Casinghino S, McCarthy TL, et al. Cloning, characterization, and expression of the transforming growth factor-beta type I receptor promoter in fetal rat bone cells. J Cell Biochem 1996;63:478–490.
Gurnell M, Wentworth JM, Agostini M, et al. A dominant-negative peroxisome proliferator-activated receptor gamma (PPARgamma) mutant is a constitutive repressor and inhibits PPARgamma-mediated adipogenesis. J Biol Chem 2000;275:5754–5759.
DasGupta R, Kaykas A, Moon RT, et al. Functional genomic analysis of the Wnt-wingless signaling pathway. Science 2005;308:826–833.
Chen A, Zhang L, Xu J, et al. The antioxidant (−)-epigallocatechin-3-gallate inhibits activated hepatic stellate cell growth and suppresses acetaldehyde-induced gene expression. Biochem J 2002;368:695–704.
Schmittgen TD, Zakrajsek BA, Mills AG, et al. Quantitative reverse transcription-polymerase chain reaction to study mRNA decay: comparison of endpoint and real-time methods. Anal Biochem 2000;285:194–204.
Schneiderhan W, Schmid-Kotsas A, Zhao J, et al. Oxidized low-density lipoproteins bind to the scavenger receptor, CD36, of hepatic stellate cells and stimulate extracellular matrix synthesis. Hepatology 2001;34:729–737.
Lin J, Chen A . Activation of peroxisome proliferator-activated receptor-gamma by curcumin blocks the signaling pathways for PDGF and EGF in hepatic stellate cells. Lab Invest 2008;88:529–540.
Zhou Y, Zheng S, Lin J, et al. The interruption of the PDGF and EGF signaling pathways by curcumin stimulates gene expression of PPARgamma in rat activated hepatic stellate cell in vitro. Lab Invest 2007;87:488–498.
Draude G, Hrboticky N, Lorenz RL . The expression of the lectin-like oxidized low-density lipoprotein receptor (LOX-1) on human vascular smooth muscle cells and monocytes and its down-regulation by lovastatin. Biochem Pharmacol 1999;57:383–386.
Duerrschmidt N, Zabirnyk O, Nowicki M, et al. Lectin-like oxidized low-density lipoprotein receptor-1-mediated autophagy in human granulosa cells as an alternative of programmed cell death. Endocrinology 2006;147:3851–3860.
Thum T, Borlak J . Mechanistic role of cytochrome P450 monooxygenases in oxidized low-density lipoprotein-induced vascular injury: therapy through LOX-1 receptor antagonism? Circ Res 2004;94:e1–e13.
Shitashige M, Hirohashi S, Yamada T . Wnt signaling inside the nucleus. Cancer Sci 2008;99:631–637.
Vlad A, Rohrs S, Klein-Hitpass L, et al. The first five years of the Wnt targetome. Cell Signal 2008;20:795–802.
Chalasani N, Deeg MA, Crabb DW . Systemic levels of lipid peroxidation and its metabolic and dietary correlates in patients with non-alcoholic steatohepatitis. Am J Gastroenterol 2004;99:1497–1502.
James O, Day C . Non-alcoholic steatohepatitis: another disease of affluence. Lancet 1999;353:1634–1636.
Shoji T, Nishizawa Y, Fukumoto M, et al. Inverse relationship between circulating oxidized low-density lipoprotein (oxLDL) and anti-oxLDL antibody levels in healthy subjects. Atherosclerosis 2000;148:171–177.
Holvoet P, Vanhaecke J, Janssens S, et al. Oxidized LDL and malondialdehyde-modified LDL in patients with acute coronary syndromes and stable coronary artery disease. Circulation 1998;98:1487–1494.
Ehara S, Ueda M, Naruko T, et al. Elevated levels of oxidized low-density lipoprotein show a positive relationship with the severity of acute coronary syndromes. Circulation 2001;103:1955–1960.
Dhaliwal BS, Steinbrecher UP . Scavenger receptors and oxidized low-density lipoproteins. Clin Chim Acta 1999;286:191–205.
Nozaki S, Kashiwagi H, Yamashita S, et al. Reduced uptake of oxidized low density lipoproteins in monocyte-derived macrophages from CD36-deficient subjects. J Clin Invest 1995;96:1859–1865.
Adachi H, Tsujimoto M . Endothelial scavenger receptors. Prog Lipid Res 2006;45:379–404.
Mehta JL, Hu B, Chen J, et al. Pioglitazone inhibits LOX-1 expression in human coronary artery endothelial cells by reducing intracellular superoxide radical generation. Arterioscler Thromb Vasc Biol 2003;23:2203–2208.
Chiba Y, Ogita T, Ando K, et al. PPARgamma ligands inhibit TNF-alpha-induced LOX-1 expression in cultured endothelial cells. Biochem Biophys Res Commun 2001;286:541–546.
Chui PC, Guan HP, Lehrke M, et al. PPARgamma regulates adipocyte cholesterol metabolism via oxidized LDL receptor 1. J Clin Invest 2005;115:2244–2256.
Kordes C, Sawitza I, Haussinger D . Canonical Wnt signaling maintains the quiescent stage of hepatic stellate cells. Biochem Biophys Res Commun 2008;367:116–123.
Thompson MD, Monga SP . WNT/beta-catenin signaling in liver health and disease. Hepatology 2007;45:1298–1305.
Tsukamoto H, She H, Hazra S, et al. Anti-adipogenic regulation underlies hepatic stellate cell transdifferentiation. J Gastroenterol Hepatol 2006;21 (Suppl 3):S102–S105.
Farmer SR . Regulation of PPARgamma activity during adipogenesis. Int J Obes (Lond) 2005;29 (Suppl 1):S13–S16.
Liu J, Wang H, Zuo Y, et al. Functional interaction between peroxisome proliferator-activated receptor gamma and beta-catenin. Mol Cell Biol 2006;26:5827–5837.
Moldes M, Zuo Y, Morrison RF, et al. Peroxisome-proliferator-activated receptor gamma suppresses Wnt/beta-catenin signalling during adipogenesis. Biochem J 2003;376:607–613.
Jaiswal AS, Marlow BP, Gupta N, et al. Beta-catenin-mediated transactivation and cell–cell adhesion pathways are important in curcumin (diferuylmethane)-induced growth arrest and apoptosis in colon cancer cells. Oncogene 2002;21:8414–8427.
Ryu MJ, Cho M, Song JY, et al. Natural derivatives of curcumin attenuate the Wnt/beta-catenin pathway through down-regulation of the transcriptional coactivator p300. Biochem Biophys Res Commun 2008;377:1304–1308.
Park CH, Hahm ER, Park S, et al. The inhibitory mechanism of curcumin and its derivative against beta-catenin/Tcf signaling. FEBS Lett 2005;579:2965–2971.
Acknowledgements
The work was supported by the grant RO1 DK 047995 from NIH/NIDDK to A Chen.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure/Conflict of interest
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Kang, Q., Chen, A. Curcumin eliminates oxidized LDL roles in activating hepatic stellate cells by suppressing gene expression of lectin-like oxidized LDL receptor-1. Lab Invest 89, 1275–1290 (2009). https://doi.org/10.1038/labinvest.2009.93
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.2009.93
Keywords
This article is cited by
-
Accumulation of free cholesterol and oxidized low-density lipoprotein is associated with portal inflammation and fibrosis in nonalcoholic fatty liver disease
Journal of Inflammation (2019)
-
Biology of pancreatic stellate cells—more than just pancreatic cancer
Pflügers Archiv - European Journal of Physiology (2017)
-
Curcumin Targets Multiple Pathways to Halt Hepatic Stellate Cell Activation: Updated Mechanisms In Vitro and In Vivo
Digestive Diseases and Sciences (2015)
-
Absence of clinical relationship between oxidized low density lipoproteins and diabetic peripheral neuropathy: a case control study
Lipids in Health and Disease (2014)
-
Protective mechanisms of medicinal plants targeting hepatic stellate cell activation and extracellular matrix deposition in liver fibrosis
Chinese Medicine (2014)