Abstract
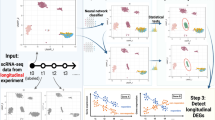
Current computational workflows for comparative analyses of single-cell datasets typically use discrete clusters as input when testing for differential abundance among experimental conditions. However, clusters do not always provide the appropriate resolution and cannot capture continuous trajectories. Here we present Milo, a scalable statistical framework that performs differential abundance testing by assigning cells to partially overlapping neighborhoods on a k-nearest neighbor graph. Using simulations and single-cell RNA sequencing (scRNA-seq) data, we show that Milo can identify perturbations that are obscured by discretizing cells into clusters, that it maintains false discovery rate control across batch effects and that it outperforms alternative differential abundance testing strategies. Milo identifies the decline of a fate-biased epithelial precursor in the aging mouse thymus and identifies perturbations to multiple lineages in human cirrhotic liver. As Milo is based on a cell–cell similarity structure, it might also be applicable to single-cell data other than scRNA-seq. Milo is provided as an open-source R software package at https://github.com/MarioniLab/miloR.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Code availability
Milo is implemented as an open-source package in R (https://github.com/MarioniLab/miloR) and is installable from Bioconductor (≥3.13; http://www.bioconductor.org/packages/release/bioc/html/miloR.html). Code used to generate figures and perform analyses can be found at https://github.com/MarioniLab/milo_analysis_2020.
References
Kiselev, V. Y., Andrews, T. S. & Hemberg, M. Challenges in unsupervised clustering of single-cell RNA-seq data. Nat. Rev. Genet. 20, 273–282 (2019).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Baran-Gale, J. et al. Ageing compromises mouse thymus function and remodels epithelial cell differentiation. eLife 9, e56221 (2020).
Pijuan-Sala, B. et al. A single-cell molecular map of mouse gastrulation and early organogenesis. Nature 566, 490–495 (2019).
Haber, A. L. et al. A single-cell survey of the small intestinal epithelium. Nature 551, 333–339 (2017).
Lun, A. T. L., Richard, A. C. & Marioni, J. C. Testing for differential abundance in mass cytometry data. Nat. Methods 14, 707–709 (2017).
Zhao, J. et al. Detection of differentially abundant cell subpopulations discriminates biological states in scRNA-seq data. Proc. Natl Acad. Sci. USA 118, e2100293118 (2021).
Burkhardt, D. B. et al. Quantifying the effect of experimental perturbations at single-cell resolution. Nat. Biotechnol. 39, 619–629 (2021).
Gut, G., Tadmor, M. D., Pe’er, D., Pelkmans, L. & Liberali, P. Trajectories of cell-cycle progression from fixed cell populations. Nat. Methods 12, 951–954 (2015).
McCarthy, D. J., Chen, Y. & Smyth, G. K. Differential expression analysis of multifactor RNA-seq experiments with respect to biological variation. Nucleic Acids Res. 40, 4288–4297 (2012).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Benjamini, Y. & Hochberg, Y. Multiple hypotheses testing with weights. Scand. J. Statist. 24, 407–418 (1997).
Soneson, C. & Robinson, M. D. Bias, robustness and scalability in single-cell differential expression analysis. Nat. Methods 15, 255–261 (2018).
Cannoodt, R., Saelens, W., Deconinck, L. & Saeys, Y. Spearheading future omics analyses using dyngen, a multi-modal simulator of single cells. Nat. Communications 12, 1–9 (2021).
Luecken, M. et al. Benchmarking atlas-level data integration in single-cell genomics. Preprint at https://www.biorxiv.org/content/10.1101/2020.05.22.111161v2 (2020).
Tran, H. T. N. et al. A benchmark of batch-effect correction methods for single-cell RNA sequencing data. Genome Biol. 21, 12 (2020).
Chazarra-Gil, R., van Dongen, S., Kiselev, V. Y. & Hemberg, M. Flexible comparison of batch correction methods for single-cell RNA-seq using BatchBench. Nucleic Acids Res. 49, e42 (2021).
Haghverdi, L., Lun, A. T. L., Morgan, M. D. & Marioni, J. C. Batch effects in single-cell RNA-sequencing data are corrected by matching mutual nearest neighbors. Nat. Biotechnol. 36, 421–427 (2018).
Stoeckius, M. et al. Cell Hashing with barcoded antibodies enables multiplexing and doublet detection for single cell genomics. Genome Biol. 19, 224 (2018).
McGinnis, C. S. et al. MULTI-seq: sample multiplexing for single-cell RNA sequencing using lipid-tagged indices. Nat. Methods 16, 619–626 (2019).
Akiyama, T. et al. The tumor necrosis factor family receptors RANK and CD40 cooperatively establish the thymic medullary microenvironment and self-tolerance. Immunity 29, 423–437 (2008).
Hikosaka, Y. et al. The cytokine RANKL produced by positively selected thymocytes fosters medullary thymic epithelial cells that express autoimmune regulator. Immunity 29, 438–450 (2008).
Wilkinson, A. L., Qurashi, M. & Shetty, S. The role of sinusoidal endothelial cells in the axis of inflammation and cancer within the liver. Front. Physiol. 11, 990 (2020).
Foldi, I. et al. Lectin-complement pathway molecules are decreased in patients with cirrhosis and constitute the risk of bacterial infections. Liver Int. 37, 1023–1031 (2017).
Ganesan, L. P. et al. FcγRIIb on liver sinusoidal endothelium clears small immune complexes. J. Immunol. 189, 4981–4988 (2012).
Sato, K. et al. Ductular reaction in liver diseases: pathological mechanisms and translational significances: liver injury and regeneration. Hepatology 69, 420–430 (2019).
Morell, C. M., Fabris, L. & Strazzabosco, M. Vascular biology of the biliary epithelium: biliary epithelium vascular biology. J. Gastroenterol. Hepatol. 28, 26–32 (2013).
Mariotti, V., Fiorotto, R., Cadamuro, M., Fabris, L. & Strazzabosco, M. New insights on the role of vascular endothelial growth factor in biliary pathophysiology. JHEP Rep. 3, 100251 (2021).
R Core Team. R: A Language and Environment for Statistical Computing. https://www.R-project.org (R Foundation for Statistical Computing, 2017).
Büttner, M., Ostner, J., Müller, C. l., Theis, F. J. & Schubert, B. scCODA: a Bayesian model for compositional single-cell data analysis. Preprint at https://www.biorxiv.org/content/10.1101/2020.12.14.422688v2 (2020).
Dixit, A. et al. Perturb-seq: dissecting molecular circuits with scalable single-cell RNA profiling of pooled genetic screens. Cell 167, 1853–1866 (2016).
Datlinger, P. et al. Pooled CRISPR screening with single-cell transcriptome readout. Nat. Methods 14, 297–301 (2017).
Jaitin, D. A. et al. Dissecting immune circuits by linking CRISPR-pooled screens with single-cell RNA-seq. Cell 167, 1883–1896 (2016).
Stoeckius, M. et al. Simultaneous epitope and transcriptome measurement in single cells. Nat. Methods 14, 865–868 (2017).
Cao, J. et al. Joint profiling of chromatin accessibility and gene expression in thousands of single cells. Science 361, 1380–1385 (2018).
Chen, S., Lake, B. B. & Zhang, K. High-throughput sequencing of the transcriptome and chromatin accessibility in the same cell. Nat. Biotechnol. 37, 1452–1457 (2019).
Zhu, C. et al. An ultra high-throughput method for single-cell joint analysis of open chromatin and transcriptome. Nat. Struct. Mol. Biol. 26, 1063–1070 (2019).
Ma, S. et al. Chromatin potential identified by shared single-cell profiling of RNA and chromatin. Cell 183, 1103–1116 (2020).
Luecken, M. D. & Theis, F. J. Current best practices in single-cell RNA-seq analysis: a tutorial. Mol. Syst. Biol. 15, e8746 (2019).
Setty, M. et al. Wishbone identifies bifurcating developmental trajectories from single-cell data. Nat. Biotechnol. 34, 637–645 (2016).
Griffiths, J. & Lun, A. MouseGastrulationData: single-cell transcriptomics data across mouse gastrulation and early organogenesis. https://github.com/MarioniLab/MouseGastrulationData (2021).
Csardi, G. & Nepusz, T. The igraph software package for complex network research. InterJournal http://www.interjournal.org/manuscript_abstract.php?361100992 (2006).
Huber, W. et al. Orchestrating high-throughput genomic analysis with Bioconductor. Nat. Methods 12, 115–121 (2015).
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004).
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Kuleshov, M. V. et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 44, W90–W97 (2016).
Lun, A. T. L., McCarthy, D. J. & Marioni, J. C. A step-by-step workflow for low-level analysis of single-cell RNA-seq data with Bioconductor. F1000Res. 5, 2122 (2016).
Yu, G., Wang, L.-G., Han, Y. & He, Q.-Y. clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16, 284–287 (2012).
Acknowledgements
We thank S. Ghazanfar for feedback on the method; N. Kumasaka for comments on the manuscript; C. Suo, V. Kedlian, R. Elmentaite, J. P. Pett, K. Tuong and B. Stewart for feedback on the software package; and D. Burkhardt, M. Luecken and W. Lewis for discussions on benchmarking. J.C.M. acknowledges core funding from the European Molecular Biology Laboratory and core funding from Cancer Research UK (C9545/A29580), which supports M.D.M. E.D. and S.A.T. acknowledge Wellcome Sanger core funding (WT206194). N.C.H. is supported by a Wellcome Trust Senior Research Fellowship in Clinical Science (ref. 219542/Z/19/Z), the Medical Research Council and a Chan Zuckerberg Initiative Seed Network Grant.
Author information
Authors and Affiliations
Contributions
E.D., M.D.M. and J.C.M. conceived the method idea. E.D. and M.D.M. developed the method, wrote the code and performed analyses. E.D., M.D.M., S.A.T. and N.C.H. interpreted the results. E.D., M.D.M., S.A.T., N.C.H. and J.C.M. wrote and approved the manuscript. M.D.M. and J.C.M. oversaw the project.
Corresponding authors
Ethics declarations
Competing interests
In the last three years, S.A.T. has received remuneration for consulting and Scientific Advisory Board membership from Genentech, Roche, Biogen, ForesiteLabs and Qiagen. All other authors have no competing interests to declare.
Additional information
Peer review information Nature Biotechnology thanks Dana Pe’er, Michael Love and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Benchmarking DA methods on simulated data.
DA analysis performance on KNN graphs from simulated datasets of different topologies: (a) discrete clusters (2700 cells, 3 populations); (b) 1-D linear trajectory (7500 cells, 7 populations); (c) Branching trajectory (7500 cells, 10 populations). Boxplots show the median with interquartile ranges (25–75%); whiskers extend to the largest value no further than 1.5x the interquartile range from the distance from the box, with outlier data points shown beyond this range.
Extended Data Fig. 2 Sensitivity of DA methods to low fold change in abundance.
(a) True positive rate (TPR, top) and false positive rate (FPR, bottom) of DA methods calculated on cells in different bins of P(C1) used to generate condition labels (bin size = 0.05, the number on the x-axis indicates the lower value in the bin). The results for 36 simulations on 2 representative populations (colors) are shown. The filled points indicate the mean of each P(C1) bin. (b) Variability in Milo power is explained by the fraction of true positive cells close to the DA threshold for definition of ground truth. Example distributions of P(C1) for cells detected as true positives (TP) or false negatives (FN) by Milo. Examples for simulations on 2 populations (rows) and 3 simulated fold changes (columns) are shown. (c, d) True Positive Rate (TPR) of DA detection for simulated DA regions of increasing size centred at the same centroid (Erythroid2 (c) and Caudal neuroectoderm (d)). Results for 3 condition simulations per population and fold change are shown.
Extended Data Fig. 3 Comparison of Milo and MELD for abundance fold change estimation.
(a–d) Scatter-plots of the true fold change at the neighbourhood index against the fold change estimated by Milo (A,C) and MELD (B,D), without batch effect (a, b) and with batch effect (magnitude = 0.5) (c, d), where LFC = log(pc’/(1 - pc’)). The neighbourhoods overlapping true DA cells (pc’ greater than the 75% quantile of P(C1) in the mouse gastrulation dataset) are highlighted in red. (e, f) Mean Squared Error (MSE) comparison for MELD and Milo for true negative neighbourhood (e) and true positive neighbourhoods (f), with increasing simulated log-Fold Change and magnitude of batch effect. Each boxplot summarises the results for n=27 simulations. Box plots show the median with interquartile ranges (25–75%); whiskers extend to the largest value no further than 1.5x the interquartile range from the distance from the box, with outlier data points shown beyond this range.
Extended Data Fig. 4 Controlling for batch effects in differential abundance analysis.
(a) In silico batch correction enhances the performance of DA methods in the presence of batch effects: comparison of performance of DA methods with no batch effect, with batch effects of increasing magnitude corrected with MNN, and uncorrected batch effects. Each boxplot summarises results from simulations on n=9 populations. (b) True Positive Rate (TPR, left) and False Discovery Rate (FDR, right) for recovery of cells in simulated DA regions for DA populations with increasing batch effect magnitude on the mouse gastrulation dataset. For each boxplot, results from 8 populations and 3 condition simulations per population are shown (n=24 simulations). Each panel represents a different DA method and a different simulated log-Fold Change. (c) Comparison of Milo performance with (~ batch + condition) or without (~ condition) accounting for the simulated batch in the NB-GLM. For each boxplot, results from 8 populations, simulated fold change > 1.5 and 3 condition simulations per population and fold change are shown (72 simulations per boxplot). In all panels, boxplots show the median with interquartile ranges (25–75%); whiskers extend to the largest value no further than 1.5x the interquartile range from the distance from the box, with outlier data points shown beyond this range.
Supplementary information
Supplementary Information
Supplementary Figs. 1–7, Supplementary Tables 1–3 and Supplementary Notes
Rights and permissions
About this article
Cite this article
Dann, E., Henderson, N.C., Teichmann, S.A. et al. Differential abundance testing on single-cell data using k-nearest neighbor graphs. Nat Biotechnol 40, 245–253 (2022). https://doi.org/10.1038/s41587-021-01033-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41587-021-01033-z
This article is cited by
-
Benchmarking differential abundance methods for finding condition-specific prototypical cells in multi-sample single-cell datasets
Genome Biology (2024)
-
Single cell analysis reveals the roles and regulatory mechanisms of type-I interferons in Parkinson’s disease
Cell Communication and Signaling (2024)
-
Kernel-based testing for single-cell differential analysis
Genome Biology (2024)
-
Cellograph: a semi-supervised approach to analyzing multi-condition single-cell RNA-sequencing data using graph neural networks
BMC Bioinformatics (2024)
-
Chromatin accessibility during human first-trimester neurodevelopment
Nature (2024)