Abstract
Studies of loss of heterozygosity (LOH) play an important role in cancer research. In this paper, we developed a two-step procedure to examine LOH by comparing unpaired tumour and normal samples. In the first step we determined which chromosomes significantly differ between the two sets of samples by using nonparametric procedures. We then used the biplot data visualisation technique and homozygosity intensity estimates to determine the regions of these chromosomes that required further examination. We illustrated our method by examining 22 autosomes in samples of 95 normal controls and 14 acute lymphoblastic leukaemia patients. The genomewide scan of LOH with the Affymetrix Human Mapping 100K Set successfully identified the important tumour suppressor gene, CDKN2A, whose deletion was validated by quantitative polymerase chain reaction in multiple patients of this study.
Similar content being viewed by others
Introduction
Loss of heterozygosity (LOH) occurs when genotypes change from a heterozygous state to a hemizygous or homozygous state, where an allele or haplotype from one parent is lost. If the lost allele plays a role in tumour suppression in tumourigenesis, then its loss results in the onset of a cancer. LOH may be caused by several biological mechanisms: DNA deletion, mitotic recombination, gene conversion, and so on. LOH in cancer-related DNA regions can be identified by comparing genotypes on the same chromosomal loci in germ line cells and cancer cells from the same patient, where the genotypes are heterozygous in the former, but hemizygous or homozygous in the latter. Conventionally, several different types of genetic markers are used to identify LOH regions such as restriction fragment length polymorphisms (Knudson 1985) and short tandem repeats polymorphisms (Rubocki et al. 2000). Recently, the rapid development of biotechnique has allowed cost-effective single nucleotide polymorphism (SNP) genotyping, which provides data on more than 100 thousand SNPs for each individual (Matsuzaki et al. 2004a, b). These dense SNPs offer a higher resolution and more accurate boundaries for the identification of LOH relative to other genetic markers (Lin et al. 2004; Huang et al. 2004). In this paper, we use the Affymetrix Human Mapping 100K Set (Affymetrix, CA, USA) providing 116,204 SNPs with a median inter-marker distance of 8.5 kb to detect LOH regions across the human genome.
Classical LOH studies determine LOH by using paired normal and tumour samples from the same patient. Lin et al. (2004) use paired normal and tumour samples to directly compare SNP sites and compute the proportion of “loss” events in a region. However, the paired data are not always available in practical studies. Huang et al. (2004) propose a model-based approach that they note is applicable when paired normal samples are not available. However, this approach depends on unrealistic independence assumptions. Our situation is between the two as we have tumour samples and independent unpaired normal samples, and we develop an alternate graphical method. In contrast to approaches based on averaging across individuals, our approach allows us to examine each SNP for each individual.
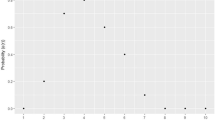
At each SNP, the genotype is either homozygous or heterozygous, and a graphical representation of raw LOH data is not revealing, although the gaps in the lower heterozygous band do indicate regions of interest. For example, in Fig. 1, we plot the homozygosity across chromosome 9 for one normal control that yields the top band, corresponding to homozygous SNPs, and bottom band, corresponding to heterozygous SNPs. Motivated by the functional data methods of Ramsay and Silverman (1997), we adopt an approach related to that of Lin et al. (2004). Functional data analysis (Ramsay and Silverman 1997) treats multivariate observations on an individual as observations on values of a function. While SNPs are discrete units, they are numerous and dense enough on the chromosome, so in practice it is reasonable to regard them as a continuum of points along the chromosome. The functional representation allows us to display the data to help detect patterns and to develop statistical procedures based on the functions themselves. To indicate the extra information available, in Fig. 1 we also plot the estimated homozygous intensity using the methods developed below. This plot gives far more information on regions of the chromosome where there may be increased or decreased homozygosity for this individual than the simple plot of homozygosity. Moreover, by estimating the underlying function we are able to compare characteristics of the chromosome between case and control subjects. To give a guide to chromosomal regions where LOH occurs in each affected individual, we consider each chromosome separately and develop a test statistic to compare homozygosity for each chromosome based on the Kullback–Leibler distance and the Wilcoxon signed rank test. Having developed a test for a given chromosome, we then adapted the gene expression biplot (GE-biplot) methodology previously applied to the visualisation of microarray data in Pittelkow and Wilson (2003).
Plot of homozygosity (1 = homozygous, 0 = heterozygous) and the estimated homozygous intensity on chromosome 9 of a normal control. Homozygosity (black points) and the estimated homozygous intensity (red points), with bandwidth 2.5%, against the SNP position (unit: Mb) of a normal control are shown. The gap from ≈45 to ≈68 Mb is the centromeric gap
The procedure is described in Sect. 2 where we consider estimation (Sect. 2.1), testing (Sect. 2.2), and data visualisation (Sect. 2.3). In Sect. 3, we apply the method to some data from acute lymphoblastic leukaemia (ALL) patients. In Sect. 4, we evaluate the performance of the proposed test using simulation studies. Section 5 contains concluding remarks of our method.
Methods
Consider a single chromosome. Let λ(t) denote the probability that a SNP at site t on this chromosome is homozygous. We call λ(t) the homozygous intensity or just the intensity. This is consistent with the approach of Lin et al. (2004). We do not observe λ(t), but rather for each individual in the sample observe a 0 (heterozygous) or 1 (homozygous) or NoCall. Here we treat the SNPs with NoCall as missing at random, and we regard them as noninformative. Thus, the observed data consist of a sequence x 1, ..., x N over the N SNP sites on the chromosome of interest, where x t takes the value 1 if the SNP at position t is homozygous, 0 if it is heterozygous, and missing if a NoCall is returned.
Estimation of the homozygous intensity
Our approach estimates the intensity of homozygosity at a given point as a weighted moving average over neighbouring points. This results in a smooth estimate of λ(t), with the smoothness depending on the weights and the size of the neighbourhood. Our model is based on local likelihood for the binomial distribution; see Chap. 4 of Loader (1999). We use the locfit package (Loader 1999) in the statistical computing language R to fit the model and take the weighted local average over the closest α percent of the SNPs to t to estimate λ(t). The locfit package is computationally efficient and allows rapid estimation of the intensities at several thousand SNPs.
Ranking the chromosomes
Estimation of the intensities allows visualisation of patterns in homozygosity across a chromosome. However, it is convenient to be able to order the chromosomes and develop a numerical measure of their degree of LOH. Let λ0 and λ1 denote intensity functions, λ0 = {λ0(t), t = 1, ..., N} and λ1 = {λ1(t), t = 1, ..., N}, where λ0(t) and λ1(t) are homozygous intensities of the controls and cases, respectively. We are interested in chromosome regions R where λ1(t) > λ0(t) for most t ∈ R. This motivates us to extend the Kullback–Leibler distance to measure the distance between two intensity functions λ0 and λ1 as follows:
This is not symmetric as we are only interested in SNPs where λ1(t) > λ0(t).
To estimate ψ(λ1,λ0), suppose we have estimated intensity functions \(\widehat\lambda_{01},\ldots,\widehat\lambda_{0n}\) for n controls and \(\widehat\lambda_{11},\ldots,\widehat\lambda_{1m}\) for m cases. Let \(\bar \lambda\) denote the estimated mean function of the pooled data from the normal controls and the cases and \(\hat\sigma\) denote the sample standard deviation function of the estimated intensities at each SNP. A nominal upper 97.5th percentile for the pooled individuals is \(U=\bar\lambda+1.96 \hat\sigma\). For each chromosome, we compute \(Y_i=\psi(\widehat\lambda_{0i},U)\) and \(Z_j=\psi(\widehat\lambda_{1j},U)\) , and compare the location of Y 1, ..., Y n and Z 1, ..., Z m using the Wilcoxon rank sum test. If the cases display more homozygosity than the controls, then the median of the Zs should be larger than the median of the Ys so we can conduct a one-sided test. We use the P value from this test to rank the chromosomes.
Data visualisation using the biplot
Pittelkow and Wilson (2003) examined the use of the biplot of Gabriel (1971) to visualise microarray data. See Jolliffe (1986) and Pittelkow and Wilson (2003) for more detailed descriptions of this approach to the visualisation of matrices. Here we employ this procedure to examine LOH on the chromosomes detected by the test developed in Sect. 2.2.
There is extensive literature on the biplots as summarised in Pittelkow and Wilson (2003). For clarity we summarise the features of interest to us. Let X be a K × p matrix, where here K is the number of individuals and p is the number of SNPs on the chromosome of interest. The singular value decomposition allows us to write X = UΛV T where Λ is a K × K diagonal matrix, U is a K × K matrix such that UU T = U TU = I K, V is a p × K matrix such that VV T = I p and V TV = I K, with I K and I p denoting the K and p dimensional identity matrices. Pittelkow and Wilson (2003) considers the following variant of the biplot that they call the GE-biplot. Write \(C=\sqrt{K}U\) and \(G=V\Lambda/\sqrt{K}\) , so that X = CG T. To understand the application of this decomposition, following Jolliffe (1986), let c Ti, i = 1, ..., K and g Tj, j = 1, ..., p denote the rows of C and G, respectively. These may be thought of as pseudo individuals and pseudo SNPs, respectively. Then the (i,j)th element of X may be written as xij = c Tig j. Let c *i and g *j denote the vectors that contain the first two elements of c i and g j, respectively. Then we approximate the (i,j)th element of X by \(\tilde x_{ij}={{\mathbf {c}}}^{*T}_i{{\mathbf {g}}}^*_j\). Considered separately, c *i and g *j provide information on the individuals and the SNPs, as observed by previous authors. However, their importance to us comes from the relationship x ij = cTig j so that x ij is the inner product of c i and g j. Thus, x ij is close to zero if c i and g j are close to orthogonal, and if x ij is distant from zero then c i and g j must lie in a similar direction. Thus, the relative positions of the approximations c*i and g*j give us information on the size of the observations xij. This is best examined in a biplot where the c*i and g*j are plotted on the same axes. This is illustrated in our application below.
Results
We illustrate our method on a set of 14 leukaemia patients (labelled P1, P2, P3, P4, P5, P6, P7, P8, P9, P10, P11, P12, P13, and P14) from the previous leukaemia studies (Batova et al. 1997, 1999; Diccianni et al. 1997; Omura-Minamisawa et al. 2000) and 95 normal controls from the previous project (Pan et al. 2006). All individuals whose samples were used in this study signed informed consent forms. Leukocyte genomic DNA of all samples was prepared and then genotyped using the Affymetrix Human Mapping 100K Set (Affymetrix, CA, USA), which provides 116,204 SNPs with a median inter-marker distance of 8.5 kb across the human genome for each individual. Genotype data of all SNPs were obtained by using the dynamic model-based algorithm (Di et al. 2005) available at the software GDAS (Affymetrix, CA, USA). For each SNP, the genotype is AA, AB, BB, or NoCall. We aimed to identify chromosomal regions with a higher probability of homozygous calls (AA or BB) in the leukaemia patients compared to the normal controls.
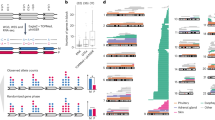
We used the nearest 2.5% of the SNPs on the chromosome to estimate the homozygous intensity. The smallest P value from the Wilcoxon rank sum test was 0.005 for chromosome 9, and we concentrate on this chromosome. The biplot of Fig. 2 identifies P2, P3, P4, P6, P9, and P13 as unusual. Further examination of the plotted intensities revealed differences in the regions ≈0−40 Mb in the p-arm of chromosome 9 and ≈68−100 Mb in the q-arm of chromosome 9 as plotted in Figs. 3 and 4. In these figures, the 90th quantile of the normal controls is plotted as a bold line. In both plots there are several cases with intensity values regularly above the 90th quantile, with P4, P6, P9, and P13 tending to be above the quantile in the ≈0−40 Mb range and P2 and P3 in the ≈68−100 Mb range. The analysis in this paper identifies the chromosomal region with LOH, including the gene locus of cyclin-dependent kinase inhibitor 2A (CDKN2A) on 9p21, whose deletion has been validated by quantitative polymerase chain reaction (qPCR) in patients P4, P6, P9, and P13 (Li et al. in preparation). CDKN2A, encoding p16INK4A and p14ARF proteins, is an important tumour suppressor gene located on 9p21 (Krimpenfort et al. 2001). The mutation of CDKN2A has been proved to involve in the tumourigenesis of leukaemia (Rasool et al. 1995).
Biplot for chromosome 9 of all samples. Each sample is represented by a red line and arranged in a new coordinate system in the biplot. For each normal control, the red line is labelled by a star sign; for each patient, the red line is labelled by the patient ID. Red lines distant to the majority of samples are regarded as potential subjects with LOH on some specific chromosome regions
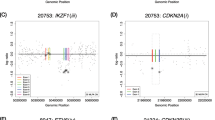
Plot of estimated homozygous intensity from 0 to 40 Mb on chromosome 9 of six leukaemia patients (P2, P3, P4, P6, P9, and P13). Estimated homozygous intensity, with bandwidth 2.5%, against the SNP position (unit: Mb) for the six cases identified in Fig. 2 is shown. Each line denotes the estimated homozygous intensity for a patient. The solid red line is the 90th quantile of the normal intensities
Plot of estimated homozygous intensity from ≈68 to 100 Mb on chromosome 9 of six leukaemia patients (P2, P3, P4, P6, P9, and P13). Estimated homozygous intensity, with bandwidth 2.5%, against the SNP position (unit: Mb) for the six cases identified in Fig. 2 is shown. Each line denotes the estimated homozygous intensity for a patient. The solid red line is the 90th quantile of the normal intensities
The proposed procedure can also be used to identify LOH caused by other mechanisms than gene deletion. For example, the long stretch of LOH identified in P13 is mainly caused by copy-neutral LOH rather than deletion. In fact, deletion-induced LOH is restricted to CDKN2A and CDKN2B (physical position ≈21.9−22.0 Mb) in this patient (data not shown). The results of our analysis justify that the proposed method is a convenient and reliable tool for a genome-wide LOH detection. Interestingly, the biplot of Fig. 2 also identifies one quite unusual control. In this normal individual, a long, contiguous stretch of homozygosity (LCSH) on chromosome 9p without copy number loss is observed. LCSH may occur in the genomes of normal individuals and most likely reflects the phenomenon of autozygosity (Li et al. 2006).
Simulation
We evaluated the performance of the proposed procedure by examining statistical power and type 1 error in simulation studies. The parameter settings in simulation study were chosen according to the real scenario of chromosome 9 discussed in Sect. 3. On chromosome 9, there were 4,796 SNPs designed in the gene chip of the Affymetrix Human Mapping 100K Set (Affymetrix, CA, USA), and the median inter-marker distance was 8.3 kb. Therefore, we generated data of 4,796 SNPs (N = 4,796) for 95 normal controls (n = 95) and 14 patients (m = 14) in the simulation study. Three parameters considered in the simulation study were (1) the percentage of SNPs that were close to the study loci and used to estimate intensity of homozygosity (α%); (2) the number of SNPs occurred in the real LOH region (n LOH); (3) mean intensity differences between case and control groups in the real LOH region (δ). Under each of simulation conditions, 500 simulation replications were performed. The simulation conditions were considered as follows:
First, in general, a conventional karyotyping has a 4-Mb resolution limitation, and more advanced platforms have a 1-Mb resolution limitation. Therefore, three lengths of real LOH regions, 1 Mb (high resolution), 2 Mb (intermediate resolution), and 4 Mb (low resolution), were considered. Out of all SNPs on chromosome 9, we selected α% = 2.5, 5, and 10% of SNPs close to the study loci. The three conditions corresponded to ≈120, 240, and 480 SNPs, respectively. The spanned lengths of corresponding regions were ≈1, 2, and 4 Mb, respectively. Second, we considered that n LOH = 120, 240, and 480 SNPs occurred in the real LOH region. Third, the biplot of Fig. 2 identified six patients (P2, P3, P4, P6, P9, and P13) and one control as unusual, where P4, P6, P9, P13, and the control had aberrant regions in the p-arm of chromosome 9. We considered δ = 4/14−1/95 ≈ 0.3. In addition, conditions of a small effect size (δ = 0.15) and a large effect size (δ = 0.6) were considered. Under a test size of S, test power was calculated for the scenario that LOH occurred on the study chromosome (δ > 0 and n LOH > 0); type 1 error was calculated for the scenario that the entire chromosomal region was free of LOH (δ = 0 and n LOH = 0). We examined and discussed the impacts of the three aforementioned parameters on power and type 1 error of the proposed test.
Results of simulation studies are shown in Table 1. The results showed that power of the proposed test varied with the magnitude of effect size (δ). The larger the effect size, the higher the power. Under test size of S = 0.050, the average power for conditions δ = 0.15, δ = 0.30, and δ = 0.60, was 0.351, 0.728, and 1.000, respectively; under test size of S = 0.025, the average power for the conditions δ = 0.15, δ = 0.30, and δ = 0.60 was 0.224, 0.601, and 1.000, respectively. Type 1 error of the proposed test was slightly inflated, probably due to multiple tests. Under test size of S = 0.050 and S = 0.025, the type 1 error was 0.085 and 0.047, respectively. Regarding the impacts of n LOH and α on power and type 1 error, we found that changes of the two parameters did not remarkably affect the proposed test under the scenario of chromosome 9 in our leukaemia study.
Discussion
Loss of heterozygosity detection plays an important role in cancer research. Identification of LOH regions across the human genome is very challenging due to the huge amount of genomic data and complex mechanism of cancer. A two-stage procedure, which consists of a genome-wide screen in the first stage and a biological confirmation in the second stage, is an efficient strategy for this work. This paper aimed to provide a convenient analysis procedure for genome-wide LOH detection based on SNP chip data. We found in Fig. 1 that raw data on homozygosity for a single individual were difficult to interpret. We used standard nonparametric procedures to estimate the homozygous intensity at each SNP for each individual. This allowed a graphical representation of homozygosity for each individual. A statistic based on the comparison of the differences of the estimated homozygosity functions of normal controls and cases from a nominal upper bound of homozygosity function in pooled samples was then computed to order the priority of chromosomes for further examination. Candidate chromosomes may be examined using a biplot. This aids in the detection of LOH regions and helps determine which cases were influenced by LOH in which region of which chromosome.
Some concluding remarks are summarised. Firstly, our method is reliable. The performance has been evaluated by simulation studies. Secondly, the analysis is biologically meaningful. The LOH regions identified by our method in the ALL study have been confirmed by qPCR experiments and are highly related to tumourigenesis. Thirdly, the method is convenient. The procedure can be implemented using standard statistical packages. Fourthly, the strategy is feasible and cost-saving. The proposed genome-wide LOH screen provides a systematical approach to scan human genome. Only the identified LOH regions require the next-stage biological examinations. Therefore, it helps reduce the effort and cost of expensive qPCR experiments.
In discussion, a small inflation of type 1 error was found in our simulation study. Combining the use of our procedure and a multiple testing correction, such as Holm’s correction (Holm 1979) and false discovery rate (Benjamini and Hochberg 1995), is suggested. In addition, an important parameter, the percentage of SNPs in a moving window, is involved in the estimation of intensity. The effect may be small in some situations, like the scenario in our leukaemia study (see Sect. 4). However, it may become critical if the LOH regions are short. In this situation, use of an over-large smoothing constant may increase an estimation bias, which leads to failure of detecting small LOH regions; use of a too small smoothing constant may increase estimation variability, which results in the false alarm of LOH regions. Currently, we are studying an optimal choice of this parameter.
References
Batova A, Diccianni MB, Omura-Minamisawa M, Yu J, Carrera CJ, Bridgeman LJ, Kung FH, Pullen J, Amylon MD, Yu AL (1999) Use of alanosine as a methylthioadenosine phosphorylase-selective therapy for T-cell acute lymphoblastic leukemia in vitro. Cancer Res 59:1492–1497
Batova A, Diccianni MB, Yu JC, Nobori T, Link MP, Pullen J, Yu AL (1997) Frequent and selective methylation of p15 and deletion of both p15 and p16 in T-cell acute lymphoblastic leukemia. Cancer Res 57:832–836
Benjamini Y, Hochberg Y (1995) Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc B 57:289–300
Di X, Matsuzaki H, Webster TA, Hubbell E, Liu G, Dong S, Bartell D, Huang J, Chiles R, Yang G, Shen MM, Kulp D, Kennedy GC, Mei R, Jones KW, Cawley S (2005) Dynamic model based algorithms for screening and genotyping over 100K SNPs on oligonucleotide microarrays. Bioinformatics 21:1958–1963
Diccianni MB, Batova A, Yu J, Vu T, Pullen J, Amylon M, Pollock B, Yu AL (1997) Shortened survival after relapse in T-cell acute lymphoblastic leukemia patients with p16/p15 deletions. Leukemia Res 21:549–558
Gabriel KR (1971) The biplot graphical display of matrices with applications to principal components analysis. Biometrika 58:453–467
Holm S (1979) A simple sequentially rejective multiple test procedure. Scand J Stat 6:65–70
Huang J, Wei W, Zhang J, Liu G, Bignell GR, Stratton MR, Futreal PA, Wooster R, Jones KW, Shapero MH (2004) Whole genome DNA copy number changes identified by high density oligonucleotide arrays. Hum Genomics 1:287–299
Jolliffe IT (1986) Principal component analysis. Springer, New York
Knudson AG (1985) Hereditary cancer, oncogenes, and anti-oncogenes. Cancer Res 45:1437–1443
Krimpenfort P, Quon KC, Mooi WJ, Loonstra A, Berns A (2001) Loss of p16Ink4a confers susceptibility to metastatic melanoma in mice. Nature 413:83–86
Li LH, Ho SF, Chen CH, Wei CY, Wong WC, Li LY, Hung SI, Chung WH, Pan WH, Lee MTM, Tsai FJ, Chang CF, We JY, Chen YT (2006) Long contiguous stretches of homozygosity in the human genome. Hum Mutat 27:1115–1121
Lin M, Wei LJ, Sellers WR, Lieberfarb M, Wong WH, Li C (2004) dChipSNP: significance curve and clustering of SNP-array-based loss-of-heterozygosity data. Bioinformatics 20:1233–1240
Loader C (1999) Local regression and likelihood. Springer, New York
Matsuzaki H, Dong S, Loi H, Di X, Liu G, Hubbell E, Law J, Berntsen T, Chadha M, Hui H, Yang G, Kennedy GC, Webster TA, Cawley S, Walsh PS, Jones KW, Fodor SPA, Mei R (2004a) Genotyping over 100,000 SNPs on a pair of oligonucleotide arrays. Nat Methods 1:109–111
Matsuzaki H, Loi H, Dong S, Tsai YY, Fang J, Law J, Di X, Liu WM, Yang G, Liu G, Huang J, Kennedy GC, Ryder TB, Marcus GA, Walsh PS, Shriver MD, Puck JM, Jones KW, Mei R (2004b) Parallel genotyping of over 10,000 SNPs using a one-primer assay on a high-density oligonucleotide array. Genome Res 14:414–425
Omura-Minamisawa M, Diccianni MB, Batova A, Chang RC, Bridgeman LJ, Yu J, Pullen J, Bowman WP, Yu AL (2000) Universal inactivation of both p16 and p15 but not downstream components is an essential event in the pathogenesis of T-cell acute lymphoblastic leukemia. Clin Cancer Res 6:1219–1228
Pan WH, Fann CSJ, Wu JY, Hung YT, Ho MS, Tai TH, Chen YJ, Liao CJ, Yang ML, Cheng ATA, Chen YT (2006) Han Chinese cell and genome bank in Taiwan: purpose, design and ethical considerations. Hum Hered 61:27–30
Pittelkow YE, Wilson SR (2003) Visualisation of gene expression data—the GE-biplot, the chip-plot and the gene-plot. Stat Appl Genet Mol Biol 2:6
Ramsay JO, Silverman BW (1997) Functional data analysis. Springer, New York
Rasool O, Heyman M, Brandter LB, Liu Y, Grander D, Soderhall S, Einhorn S (1995) p15ink4B and p16ink4 gene inactivation in acute lymphocytic leukemia. Blood 85:3431–3436
Rubocki RJ, Duffy KJ, Shepard KL, McCue BJ, Shepard SJ, Wisecarver JL (2000) Loss of heterozygosity detected in a short tandem repeat (STR) locus commonly used for human DNA identification. J Forensic Sci 45:1087–1089
Acknowledgments
We appreciate two anonymous reviewers for their insightful suggestions and comments, which have improved this paper. This work commenced when the first author visited the Institute of Statistical Science, Academia Sinica, Taipei, Taiwan. The authors are grateful to the National Science Council of Taiwan for funding that made the visit possible. We also would like to acknowledge the National Genotyping Centre of National Research Programme for Genomic Medicine, NSC, for genotyping support. This work was partially supported by a National Science Council of Taiwan grant (NSC 97-2314-B-001-006-MY3) and a National Research Program of Taiwan for Genomic Medicine grant (NSC 97-3112-B-001-027).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Huggins, R., Li, LH., Lin, YC. et al. Nonparametric estimation of LOH using Affymetrix SNP genotyping arrays for unpaired samples. J Hum Genet 53, 983–990 (2008). https://doi.org/10.1007/s10038-008-0340-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10038-008-0340-9
Keywords
This article is cited by
-
Homozygosity disequilibrium and its gene regulation
BMC Proceedings (2016)
-
SAQC: SNP Array Quality Control
BMC Bioinformatics (2011)