Abstract
Natural biopolymers have inspired the development of synthetic analogues that can adopt defined conformations and assemble into 3D architectures. These compounds are mainly based on peptides and nucleic acids, which are well understood at the molecular level and can be rationally synthesized to obtain functional artificial systems. By contrast, synthetic glycans have remained relatively unexplored in materials science and supramolecular chemistry, a paradox considering that carbohydrates are the most abundant structural materials on Earth. Recent advances in automated synthesis and analytical techniques have improved our understanding of glycan structures, transforming our perception of glycans from shapeless molecules to polymers adopting several distinct conformations. This newly available knowledge suggests that glycans capable of folding and assembling into defined architectures can be created. In this Perspective, we explore the emerging field of programmable carbohydrate architectures, highlighting the breakthroughs in synthesis and analysis that have made the design of artificial glycan foldamers and assemblies possible. Finally, we discuss the future prospects of this field, analysing which challenges must be overcome to establish design rules for building carbohydrate materials on demand.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
Glycan 3D features are either obtained from Polysac3-DB and BioOligo3-DB, both are sub-databases of Glyco3D (https://glyco3d.cermav.cnrs.fr), or directly from the cited publication. Peptide 3D features are obtained from the Protein Data Bank (https://www.rcsb.org). All 3D structures have been established experimentally from X-ray, neutron electron diffraction, high-resolution NMR and molecular modelling. All PDB files can also be downloaded from https://doi.org/10.17617/3.79M9Z6.
References
Hong, F., Zhang, F., Liu, Y. & Yan, H. DNA origami: scaffolds for creating higher order structures. Chem. Rev. 117, 12584–12640 (2017).
Huang, P.-S., Boyken, S. E. & Baker, D. The coming of age of de novo protein design. Nature 537, 320 (2016).
Gellman, S. H. Foldamers: a manifesto. Acc. Chem. Res. 31, 173–180 (1998).
Hill, D. J., Mio, M. J., Prince, R. B., Hughes, T. S. & Moore, J. S. A field guide to foldamers. Chem. Rev. 101, 3893–4012 (2001).
Martinek, T. A. & Fülöp, F. Peptidic foldamers: ramping up diversity. Chem. Soc. Rev. 41, 687–702 (2012).
Goodman, C. M., Choi, S., Shandler, S. & DeGrado, W. F. Foldamers as versatile frameworks for the design and evolution of function. Nat. Chem. Biol. 3, 252–262 (2007).
Girvin, Z. C., Andrews, M. K., Liu, X. & Gellman, S. H. Foldamer-templated catalysis of macrocycle formation. Science 366, 1528–1531 (2019).
Schneider, J. P. et al. Responsive hydrogels from the intramolecular folding and self-assembly of a designed peptide. J. Am. Chem. Soc. 124, 15030–15037 (2002).
Yoo, S. H. & Lee, H.-S. Foldectures: 3D molecular architectures from self-assembly of peptide foldamers. Acc. Chem. Res. 50, 832–841 (2017).
Tyrikos-Ergas, T., Fittolani, G., Seeberger, P. H. & Delbianco, M. Structural studies using unnatural oligosaccharides: toward sugar foldamers. Biomacromolecules 21, 18–29 (2020).
Diener, M. et al. Primary, secondary, tertiary and quaternary structure levels in linear polysaccharides: from random coil, to single helix to supramolecular assembly. Biomacromolecules 20, 1731–1739 (2019).
Fittolani, G., Seeberger, P. H. & Delbianco, M. Helical polysaccharides. Pept. Sci. 112, e24124 (2020).
Delbianco, M., Bharate, P., Varela-Aramburu, S. & Seeberger, P. H. Carbohydrates in supramolecular chemistry. Chem. Rev. 116, 1693–1752 (2016).
Brito, A. et al. Carbohydrate amphiphiles for supramolecular biomaterials: design, self-assembly, and applications. Chem 7, 2943–2964 (2021).
Balijepalli, A. S. & Grinstaff, M. W. Poly-amido-saccharides (PASs): functional synthetic carbohydrate polymers inspired by nature. Acc. Chem. Res. 53, 2167–2179 (2020).
Gim, S. et al. Supramolecular assembly and chirality of synthetic carbohydrate materials. Angew. Chem. Int. Ed. 59, 22577–22583 (2020).
He, C. et al. Glycopeptide self-assembly modulated by glycan stereochemistry through glycan–aromatic interactions. J. Am. Chem. Soc. 142, 17015–17023 (2020).
Pires, R. A. et al. Controlling cancer cell fate using localized biocatalytic self-assembly of an aromatic carbohydrate amphiphile. J. Am. Chem. Soc. 137, 576–579 (2015).
Liu, R. et al. A comprehensive landscape for fibril association behaviors encoded synergistically by saccharides and peptides. J. Am. Chem. Soc. 143, 6622–6633 (2021).
Guberman, M. & Seeberger, P. H. Automated glycan assembly: a perspective. J. Am. Chem. Soc. 141, 5581–5592 (2019).
Varki, A. et al. (eds) Essentials of Glycobiology 3rd edn (Cold Spring Harbor Laboratory, 2017).
Bertozzi, C. R. & Kiessling, L. L. Chemical glycobiology. Science 291, 2357–2364 (2001).
Smith, B. A. H. & Bertozzi, C. R. The clinical impact of glycobiology: targeting selectins, Siglecs and mammalian glycans. Nat. Rev. Drug Discov. 20, 217–243 (2021).
Wu, X. et al. Imaging single glycans. Nature 582, 375–378 (2020).
Gray, C. J. et al. Advancing solutions to the carbohydrate sequencing challenge. J. Am. Chem. Soc. 141, 14463–14479 (2019).
Woods, R. J. Predicting the structures of glycans, glycoproteins, and their complexes. Chem. Rev. 118, 8005–8024 (2018).
Kirschner, K. N. et al. GLYCAM06: a generalizable biomolecular force field. Carbohydrates. J. Comput. Chem. 29, 622–655 (2008).
Casalino, L. et al. Beyond shielding: the roles of glycans in the SARS-CoV-2 spike protein. ACS Cent. Sci. 6, 1722–1734 (2020).
Zhang, Q. et al. Synthetic, zwitterionic sp1 oligosaccharides adopt a helical structure crucial for antibody interaction. ACS Cent. Sci. 5, 1407–1416 (2019).
Alabugin, I. V., Kuhn, L., Krivoshchapov, N. V., Mehaffy, P. & Medvedev, M. G. Anomeric effect, hyperconjugation and electrostatics: lessons from complexity in a classic stereoelectronic phenomenon. Chem. Soc. Rev. 50, 10212–10252 (2021).
Yu, Y. & Delbianco, M. Conformational studies of oligosaccharides. Chem. Eur. J. 26, 9814–9825 (2020).
Almond, A. & Sheehan, J. K. Predicting the molecular shape of polysaccharides from dynamic interactions with water. Glycobiology 13, 255–264 (2003).
Almond, A. Towards understanding the interaction between oligosaccharides and water molecules. Carbohydr. Res. 340, 907–920 (2005).
Aeschbacher, T. et al. A secondary structural element in a wide range of fucosylated glycoepitopes. Chem. Eur. J. 23, 11598–11610 (2017).
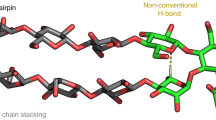
Kwon, J. et al. Glycan stability and flexibility: thermodynamic and kinetic characterization of nonconventional hydrogen bonding in Lewis antigens. J. Am. Chem. Soc. 145, 10022–10034 (2023).
Zhang, Y. et al. Synthesis and structural analysis of Aspergillus fumigatus galactosaminogalactans featuring α-galactose, α-galactosamine and α-N-acetyl galactosamine linkages. Angew. Chem. Int. Ed. 59, 12746–12750 (2020).
Seidi, F. et al. Crystalline polysaccharides: a review. Carbohydr. Polym. 275, 118624 (2022).
Pylkkänen, R. et al. β-1,3-Glucan synthesis, novel supramolecular self-assembly, characterization and application. Nanoscale 14, 15533–15541 (2022).
Panza, M., Pistorio, S. G., Stine, K. J. & Demchenko, A. V. Automated chemical oligosaccharide synthesis: novel approach to traditional challenges. Chem. Rev. 118, 8105–8150 (2018).
Fittolani, G., Tyrikos-Ergas, T., Vargová, D., Chaube, M. A. & Delbianco, M. Progress and challenges in the synthesis of sequence controlled polysaccharides. Beilstein J. Org. Chem. 17, 1981–2025 (2021).
Joseph, A. A., Pardo-Vargas, A. & Seeberger, P. H. Total synthesis of polysaccharides by automated glycan assembly. J. Am. Chem. Soc. 142, 8561–8564 (2020).
Delbianco, M. et al. Well-defined oligo- and polysaccharides as ideal probes for structural studies. J. Am. Chem. Soc. 140, 5421–5426 (2018).
Fittolani, G., Vargová, D., Seeberger, P. H., Ogawa, Y. & Delbianco, M. Bottom-up approach to understand chirality transfer across scales in cellulose assemblies. J. Am. Chem. Soc. 144, 12469–12475 (2022).
Dal Colle, M. C. S., Fittolani, G. & Delbianco, M. Synthetic approaches to break the chemical shift degeneracy of glycans. ChemBioChem 23, e202200416 (2022).
Yu, Y. et al. Systematic hydrogen-bond manipulations to establish polysaccharide structure–property correlations. Angew. Chem. Int. Ed. 58, 1433–7851 (2019).
Poveda, A., Fittolani, G., Seeberger, P. H., Delbianco, M. & Jiménez-Barbero, J. The flexibility of oligosaccharides unveiled through residual dipolar coupling analysis. Front. Mol. Biosci. 8, 784318 (2021).
Huang, J.-Y. & Delbianco, M. Recent developments in solid-phase glycan synthesis. Synthesis 55, 1337–1354 (2022).
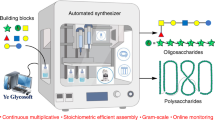
Yao, W. et al. Automated solution-phase multiplicative synthesis of complex glycans up to a 1,080-mer. Nat. Synth. 1, 854–863 (2022).
Wu, Y., Qiu, Y., Feng, Y. & Stoddart, J. F. Automating glycan assembly in solution. ACS Cent. Sci. 8, 1369–1372 (2022).
Wu, L. et al. Precision native polysaccharides from living polymerization of anhydrosugars. Nat. Chem. 15, 1276–1284 (2023).
Li, T. et al. An automated platform for the enzyme-mediated assembly of complex oligosaccharides. Nat. Chem. 11, 229–236 (2019).
Wen, L. et al. Toward automated enzymatic synthesis of oligosaccharides. Chem. Rev. 17, 8151–8187 (2018).
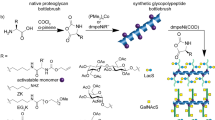
Zhang, J. et al. Machine-driven chemoenzymatic synthesis of glycopeptide. Angew. Chem. Int. Ed. 59, 19825–19829 (2020).
Smith, P. J. et al. Enzymatic synthesis of artificial polysaccharides. ACS Sustain. Chem. Eng. 8, 11853–11871 (2020).
Bell, E. L. et al. Biocatalysis. Nat. Rev. Methods Primers 1, 46 (2021).
Pallister, E., Gray, C. J. & Flitsch, S. L. Enzyme promiscuity of carbohydrate active enzymes and their applications in biocatalysis. Curr. Opin. Struct. Biol. 65, 184–192 (2020).
Gimeno, A., Valverde, P., Ardá, A. & Jiménez-Barbero, J. Glycan structures and their interactions with proteins. A NMR view. Curr. Opin. Struct. Biol. 62, 22–30 (2020).
Fontana, C. & Widmalm, G. Primary structure of glycans by NMR spectroscopy. Chem. Rev. 123, 1040–1102 (2023).
Quinn, C. M., Wang, M. & Polenova, T. in Protein NMR: Methods in Molecular Biology Vol. 1688 (ed. Ghose, R.) 1–35 (2018).
Canales, A. et al. Breaking the limits in analyzing carbohydrate recognition by NMR spectroscopy: resolving branch-selective interaction of a tetra-antennary N-glycan with lectins. Angew. Chem. Int. Ed. 56, 14987–14991 (2017).
Canales, A. et al. Breaking pseudo-symmetry in multiantennary complex N-glycans using lanthanide-binding tags and NMR pseudo-contact shifts. Angew. Chem. Int. Ed. 52, 13789–13793 (2013).
Battistel, M. D., Shangold, M., Trinh, L., Shiloach, J. & Freedberg, D. I. Evidence for helical structure in a tetramer of α2-8 sialic acid: unveiling a structural antigen. J. Am. Chem. Soc. 134, 10717–10720 (2012).
Brisson, J. R., Baumann, H., Imberty, A., Perez, S. & Jennings, H. J. Helical epitope of the group B meningococcal α(2-8)-linked sialic acid polysaccharide. Biochemistry 31, 4996–5004 (1992).
Wang, Z. et al. Synthetic zwitterionic Streptococcus pneumoniae type 1 oligosaccharides carrying labile O-acetyl esters. Angew. Chem. Int. Ed. 62, e202211940 (2023).
Li, W. et al. Synthesis of bradyrhizose oligosaccharides relevant to the Bradyrhizobium O-antigen. Angew. Chem. Int. Ed. 56, 2092–2096 (2017).
Wang, Z. et al. Total synthesis and structural studies of zwitterionic Bacteroides fragilis polysaccharide A1 fragments. J. Am. Chem. Soc. 145, 14052–14063 (2023).
Deng, Z. et al. A close look at proteins: submolecular resolution of two- and three-dimensionally folded cytochrome c at surfaces. Nano Lett. 12, 2452–2458 (2012).
Anggara, K. et al. Direct observation of glycans bonded to proteins and lipids at the single-molecule level. Science 382, 219–223 (2023).
Anggara, K. et al. Exploring the molecular conformation space by soft molecule–surface collision. J. Am. Chem. Soc. 142, 21420–21427 (2020).
Seibel, J. et al. Visualizing chiral interactions in carbohydrates adsorbed on Au(111) by high-resolution STM imaging. Angew. Chem. Int. Ed. 135, e202305733 (2023).
Jonas, E., Kuhn, S. & Schlörer, N. Prediction of chemical shift in NMR: a review. Magn. Reson. Chem. 60, 1021–1031 (2022).
Pérez, S., Sarkar, A., Rivet, A., Breton, C. & Imberty, A. in Glycoinformatics Vol. 1273 (eds Lütteke, T. & Frank, M.) 241–258 (Springer, 2015).
Ogawa, Y. & Putaux, J.-L. Recent advances in electron microscopy of carbohydrate nanoparticles. Front. Chem. 10, 835663 (2022).
Ogawa, Y., Putaux, J.-L. & Nishiyama, Y. Crystallography of polysaccharides: current state and challenges. Curr. Opin. Chem. Biol. 70, 102183 (2022).
Ercius, P., Alaidi, O., Rames, M. J. & Ren, G. Electron tomography: a three-dimensional analytic tool for hard and soft materials research. Adv. Mater. 27, 5638–5663 (2015).
Baker, L. A. & Rubinstein, J. L. in Methods in Enzymology Vol. 481 (ed. Jensen, G. J.) 371–388 (Academic, 2010).
Ogawa, Y. Electron microdiffraction reveals the nanoscale twist geometry of cellulose nanocrystals. Nanoscale 11, 21767–21774 (2019).
Ophus, C. Four-dimensional scanning transmission electron microscopy (4D-STEM): from scanning nanodiffraction to ptychography and beyond. Microsc. Microanal. 25, 563–582 (2019).
Lim, J. H. et al. Structural anisotropy governs the kink formation in cellulose nanocrystals. J. Phys. Chem. Lett. 14, 3961–3969 (2023).
Wu, C., Wang, X., Chu, B., Tang, S. & Wang, Y. Self-assembly of core–corona β-glucan into stiff and metalizable nanostructures from 1D to 3D. ACS Nano 12, 10545–10553 (2018).
Numata, M. et al. Inclusion of cut and as-grown single-walled carbon nanotubes in the helical superstructure of schizophyllan and curdlan (β-1,3-glucans). J. Am. Chem. Soc. 127, 5875–5884 (2005).
Numata, M. et al. β-1,3-Glucan polysaccharide can act as a one-dimensional host to create novel silica nanofiber structures. Chem. Commun. 37, 4655–4657 (2005).
Bae, A.-H. et al. 1D arrangement of Au nanoparticles by the helical structure of schizophyllan: a unique encounter of a natural product with inorganic compounds. Angew. Chem. Int. Ed. 44, 2030–2033 (2005).
Shiraki, T., Dawn, A., Tsuchiya, Y. & Shinkai, S. Thermo- and solvent-responsive polymer complex created from supramolecular complexation between a helix-forming polysaccharide and a cationic polythiophene. J. Am. Chem. Soc. 132, 13928–13935 (2010).
Smith, P. J. et al. Enzymatic synthesis of xylan microparticles with tunable morphologies. ACS Mater. Au 2, 440–452 (2022).
Binnig, G., Quate, C. F. & Gerber, C. Atomic force microscope. Phys. Rev. Lett. 56, 930–933 (1986).
Viljoen, A. et al. Force spectroscopy of single cells using atomic force microscopy. Nat. Rev. Methods Primers 1, 63 (2021).
Abu-Lail, N. I. & Camesano, T. A. Polysaccharide properties probed with atomic force microscopy. J. Microsc. 212, 217–238 (2003).
Schefer, L., Adamcik, J., Diener, M. & Mezzenga, R. Supramolecular chiral self-assembly and supercoiling behavior of carrageenans at varying salt conditions. Nanoscale 7, 16182–16188 (2015).
Fittolani, G. et al. Synthesis of a glycan hairpin. Nat. Chem. 15, 1461–1469 (2023).
Girvin, Z. C. & Gellman, S. H. Foldamer catalysis. J. Am. Chem. Soc. 142, 17211–17223 (2020).
Levin, A. et al. Biomimetic peptide self-assembly for functional materials. Nat. Rev. Chem. 4, 615–634 (2020).
Yu, Y. et al. Oligosaccharides self-assemble and show intrinsic optical properties. J. Am. Chem. Soc. 141, 4833–4838 (2019).
Gim, S. et al. Targeted chemical modifications identify key features of carbohydrate assemblies and generate tailored carbohydrate materials. Chem. Eur. J. 27, 13139–13143 (2021).
Yao, Y., Meng, X., Li, C., Bernaerts, K. V. & Zhang, K. Tuning the chiral structures from self-assembled carbohydrate derivatives. Small 19, 2208286 (2023).
Hendrikse, S. I. S. et al. Elucidating the ordering in self-assembled glycocalyx mimicking supramolecular copolymers in water. J. Am. Chem. Soc. 141, 13877–13886 (2019).
Su, L., Hendrikse, S. I. S. & Meijer, E. W. Supramolecular glycopolymers: how carbohydrates matter in structure, dynamics, and function. Curr. Opin. Chem. Biol. 69, 102171 (2022).
Richaud, A. D., Zhao, G., Hobloss, S. & Roche, S. P. Folding in place: design of β-strap motifs to stabilize the folding of hairpins with long loops. J. Org. Chem. 86, 13535–13547 (2021).
Colesnic, D. et al. Precisely designed difunctionalized cyclodextrin produces a solid-state organic porous hierarchical supramolecular assembly. Chem. Eur. J. 29, e202300150 (2023).
Davis, M. E. & Brewster, M. E. Cyclodextrin-based pharmaceutics: past, present and future. Nat. Rev. Drug Discov. 3, 1023 (2004).
Liu, J. et al. Programmed synthesis of hepta-differentiated β-cyclodextrin: 1 out of 117655 arrangements. Angew. Chem. Int. Ed. 60, 12090–12096 (2021).
Erichsen, A., Peters, G. H. J. & Beeren, S. R. Templated enzymatic synthesis of δ-cyclodextrin. J. Am. Chem. Soc. 145, 4882–4891 (2023).
Ikuta, D. et al. Conformationally supple glucose monomers enable synthesis of the smallest cyclodextrins. Science 364, 674–677 (2019).
Li, X. et al. Promoter-controlled synthesis and conformational analysis of cyclic mannosides up to a 32-mer. Angew. Chem. Int. Ed. 62, e202307851 (2023).
Ricardo, M. G. et al. Design, synthesis, and characterization of stapled oligosaccharides. J. Am. Chem. Soc. 144, 18429–18434 (2022).
Naik, R. R. & Singamaneni, S. Introduction: bioinspired and biomimetic materials. Chem. Rev. 117, 12581–12583 (2017).
Sinha, N. J., Langenstein, M. G., Pochan, D. J., Kloxin, C. J. & Saven, J. G. Peptide design and self-assembly into targeted nanostructure and functional materials. Chem. Rev. 121, 13915–13935 (2021).
Jiang, T., Vail, O. A., Jiang, Z., Zuo, X. & Conticello, V. P. Rational design of multilayer collagen nanosheets with compositional and structural control. J. Am. Chem. Soc. 137, 7793–7802 (2015).
Tanrikulu, I. C., Forticaux, A., Jin, S. & Raines, R. T. Peptide tessellation yields micrometre-scale collagen triple helices. Nat. Chem. 8, 1008–1014 (2016).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Ke, P. C. et al. Half a century of amyloids: past, present and future. Chem. Soc. Rev. 49, 5473–5509 (2020).
Kuhlman, B. et al. Design of a novel globular protein fold with atomic-level accuracy. Science 302, 1364–1368 (2003).
Anishchenko, I. et al. De novo protein design by deep network hallucination. Nature 600, 547–552 (2021).
Kumar, P., Paterson, N. G., Clayden, J. & Woolfson, D. N. De novo design of discrete, stable 310-helix peptide assemblies. Nature 607, 387–392 (2022).
Hu, K.-N. & Tycko, R. What can solid state NMR contribute to our understanding of protein folding? Biophys. Chem. 151, 10–21 (2010).
Hamley, I. W. & Castelletto, V. Small-angle scattering techniques for peptide and peptide hybrid nanostructures and peptide-based biomaterials. Adv. Colloid Interface Sci. 318, 102959 (2023).
Clarke, D. E., Parmenter, C. D. J. & Scherman, O. A. Tunable pentapeptide self-assembled β-sheet hydrogels. Angew. Chem. Int. Ed. 57, 7709–7713 (2018).
Mitić, Ž. et al. Instrumental methods and techniques for structural and physicochemical characterization of biomaterials and bone tissue: a review. Mater. Sci. Eng. C 79, 930–949 (2017).
Baek, M. et al. Accurate prediction of protein structures and interactions using a three-track neural network. Science 373, 871–876 (2021).
Jumper, J. et al. Highly accurate protein structure prediction with AlphaFold. Nature 596, 583–589 (2021).
Hribernik, N. et al. Controlling the assembly of cellulose-based oligosaccharides through sequence modifications. Angew. Chem. Int. Ed. 62, e202310357 (2023).
Tamburrini, A., Colombo, C. & Bernardi, A. Design and synthesis of glycomimetics: recent advances. Med. Res. Rev. 40, 495–531 (2020).
Zhang, P. et al. Artificial chiral metallo-pockets including a single metal serving as structural probe and catalytic center. Chem 3, 174–191 (2017).
Breslow, R. & Dong, S. D. Biomimetic reactions catalyzed by cyclodextrins and their derivatives. Chem. Rev. 98, 1997–2012 (1998).
Szente, L. & Szemán, J. Cyclodextrins in analytical chemistry: host–guest type molecular recognition. Anal. Chem. 85, 8024–8030 (2013).
Hu, J., Seeberger, P. H. & Yin, J. Using carbohydrate-based biomaterials as scaffolds to control human stem cell fate. Org. Biomol. Chem. 14, 8648–8658 (2016).
Acknowledgements
The authors thank the Max Planck Society, the German Federal Ministry of Education and Research (BMBF, grant number 13XP5114, to M. D.), the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation; SFB 1449, project number 431232613, sub-project C2) and the European Research Council (ERC) under the Horizon Europe research and innovation programme (project number 101075357 GLYCOFOLD) for the generous financial support.
Author information
Authors and Affiliations
Contributions
M.D. conceived the article. S.D. designed the figures. All authors contributed to the discussion and writing of the article.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Reviews Materials thanks Alexei Demchenko and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Djalali, S., Yadav, N. & Delbianco, M. Towards glycan foldamers and programmable assemblies. Nat Rev Mater 9, 190–201 (2024). https://doi.org/10.1038/s41578-023-00638-x
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41578-023-00638-x