Abstract
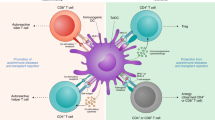
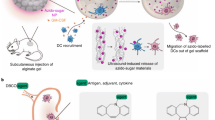
Using siRNAs to genetically manipulate immune cells is important to both basic immunological studies and therapeutic applications. However, siRNA delivery is challenging because primary immune cells are often sensitive to the delivery materials and generate immune responses. We have recently developed an amphiphilic dendrimer that is able to deliver siRNA to a variety of cells, including primary immune cells. We provide here a protocol for the synthesis of this dendrimer, as well as siRNA delivery to immune cells such as primary T and B cells, natural killer cells, macrophages, and primary microglia. The dendrimer synthesis entails straightforward click coupling followed by an amidation reaction, and the siRNA delivery protocol requires simple mixing of the siRNA and dendrimer in buffer, with subsequent application to the primary immune cells to achieve effective and functional siRNA delivery. This dendrimer-mediated siRNA delivery largely outperforms the standard electroporation technique, opening a new avenue for functional and therapeutic studies of the immune system. The whole protocol encompasses the dendrimer synthesis, which takes 10 days; the primary immune cell preparation, which takes 3–10 d, depending on the tissue source and cell type; the dendrimer-mediated siRNA delivery; and subsequent functional assays, which take an additional 3–6 d.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Wilson, R. C. & Doudna, J. A. Molecular mechanisms of RNA interference. Annu. Rev. Biophys. 42, 217–239 (2013).
Kim, D. & Rossi, J. RNAi mechanisms and applications. Biotechniques 44, 613–616 (2008).
Setten, R. L., Rossi, J. J. & Han, S. P. The current state and future directions of RNAi-based therapeutics. Nat. Rev. Drug Discov. 18, 421–446 (2019).
Sioud, M. Releasing the immune system brakes using siRNAs enhances cancer immunotherapy. Cancers (Basel) 11, 176 (2019).
Riley, R. S., June, C. H., Langer, R. & Mitchell, M. J. Delivery technologies for cancer immunotherapy. Nat. Rev. Drug Discov. 18, 175–196 (2019).
Kanasty, R., Dorkin, J. R., Vegas, A. & Anderson, D. Delivery materials for siRNA therapeutics. Nat. Mater. 12, 967–977 (2013).
Whitehead, K. A., Langer, R. & Anderson, D. G. Knocking down barriers: advances in siRNA delivery. Nat. Rev. Drug Discov. 8, 129–138 (2009).
Judge, A. D. et al. Sequence-dependent stimulation of the mammalian innate immune response by synthetic siRNA. Nat. Biotechnol. 23, 457–462 (2005).
Judge, A. & MacLachlan, I. Overcoming the innate immune response to small interfering RNA. Hum. Gene Ther. 19, 111–124 (2008).
Whitehead, K. A., Dahlman, J. E., Langer, R. S. & Anderson, D. G. Silencing or stimulation? siRNA delivery and the immune system. Annu. Rev. Chem. Biomol. Eng. 2, 77–96 (2011).
Zhang, X., Edwards, J. P. & Mosser, D. M. The expression of exogenous genes in macrophages: obstacles and opportunities. Methods Mol. Biol. 531, 123–143 (2009).
Freeley, M. & Long, A. Advances in siRNA delivery to T-cells: potential clinical applications for inflammatory disease, cancer and infection. Biochem. J. 455, 133–147 (2013).
Nakamura, T. et al. Small-sized, stable lipid nanoparticle for the efficient delivery of siRNA to human immune cell lines. Sci. Rep. 6, 37849 (2016).
Nakamura, T., Yamada, K., Fujiwara, Y., Sato, Y. & Harashima, H. Reducing the cytotoxicity of lipid nanoparticles associated with a fusogenic cationic lipid in a natural killer cell line by introducing a polycation-based siRNA core. Mol. Pharmaceutics 15, 2142–2150 (2018).
Davis, M. E. et al. Evidence of RNAi in humans from systemically administered siRNA via targeted nanoparticles. Nature 464, 1067–1070 (2010).
Yu, T. et al. An amphiphilic dendrimer for effective delivery of small interfering RNA and gene silencing in vitro and in vivo. Angew. Chem. Int. Ed. Engl. 51, 8478–8484 (2012).
Liu, X. X. et al. Adaptive amphiphilic dendrimer-based nanoassemblies as robust and versatile siRNA delivery systems. Angew. Chem. Int. Ed. Engl 53, 11822–11827 (2014).
Chen, C. et al. Mastering dendrimer self-assembly for efficient siRNA delivery: from conceptual design to in vivo efficient gene silencing. Small 12, 3667–3676 (2016).
Dong, Y. et al. A dual targeting dendrimer-mediated siRNA Delivery system for effective gene silencing in cancer therapy. J. Am. Chem. Soc. 140, 16264–16274 (2018).
Ellert-Miklaszewska, A. et al. Efficient and innocuous delivery of small interfering RNA to microglia using an amphiphilic dendrimer nanovector. Nanomedicine (Lond.) 14, 2441–2458 (2019).
Garofalo, S. et al. Natural killer cells modulate motor neuron-immune cell cross talk in models of amyotrophic lateral sclerosis. Nat. Commun. 11, 1773 (2020).
Kim, D. H. et al. Synthetic dsRNA Dicer substrates enhance RNAi potency and efficacy. Nat. Biotechnol. 23, 222–226 (2005).
Amarzguioui, M. et al. Rational design and in vitro and in vivo delivery of Dicer substrate siRNA. Nat. Protoc. 1, 508–517 (2006).
Tomalia, D. A. et al. A new class of polymers - starburst-dendritic macromolecules. Polym. J. 17, 117–132 (1985).
Peterson, J., Allikmaa, V., Subbi, J., Pehk, T. & Lopp, M. Structural deviations in poly(amidoamine) dendrimers: a MALDI-TOF MS analysis. Eur. Polym. J. 39, 33–42 (2003).
van Dongen, M. A., Desai, A., Orr, B. G., Baker, J. R. & Holl, M. M. B. Quantitative analysis of generation and branch defects in G5 poly(amidoamine) dendrimer. Polymer 54, 4126–4133 (2013).
Rastogi, A. & Nayan, R. Studies on copper (II) complexes of some polyaza macrocycles derived from 1, 2-diaminoethane. J. Coord. Chem. 62, 3366–3376 (2009).
Trouplin, V. et al. Bone marrow-derived macrophage production. J. Vis. Exp 2013, e50966 (2013).
Armarego, W. L. F. & Perrin, D. D. Purification of Laboratory Chemicals (Butterworth Heinemann, 1996).
Vogel, A. I., Furniss, B. S., Hannaford, A. J., Smith, P. W. & Tatchell, A. R. Vogel’s Textbook of Practical Organic Chemistry (Longman Scientific & Technical, 1989).
Fauci, A. S. The human immunodeficiency virus: infectivity and mechanisms of pathogenesis. Science 239, 617–622 (1988).
Frankel, A. D. & Young, J. A. HIV-1: fifteen proteins and an RNA. Annu. Rev. Biochem. 67, 1–25 (1998).
Molfetta, R., Quatrini, L., Santoni, A. & Paolini, R. Regulation of NKG2D-dependent NK cell functions: the yin and the yang of receptor endocytosis. Int. J. Mol. Sci. 18, 1677 (2017).
Malemud, C. J. The role of the JAK/STAT signal pathway in rheumatoid arthritis. Ther. Adv. Musculoskelet. Dis. 10, 117–127 (2018).
De Vries, L. C. S., Wildenberg, M. E., De Jonge, W. J. & D’Haens, G. R. The future of Janus kinase inhibitors in inflammatory bowel disease. J. Crohns Colitis 11, 885–893 (2017).
Triyangkulsri, K. & Suchonwanit, P. Role of Janus kinase inhibitors in the treatment of alopecia areata. Drug Des. Devel. Ther. 12, 2323–2335 (2018).
Guo, X. et al. Transfection reagent Lipofectamine triggers type I interferon signaling activation in macrophages. Immunol. Cell Biol. 97, 92–96 (2019).
Papaspyridonos, M. et al. Id1 suppresses anti-tumour immune responses and promotes tumour progression by impairing myeloid cell maturation. Nat. Commun. 6, 6840 (2015).
Zebedee, Z. & Hara, E. Id proteins in cell cycle control and cellular senescence. Oncogene 20, 8317–8325 (2001).
Acknowledgements
This work was supported by La Ligue Nationale Contre le Cancer (EL2016.LNCC/LPP to L.P.); the NIH (grants R01AI29329, R01AI42552, and R01HL07470 to J.J.R., as well as grant P30CA033572 for City of Hope Core Facility support); National Science Centre Poland (no. 2017/25/B/NZ3/02483 to A.E.-M.); the French National Research Agency and the Italian Ministry of Health under the frame of the Era-Net EURONANOMED European Research projects ‘NANOGLIO’ (L.P., A.S., C.L., P.N.M.), ‘TARBRAINFECT’ (L.P.) and ‘NAN-4-TUM’ (L.P.); the China Scholarship Council (J.C., J.T.) and Bourse Eiffel du Campus France (J.C.); the Italian Association for Cancer Research (AIRC) (IG 2015 and IG 2019 to C.L; 22329 2018 to S.G.); and the Italian Research Foundation for ALS (AriSLA) (Pilot NKINALS 2019 to S.G.). This project has received funding from the European Union’s Horizon 2020 research and innovation program H2020 NMBP ‘SAFE-N-MEDTECH’ (L.P.; under grant agreement no. 814607) and ‘NEWDEAL’ (A.D., F.C., P.N.M.; under grant agreement no. 720905). This publication reflects only the authors’ views, and the Commission is not responsible for any use that may be made of the information it contains. This article is based upon work from COST Action CA 17140 ‘Cancer Nanomedicine from the Bench to the Bedside’, supported by COST (European Cooperation in Science and Technology). We thank A. Tintaru from Aix-Marseille University in France for NMR recording, G. Bernardini from the University of Rome La Sapienza in Italy for FACS analysis, and E. Sulpice and X. Gidrol from CEA/IRIG in Grenoble France for generously providing mouse JAK1 siRNA.
Author information
Authors and Affiliations
Contributions
Conceptualization, J.Z., L.P.; writing and editing, J.Z., L.P., J.C., Y.J., J.T., A.E.-M., B.K., C.L., S.G., A.S., A.K.D., F.C., P.N.M.; funding acquisition, L.P., J.J.R., B.K., A.E.-M., C.L., P.N.M.; all authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Protocols thanks Jørn B. Christensen and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Related links
Key references using this protocol
Liu, X. et al. Angew. Chem. Int. Ed. 53, 11822–11827 (2014): https://doi.org/10.1002/anie.201406764
Ellert-Miklaszewska, A. et al. Nanomedicine (Lond.) 14, 2441–2458 (2019): https://doi.org/10.2217/nnm-2019-0176
Garofalo, S. et al. Nat. Commun. 11, 1773 (2020): https://doi.org/10.1038/s41467-020-15644-8
Supplementary information
Supplementary Information
Supplementary Figs. 1–8, Supplementary Results, Supplementary Methods and Supplementary References.
Source data
Source Data Fig. 4
Statistical source data for Fig. 4a,b,c, d, f, g.
Source Data Fig. 4
Unprocessed western blots (gels) for Figs. 4e and 4h.
Rights and permissions
About this article
Cite this article
Chen, J., Ellert-Miklaszewska, A., Garofalo, S. et al. Synthesis and use of an amphiphilic dendrimer for siRNA delivery into primary immune cells. Nat Protoc 16, 327–351 (2021). https://doi.org/10.1038/s41596-020-00418-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41596-020-00418-9
This article is cited by
-
Engineering siRNA therapeutics: challenges and strategies
Journal of Nanobiotechnology (2023)
-
Synthesis of siRNA nanoparticles to silence plaque-destabilizing gene in atherosclerotic lesional macrophages
Nature Protocols (2022)
-
From Bench to the Clinic: The Path to Translation of Nanotechnology-Enabled mRNA SARS-CoV-2 Vaccines
Nano-Micro Letters (2022)
-
An ionizable supramolecular dendrimer nanosystem for effective siRNA delivery with a favorable safety profile
Nano Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.